Abstract
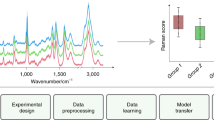
Spectroscopic techniques such as Fourier-transform infrared (FTIR) spectroscopy are used to study interactions of light with biological materials. This interaction forms the basis of many analytical assays used in disease screening/diagnosis, microbiological studies, and forensic/environmental investigations. Advantages of spectrochemical analysis are its low cost, minimal sample preparation, non-destructive nature and substantially accurate results. However, an urgent need exists for repetition and validation of these methods in large-scale studies and across different research groups, which would bring the method closer to clinical and/or industrial implementation. For this to succeed, it is important to understand and reduce the effect of random spectral alterations caused by inter-individual, inter-instrument and/or inter-laboratory variations, such as variations in air humidity and CO2 levels, and aging of instrument parts. Thus, it is evident that spectral standardization is critical to the widespread adoption of these spectrochemical technologies. By using calibration transfer procedures, in which the spectral response of a secondary instrument is standardized to resemble the spectral response of a primary instrument, different sources of variation can be normalized into a single model using computational-based methods, such as direct standardization (DS) and piecewise direct standardization (PDS); therefore, measurements performed under different conditions can generate the same result, eliminating the need for a full recalibration. Here, we have constructed a protocol for model standardization using different transfer technologies described for FTIR spectrochemical applications. This is a critical step toward the construction of a practical spectrochemical analysis model for daily routine analysis, where uncertain and random variations are present.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding authors on reasonable request.
Software availability
Outlier detection algorithm: https://doi.org/10.6084/m9.figshare.7066613.v1
References
Baker, M. J. et al. Clinical applications of infrared and Raman spectroscopy: state of play and future challenges. Analyst 143, 1735–1757 (2018).
Melin, A. M., Perromat, A. & Déléris, G. Pharmacologic application of Fourier transform IR spectroscopy: in vivo toxicity of carbon tetrachloride on rat liver. Biopolymers 57, 160–168 (2000).
Eliasson, C. & Matousek, P. Noninvasive authentication of pharmaceutical products through packaging using spatially offset Raman spectroscopy. Anal. Chem. 79, 1696–1701 (2007).
Baker, M. J. et al. Using Fourier transform IR spectroscopy to analyze biological materials. Nat. Protoc. 9, 1771–1791 (2014).
Llabjani, V. et al. Polybrominated diphenyl ether-associated alterations in cell biochemistry as determined by attenuated total reflection Fourier-transform infrared spectroscopy: a comparison with DNA-reactive and/or endocrine-disrupting agents. Environ. Sci. Technol. 43, 3356–3364 (2009).
Hofmann-Wellenhof, B., Lichtenegger, H. & Collins, J. Global Positioning System: Theory and Practice (Springer Science & Business Media, Vienna, 2012).
Morris, P. & Perkins, A. Diagnostic imaging. Lancet 379, 1525–1533 (2012).
Lee, S. S. et al. Crohn disease of the small bowel: comparison of CT enterography, MR enterography, and small-bowel follow-through as diagnostic techniques. Radiology 251, 751–761 (2009).
Lagleyre, S. et al. Reliability of high-resolution CT scan in diagnosis of otosclerosis. Otol. Neurotol. 30, 1152–1159 (2009).
Kalita, J. & Misra, U. Comparison of CT scan and MRI findings in the diagnosis of Japanese encephalitis. J. Neurol. Sci. 174, 3–8 (2000).
Schrevens, L., Lorent, N., Dooms, C. & Vansteenkiste, J. The role of PET scan in diagnosis, staging, and management of non-small cell lung cancer. Oncologist 9, 633–643 (2004).
Jagust, W., Reed, B., Mungas, D., Ellis, W. & Decarli, C. What does fluorodeoxyglucose PET imaging add to a clinical diagnosis of dementia? Neurology 69, 871–877 (2007).
Zhou, M. et al. Clinical utility of breast-specific gamma imaging for evaluating disease extent in the newly diagnosed breast cancer patient. Am. J. Surg. 197, 159–163 (2009).
Wallace, B. A. et al. Biomedical applications of synchrotron radiation circular dichroism spectroscopy: identification of mutant proteins associated with disease and development of a reference database for fold motifs. Faraday Discuss. 126, 237–243 (2004).
Greenfield, N. J. Using circular dichroism spectra to estimate protein secondary structure. Nat. Protoc. 1, 2876–2890 (2006).
Micsonai, A. et al. Accurate secondary structure prediction and fold recognition for circular dichroism spectroscopy. Proc. Natl. Acad. Sci. USA 112, E3095–E3103 (2015).
Miles, A. J. & Wallace, B. A. Circular dichroism spectroscopy of membrane proteins. Chem. Soc. Rev. 45, 4859–4872 (2016).
Brown, J. Q., Vishwanath, K., Palmer, G. M. & Ramanujam, N. Advances in quantitative UV–visible spectroscopy for clinical and pre-clinical application in cancer. Curr. Opin. Biotechnol. 20, 119–131 (2009).
Yang, P.-W. et al. Visible-absorption spectroscopy as a biomarker to predict treatment response and prognosis of surgically resected esophageal cancer. Sci. Rep. 6, 33414 (2016).
World Health Organization. Fluorescence microscopy for disease diagnosis and environmental monitoring. https://apps.who.int/iris/handle/10665/119734 (2005).
Shahzad, A. et al. Diagnostic application of fluorescence spectroscopy in oncology field: hopes and challenges. Appl. Spectrosc. Rev. 45, 92–99 (2010).
Sieroń, A. et al. The role of fluorescence diagnosis in clinical practice. Onco Targets Ther. 6, 977 (2013).
Shin, D., Vigneswaran, N., Gillenwater, A. & Richards-Kortum, R. Advances in fluorescence imaging techniques to detect oral cancer and its precursors. Future Oncol. 6, 1143–1154 (2010).
Shahzad, A. et al. Emerging applications of fluorescence spectroscopy in medical microbiology field. J. Transl. Med. 7, 99 (2009).
Möller-Hartmann, W. et al. Clinical application of proton magnetic resonance spectroscopy in the diagnosis of intracranial mass lesions. Neuroradiology 44, 371–381 (2002).
Gowda, G. N. et al. Metabolomics-based methods for early disease diagnostics. Expert Rev. Mol. Diagn. 8, 617–633 (2008).
Frisoni, G. B., Fox, N. C., Jack, C. R., Scheltens, P. & Thompson, P. M. The clinical use of structural MRI in Alzheimer disease. Nat. Rev. Neurol. 6, 67–77 (2010).
Chan, A. W. et al. 1 H-NMR urinary metabolomic profiling for diagnosis of gastric cancer. Br. J. Cancer 114, 59–62 (2016).
Palmnas, M. S. & Vogel, H. J. The future of NMR metabolomics in cancer therapy: towards personalizing treatment and developing targeted drugs? Metabolites 3, 373–396 (2013).
Patil, P. & Dasgupta, B. Role of diagnostic ultrasound in the assessment of musculoskeletal diseases. Ther. Adv. Musculoskelet. Dis. 4, 341–355 (2012).
Navani, N. et al. Lung cancer diagnosis and staging with endobronchial ultrasound-guided transbronchial needle aspiration compared with conventional approaches: an open-label, pragmatic, randomised controlled trial. Lancet Respir. Med. 3, 282–289 (2015).
Menon, U. et al. Sensitivity and specificity of multimodal and ultrasound screening for ovarian cancer, and stage distribution of detected cancers: results of the prevalence screen of the UK Collaborative Trial of Ovarian Cancer Screening (UKCTOCS). Lancet Oncol. 10, 327–340 (2009).
Smith-Bindman, R. et al. Endovaginal ultrasound to exclude endometrial cancer and other endometrial abnormalities. JAMA 280, 1510–1517 (1998).
Gajjar, K. et al. Diagnostic segregation of human brain tumours using Fourier-transform infrared and/or Raman spectroscopy coupled with discriminant analysis. Anal. Methods 5, 89–102 (2013).
Bury, D. et al. Phenotyping metastatic brain tumors applying spectrochemical analyses: segregation of different cancer types. Anal. Lett. 52, 575–587 (2019).
Hands, J. R. et al. Attenuated Total Reflection Fourier Transform Infrared (ATR-FTIR) spectral discrimination of brain tumour severity from serum samples. J. Biophotonics 7, 189–199 (2014).
Hands, J. R. et al. Brain tumour differentiation: rapid stratified serum diagnostics via attenuated total reflection Fourier-transform infrared spectroscopy. J. Neurooncol. 127, 463–472 (2016).
Walsh, M. J., Kajdacsy-Balla, A., Holton, S. E. & Bhargava, R. Attenuated total reflectance Fourier-transform infrared spectroscopic imaging for breast histopathology. Vib. Spectrosc. 60, 23–28 (2012).
Lane, R. & Seo, S. S. Attenuated total reflectance fourier transform infrared spectroscopy method to differentiate between normal and cancerous breast cells. J. Nanosci. Nanotechnol. 12, 7395–7400 (2012).
Backhaus, J. et al. Diagnosis of breast cancer with infrared spectroscopy from serum samples. Vib. Spectrosc. 52, 173–177 (2010).
Wang, J.-S. et al. FT-IR spectroscopic analysis of normal and cancerous tissues of esophagus. World J. Gastroenterol. 9, 1897–1899 (2003).
Maziak, D. E. et al. Fourier-transform infrared spectroscopic study of characteristic molecular structure in cancer cells of esophagus: an exploratory study. Cancer Detect. Prev. 31, 244–253 (2007).
McIntosh, L. M. et al. Infrared spectra of basal cell carcinomas are distinct from non-tumor-bearing skin components. J. Invest. Dermatol. 112, 951–956 (1999).
McIntosh, L. M. et al. Towards non-invasive screening of skin lesions by near-infrared spectroscopy. J. Invest. Dermatol. 116, 175–181 (2001).
Mostaço-Guidolin, L. B., Murakami, L. S., Nomizo, A. & Bachmann, L. Fourier transform infrared spectroscopy of skin cancer cells and tissues. Appl. Spectrosc. Rev. 44, 438–455 (2009).
Mordechai, S. et al. Possible common biomarkers from FTIR microspectroscopy of cervical cancer and melanoma. J. Microsc. 215, 86–91 (2004).
Hammody, Z., Sahu, R. K., Mordechai, S., Cagnano, E. & Argov, S. Characterization of malignant melanoma using vibrational spectroscopy. Sci. World J. 5, 173–182 (2005).
Kondepati, V. R., Keese, M., Mueller, R., Manegold, B. C. & Backhaus, J. Application of near-infrared spectroscopy for the diagnosis of colorectal cancer in resected human tissue specimens. Vib. Spectrosc. 44, 236–242 (2007).
Rigas, B., Morgello, S., Goldman, I. S. & Wong, P. Human colorectal cancers display abnormal Fourier-transform infrared spectra. Proc. Natl. Acad. Sci. USA 87, 8140–8144 (1990).
Yao, H., Shi, X. & Zhang, Y. The use of FTIR-ATR spectrometry for evaluation of surgical resection margin in colorectal cancer: a pilot study of 56 samples. J. Spectrosc. 2014, 4 (2014).
Lewis, P. D. et al. Evaluation of FTIR Spectroscopy as a diagnostic tool for lung cancer using sputum. BMC Cancer 10, 640 (2010).
Akalin, A. et al. Classification of malignant and benign tumors of the lung by infrared spectral histopathology (SHP). Lab. Invest. 95, 406–421 (2015).
Großerueschkamp, F. et al. Marker-free automated histopathological annotation of lung tumour subtypes by FTIR imaging. Analyst 140, 2114–2120 (2015).
Owens, G. L. et al. Vibrational biospectroscopy coupled with multivariate analysis extracts potentially diagnostic features in blood plasma/serum of ovarian cancer patients. J. Biophotonics 7, 200–209 (2014).
Gajjar, K. et al. Fourier-transform infrared spectroscopy coupled with a classification machine for the analysis of blood plasma or serum: a novel diagnostic approach for ovarian cancer. Analyst 138, 3917–3926 (2013).
Theophilou, G., Lima, K. M. G., Martin-Hirsch, P. L., Stringfellow, H. F. & Martin, F. L. ATR-FTIR spectroscopy coupled with chemometric analysis discriminates normal, borderline and malignant ovarian tissue: classifying subtypes of human cancer. Analyst 141, 585–594 (2016).
Mehrotra, R., Tyagi, G., Jangir, D. K., Dawar, R. & Gupta, N. Analysis of ovarian tumor pathology by Fourier Transform Infrared Spectroscopy. J. Ovarian Res. 3, 27 (2010).
Paraskevaidi, M. et al. Potential of mid-infrared spectroscopy as a non-invasive diagnostic test in urine for endometrial or ovarian cancer. Analyst 143, 3156–3163 (2018).
Taylor, S. E. et al. Infrared spectroscopy with multivariate analysis to interrogate endometrial tissue: a novel and objective diagnostic approach. Br. J. Cancer 104, 790–797 (2011).
Paraskevaidi, M. et al. Aluminium foil as an alternative substrate for the spectroscopic interrogation of endometrial cancer. J. Biophotonics 11, e201700372 (2018).
Gajjar, K. et al. Histology verification demonstrates that biospectroscopy analysis of cervical cytology identifies underlying disease more accurately than conventional screening: removing the confounder of discordance. PLoS ONE 9, e82416 (2014).
Walsh, M. J. et al. IR microspectroscopy: potential applications in cervical cancer screening. Cancer Lett. 246, 1–11 (2007).
Wood, B. R., Quinn, M. A., Burden, F. R. & McNaughton, D. An investigation into FTIR spectroscopy as a biodiagnostic tool for cervical cancer. Biospectroscopy 2, 143–153 (1996).
Podshyvalov, A. et al. Distinction of cervical cancer biopsies by use of infrared microspectroscopy and probabilistic neural networks. Appl. Opt. 44, 3725–3734 (2005).
Theophilou, G. et al. A biospectroscopic analysis of human prostate tissue obtained from different time periods points to a trans-generational alteration in spectral phenotype. Sci. Rep. 5, 13465 (2015).
Baker, M. J. et al. Investigating FTIR based histopathology for the diagnosis of prostate cancer. J. Biophotonics 2, 104–113 (2009).
Derenne, A., Gasper, R. & Goormaghtigh, E. The FTIR spectrum of prostate cancer cells allows the classification of anticancer drugs according to their mode of action. Analyst 136, 1134–1141 (2011).
Gazi, E. et al. A correlation of FTIR spectra derived from prostate cancer biopsies with Gleason grade and tumour stage. Eur. Urol. 50, 750–761 (2006).
Paraskevaidi, M. et al. Differential diagnosis of Alzheimer’s disease using spectrochemical analysis of blood. Proc. Natl. Acad. Sci. USA 114, E7929–E7938 (2017).
Carmona, P. et al. Discrimination analysis of blood plasma associated with Alzheimer’s disease using vibrational spectroscopy. J. Alzheimers Dis. 34, 911–920 (2013).
Carmona, P., Molina, M., López-Tobar, E. & Toledano, A. Vibrational spectroscopic analysis of peripheral blood plasma of patients with Alzheimer’s disease. Anal. Bioanal. Chem. 407, 7747–7756 (2015).
Paraskevaidi, M. et al. Blood-based near-infrared spectroscopy for the rapid low-cost detection of Alzheimer’s disease. Analyst 143, 5959–5964 (2018).
Sitole, L., Steffens, F., Krüger, T. P. J. & Meyer, D. Mid-ATR-FTIR spectroscopic profiling of HIV/AIDS sera for novel systems diagnostics in global health. OMICS 18, 513–523 (2014).
Coopman, R. et al. Glycation in human fingernail clippings using ATR-FTIR spectrometry, a new marker for the diagnosis and monitoring of diabetes mellitus. Clin. Biochem. 50, 62–67 (2017).
Scott, D. A. et al. Diabetes-related molecular signatures in infrared spectra of human saliva. Diabetol. Metab. Syndr. 2, 48 (2010).
Varma, V. K., Kajdacsy-Balla, A., Akkina, S. K., Setty, S. & Walsh, M. J. A label-free approach by infrared spectroscopic imaging for interrogating the biochemistry of diabetic nephropathy progression. Kidney Int. 89, 1153–1159 (2016).
Lechowicz, L., Chrapek, M., Gaweda, J., Urbaniak, M. & Konieczna, I. Use of Fourier-transform infrared spectroscopy in the diagnosis of rheumatoid arthritis: a pilot study. Mol. Biol. Rep. 43, 1321–1326 (2016).
Canvin, J. et al. Infrared spectroscopy: shedding light on synovitis in patients with rheumatoid arthritis. Rheumatology 42, 76–82 (2003).
Oemrawsingh, R. M. et al. Near-infrared spectroscopy predicts cardiovascular outcome in patients with coronary artery disease. J. Am. Coll. Cardiol. 64, 2510–2518 (2014).
Wang, J. et al. Near-infrared spectroscopic characterization of human advanced atherosclerotic plaques. J. Am. Coll. Cardiol. 39, 1305–1313 (2002).
Martin, M. et al. The effect of common anticoagulants in detection and quantification of malaria parasitemia in human red blood cells by ATR-FTIR spectroscopy. Analyst 142, 1192–1199 (2017).
Khoshmanesh, A. et al. Detection and quantification of early-stage malaria parasites in laboratory infected erythrocytes by attenuated total reflectance infrared spectroscopy and multivariate analysis. Anal. Chem. 86, 4379–4386 (2014).
Roy, S. et al. Simultaneous ATR-FTIR based determination of malaria parasitemia, glucose and urea in whole blood dried onto a glass slide. Anal. Chem. 89, 5238–5245 (2017).
Markus, A. P. J. et al. New technique for diagnosis and monitoring of alcaptonuria: quantification of homogentisic acid in urine with mid-infrared spectrometry. Anal. Chim. Acta 429, 287–292 (2001).
Grimard, V. et al. Phosphorylation-induced conformational changes of cystic fibrosis transmembrane conductance regulator monitored by Attenuated Total Reflection-Fourier Transform IR Spectroscopy and Fluorescence Spectroscopy. J. Biol. Chem. 279, 5528–5536 (2004).
Aksoy, C., Guliyev, A., Kilic, E., Uckan, D. & Severcan, F. Bone marrow mesenchymal stem cells in patients with beta thalassemia major: molecular analysis with attenuated total reflection-Fourier transform infrared spectroscopy study as a novel method. Stem Cells Dev. 21, 2000–2011 (2012).
Graça, G. et al. Mid-infrared (MIR) metabolic fingerprinting of amniotic fluid: a possible avenue for early diagnosis of prenatal disorders? Anal. Chim. Acta 764, 24–31 (2013).
Hasegawa, J. et al. Evaluation of placental function using near infrared spectroscopy during fetal growth restriction. J. Perinat. Med. 38, 29–32 (2010).
Theelen, T., Berendschot, T. T., Hoyng, C. B., Boon, C. J. & Klevering, B. J. Near-infrared reflectance imaging of neovascular age-related macular degeneration. Graefe’s Arch. Clin. Exp. Ophthalmol. 247, 1625–1633 (2009).
Semoun, O. et al. Infrared features of classic choroidal neovascularisation in exudative age-related macular degeneration. Br. J. Ophthalmol. 93, 182–185 (2009).
Peters, A. S. et al. Serum-infrared spectroscopy is suitable for diagnosis of atherosclerosis and its clinical manifestations. Vib. Spectrosc. 92, 20–26 (2017).
Afara, I. O., Prasadam, I., Arabshahi, Z., Xiao, Y. & Oloyede, A. Monitoring osteoarthritis progression using near infrared (NIR) spectroscopy. Sci. Rep. 7, 11463 (2017).
Bi, X. et al. Fourier transform infrared imaging and MR microscopy studies detect compositional and structural changes in cartilage in a rabbit model of osteoarthritis. Anal. Bioanal. Chem. 387, 1601–1612 (2007).
David-Vaudey, E. et al. Fourier Transform Infrared Imaging of focal lesions in human osteoarthritic cartilage. Eur. Cell. Mater. 10, 51–60 (2005).
Trevisan, J., Angelov, P. P., Carmichael, P. L., Scott, A. D. & Martin, F. L. Extracting biological information with computational analysis of Fourier-transform infrared (FTIR) biospectroscopy datasets: current practices to future perspectives. Analyst 137, 3202–3215 (2012).
Andrew Chan, K. L. & Kazarian, S. G. Attenuated total reflection Fourier-transform infrared (ATR-FTIR) imaging of tissues and live cells. Chem. Soc. Rev. 45, 1850–1864 (2016).
Pilling, M. & Gardner, P. Fundamental developments in infrared spectroscopic imaging for biomedical applications. Chem. Soc. Rev. 45, 1935–1957 (2016).
Martin, F. L. et al. Distinguishing cell types or populations based on the computational analysis of their infrared spectra. Nat. Protoc. 5, 1748–1760 (2010).
Butler, H. J. et al. Using Raman spectroscopy to characterize biological materials. Nat. Protoc. 11, 664–687 (2016).
Kong, L. et al. Characterization of bacterial spore germination using phase-contrast and fluorescence microscopy, Raman spectroscopy and optical tweezers. Nat. Protoc. 6, 625–639 (2011).
Harmsen, S., Wall, M. A., Huang, R. & Kircher, M. F. Cancer imaging using surface-enhanced resonance Raman scattering nanoparticles. Nat. Protoc. 12, 1400–1414 (2017).
Beckonert, O. et al. Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nat. Protoc. 2, 2692–2703 (2007).
Felten, J. et al. Vibrational spectroscopic image analysis of biological material using multivariate curve resolution–alternating least squares (MCR-ALS). Nat. Protoc. 10, 217–240 (2015).
Yang, H., Yang, S., Kong, J., Dong, A. & Yu, S. Obtaining information about protein secondary structures in aqueous solution using Fourier transform IR spectroscopy. Nat. Protoc. 10, 382–396 (2015).
Sreedhar, H. et al. High-definition Fourier transform infrared (FT-IR) spectroscopic imaging of human tissue sections towards improving pathology. J. Vis. Exp. 2015, 52332 (2015).
Varriale, A. et al. Fluorescence correlation spectroscopy assay for gliadin in food. Anal. Chem. 79, 4687–4689 (2007).
Song, X., Li, H., Al-Qadiri, H. M. & Lin, M. Detection of herbicides in drinking water by surface-enhanced Raman spectroscopy coupled with gold nanostructures. J. Food Meas. Charact. 7, 107–113 (2013).
Osborne, B. G. & Fearn, T. Near-infrared spectroscopy in food analysis. Encyclopedia Anal. Chem. 5, 4069–4082 (2000).
Qu, J.-H. et al. Applications of near-infrared spectroscopy in food safety evaluation and control: a review of recent research advances. Crit. Rev. Food Sci. Nutr. 55, 1939–1954 (2015).
Penido, C. A. F., Pacheco, M. T. T., Lednev, I. K. & Silveira, L. Raman spectroscopy in forensic analysis: identification of cocaine and other illegal drugs of abuse. J. Raman Spectrosc. 47, 28–38 (2016).
Ryder, A. G. Classification of narcotics in solid mixtures using principal component analysis and Raman spectroscopy. J. Forensic Sci. 47, 275–284 (2002).
Harrigan, G. G. et al. Application of high-throughput Fourier-transform infrared spectroscopy in toxicology studies: contribution to a study on the development of an animal model for idiosyncratic toxicity. Toxicol. Lett. 146, 197–205 (2004).
Choo-Smith, L.-P. et al. Investigating microbial (micro) colony heterogeneity by vibrational spectroscopy. Appl. Environ. Microbiol. 67, 1461–1469 (2001).
Helm, D., Labischinski, H., Schallehn, G. & Naumann, D. Classification and identification of bacteria by Fourier-transform infrared spectroscopy. Microbiology 137, 69–79 (1991).
Carmona, P., Monzon, M., Monleon, E., Badiola, J. J. & Monreal, J. In vivo detection of scrapie cases from blood by infrared spectroscopy. J. Gen. Virol. 86, 3425–3431 (2005).
Cui, L. et al. Functional single-cell approach to probing nitrogen-fixing bacteria in soil communities by resonance Raman spectroscopy with 15N2 labeling. Anal. Chem. 90, 5082–5089 (2018).
Lasch, P. & Naumann, D. Infrared spectroscopy in microbiology. in Encyclopedia of Analytical Chemistry (eds Brown, J. & Pawlu, T.) (Arcler Press, Oakville, ON, Canada, 2015).
Maquelin, K. et al. Identification of medically relevant microorganisms by vibrational spectroscopy. J. Microbiol. Methods 51, 255–271 (2002).
Day, J. S., Edwards, H. G., Dobrowski, S. A. & Voice, A. M. The detection of drugs of abuse in fingerprints using Raman spectroscopy I: latent fingerprints. Spectrochim. Acta A 60, 563–568 (2004).
Macleod, N. A. & Matousek, P. Emerging non-invasive raman methods in process control and forensic applications. Pharm. Res. 25, 2205–2215 (2008).
Lewis, I., Daniel, N. Jr, Chaffin, N., Griffiths, P. & Tungol, M. Raman spectroscopic studies of explosive materials: towards a fieldable explosives detector. Spectrochim. Acta A 51, 1985–2000 (1995).
Hargreaves, M. D. & Matousek, P. Threat detection of liquid explosive precursor mixtures by Spatially Offset Raman Spectroscopy (SORS). in Optics and Photonics for Counterterrorism and Crime Fighting V (ed. Lewis, C.) Proceedings of SPIE, Vol. 7486, 74860B (International Society for Optics and Photonics, Bellingham, WA, 2009).
Ali, E. M., Edwards, H. G., Hargreaves, M. D. & Scowen, I. J. Raman spectroscopic investigation of cocaine hydrochloride on human nail in a forensic context. Anal. Bioanal. Chem. 390, 1159–1166 (2008).
Vergote, G. J., Vervaet, C., Remon, J. P., Haemers, T. & Verpoort, F. Near-infrared FT-Raman spectroscopy as a rapid analytical tool for the determination of diltiazem hydrochloride in tablets. Eur. J. Pharm. Sci. 16, 63–67 (2002).
Lohr, D. et al. Non-destructive determination of carbohydrate reserves in leaves of ornamental cuttings by near-infrared spectroscopy (NIRS) as a key indicator for quality assessments. Biosyst. Eng. 158, 51–63 (2017).
Heys, K. A., Shore, R. F., Pereira, M. G. & Martin, F. L. Levels of organochlorine pesticides are associated with amyloid aggregation in apex avian brains. Environ. Sci. Technol. 51, 8672–8681 (2017).
Comino, F., Aranda, V., García-Ruiz, R. & Domínguez-Vidal, A. Infrared spectroscopy as a tool for the assessment of soil biological quality in agricultural soils under contrasting management practices. Ecol. Indic. 87, 117–126 (2018).
Eliasson, C., Macleod, N. & Matousek, P. Noninvasive detection of concealed liquid explosives using Raman spectroscopy. Anal. Chem. 79, 8185–8189 (2007).
Liu, H.-B., Zhong, H., Karpowicz, N., Chen, Y. & Zhang, X.-C. Terahertz spectroscopy and imaging for defense and security applications. Proc. IEEE 95, 1514–1527 (2007).
Golightly, R. S., Doering, W. E. & Natan, M. J. Surface-enhanced Raman spectroscopy and homeland security: a perfect match? ACS Nano 3, 2859–2869 (2009).
Wang, Y., Veltkamp, D. J. & Kowalski, B. R. Multivariate instrument standardization. Anal. Chem. 63, 2750–2756 (1991).
Brouckaert, D., Uyttersprot, J.-S., Broeckx, W. & De Beer, T. Calibration transfer of a Raman spectroscopic quantification method for the assessment of liquid detergent compositions from at-line laboratory to in-line industrial scale. Talanta 179, 386–392 (2018).
Vasconcelos de Andrade, E. W., Medeiros de Morais, C. L., Lopes da Costa, F. S. & Gomes de Lima, K. M. A multivariate control chart approach for calibration transfer between NIR spectrometers for simultaneous determination of rifampicin and isoniazid in pharmaceutical formulation. Curr. Anal. Chem. 14, 488–494 (2018).
Zamora-Rojas, E., Pérez-Marín, D., De Pedro-Sanz, E., Guerrero-Ginel, J. & Garrido-Varo, A. Handheld NIRS analysis for routine meat quality control: database transfer from at-line instruments. Chemom. Intellig. Lab. Syst. 114, 30–35 (2012).
Panchuk, V., Kirsanov, D., Oleneva, E., Semenov, V. & Legin, A. Calibration transfer between different analytical methods. Talanta 170, 457–463 (2017).
de Morais, Cd. L. M. & de Lima, K. M. G. Determination and analytical validation of creatinine content in serum using image analysis by multivariate transfer calibration procedures. Anal. Methods 7, 6904–6910 (2015).
Khaydukova, M. et al. Multivariate calibration transfer between two different types of multisensor systems. Sens. Actuators B Chem. 246, 994–1000 (2017).
Barreiro, P., Herrero, D., Hernández, N., Gracia, A. & León, L. Calibration transfer between portable and laboratory NIR spectrophotometers. Acta Hortic. 802, 373–378 (2008).
Sulub, Y., LoBrutto, R., Vivilecchia, R. & Wabuyele, B. W. Content uniformity determination of pharmaceutical tablets using five near-infrared reflectance spectrometers: a process analytical technology (PAT) approach using robust multivariate calibration transfer algorithms. Anal. Chim. Acta 611, 143–150 (2008).
Zhang, L., Small, G. W. & Arnold, M. A. Multivariate calibration standardization across instruments for the determination of glucose by Fourier transform near-infrared spectrometry. Anal. Chem. 75, 5905–5915 (2003).
Koehler, F. W. IV, Small, G. W., Combs, R. J., Knapp, R. B. & Kroutil, R. T. Calibration transfer algorithm for automated qualitative analysis by passive Fourier transform infrared spectrometry. Anal. Chem. 72, 1690–1698 (2000).
Martens, H., Høy, M., Wise, B. M., Bro, R. & Brockhoff, P. B. Pre-whitening of data by covariance-weighted pre-processing. J. Chemom. 17, 153–165 (2003).
Feudale, R. N. et al. Transfer of multivariate calibration models: a review. Chemom. Intellig. Lab. Syst. 64, 181–192 (2002).
Woody, N. A., Feudale, R. N., Myles, A. J. & Brown, S. D. Transfer of multivariate calibrations between four near-infrared spectrometers using orthogonal signal correction. Anal. Chem. 76, 2595–2600 (2004).
Greensill, C., Wolfs, P., Spiegelman, C. & Walsh, K. Calibration transfer between PDA-based NIR spectrometers in the NIR assessment of melon soluble solids content. Appl. Spectrosc. 55, 647–653 (2001).
Sjöblom, J., Svensson, O., Josefson, M., Kullberg, H. & Wold, S. An evaluation of orthogonal signal correction applied to calibration transfer of near infrared spectra. Chemom. Intellig. Lab. Syst. 44, 229–244 (1998).
Rodrigues, R. R. et al. Evaluation of calibration transfer methods using the ATR-FTIR technique to predict density of crude oil. Chemom. Intellig. Lab. Syst. 166, 7–13 (2017).
Andrews, D. T. & Wentzell, P. D. Applications of maximum likelihood principal component analysis: incomplete data sets and calibration transfer. Anal. Chim. Acta 350, 341–352 (1997).
Bouveresse, E., Massart, D. & Dardenne, P. Calibration transfer across near-infrared spectrometric instruments using Shenk’s algorithm: effects of different standardisation samples. Anal. Chim. Acta 297, 405–416 (1994).
Shenk, J. S. & Westerhaus, M. O. Populations structuring of near infrared spectra and modified partial least squares regression. Crop Sci. 31, 1548–1555 (1991).
Paatero, P. & Tapper, U. Positive matrix factorization: A non-negative factor model with optimal utilization of error estimates of data values. Environmetrics 5, 111–126 (1994).
Xie, Y. & Hopke, P. K. Calibration transfer as a data reconstruction problem. Anal. Chim. Acta 384, 193–205 (1999).
Goodacre, R. et al. On mass spectrometer instrument standardization and interlaboratory calibration transfer using neural networks. Anal. Chim. Acta 348, 511–532 (1997).
Chen, W.-R., Bin, J., Lu, H.-M., Zhang, Z.-M. & Liang, Y.-Z. Calibration transfer via an extreme learning machine auto-encoder. Analyst 141, 1973–1980 (2016).
Hu, Y., Peng, S., Bi, Y. & Tang, L. Calibration transfer based on maximum margin criterion for qualitative analysis using Fourier transform infrared spectroscopy. Analyst 137, 5913–5918 (2012).
Fan, W., Liang, Y., Yuan, D. & Wang, J. Calibration model transfer for near-infrared spectra based on canonical correlation analysis. Anal. Chim. Acta 623, 22–29 (2008).
Isabelle, M. et al. Multi-centre Raman spectral mapping of oesophageal cancer tissues: a study to assess system transferability. Faraday Discuss. 187, 87–103 (2016).
Wang, Z., Dean, T. & Kowalski, B. R. Additive background correction in multivariate instrument standardization. Anal. Chem. 67, 2379–2385 (1995).
Kennard, R. W. & Stone, L. A. Computer aided design of experiments. Technometrics 11, 137–148 (1969).
Palonpon, A. F. et al. Raman and SERS microscopy for molecular imaging of live cells. Nat. Protoc. 8, 677–692 (2013).
Witze, E. S., Old, W. M., Resing, K. A. & Ahn, N. G. Mapping protein post-translational modifications with mass spectrometry. Nat. Methods 4, 798–806 (2007).
Aebersold, R. & Mann, M. Mass spectrometry-based proteomics. Nature 422, 198–207 (2003).
Pence, I. & Mahadevan-Jansen, A. Clinical instrumentation and applications of Raman spectroscopy. Chem. Soc. Rev. 45, 1958–1979 (2016).
Ibrahim, O. et al. Improved protocols for pre-processing Raman spectra of formalin fixed paraffin preserved tissue sections. Anal. Methods 9, 4709–4717 (2017).
Tfayli, A. et al. Digital dewaxing of Raman signals: discrimination between nevi and melanoma spectra obtained from paraffin-embedded skin biopsies. Appl. Spectrosc. 63, 564–570 (2009).
Byrne, H. J., Knief, P., Keating, M. E. & Bonnier, F. Spectral pre and post processing for infrared and Raman spectroscopy of biological tissues and cells. Chem. Soc. Rev. 45, 1865–1878 (2016).
Meade, A. D. et al. Studies of chemical fixation effects in human cell lines using Raman microspectroscopy. Anal. Bioanal. Chem. 396, 1781–1791 (2010).
Baker, M. J. et al. Developing and understanding biofluid vibrational spectroscopy: a critical review. Chem. Soc. Rev. 45, 1803–1818 (2016).
Bonifacio, A., Cervo, S. & Sergo, V. Label-free surface-enhanced Raman spectroscopy of biofluids: fundamental aspects and diagnostic applications. Anal. Bioanal. Chem. 407, 8265–8277 (2015).
Mitchell, A. L., Gajjar, K. B., Theophilou, G., Martin, F. L. & Martin-Hirsch, P. L. Vibrational spectroscopy of biofluids for disease screening or diagnosis: translation from the laboratory to a clinical setting. J. Biophotonics 7, 153–165 (2014).
Lovergne, L. et al. Biofluid infrared spectro-diagnostics: pre-analytical considerations for clinical applications. Faraday Discuss. 187, 521–537 (2016).
Bonifacio, A. et al. Surface-enhanced Raman spectroscopy of blood plasma and serum using Ag and Au nanoparticles: a systematic study. Anal. Bioanal. Chem. 406, 2355–2365 (2014).
Paraskevaidi, M., Martin-Hirsch, P. L. & Martin, F. L. ATR-FTIR spectroscopy tools for medical diagnosis and disease investigation. in Nanotechnology Characterization Tools for Biosensing and Medical Diagnosis (ed. Kumar, C. S. S. R.) 163–211 (Springer, Berlin, 2017).
Mitchell, B. L., Yasui, Y., Li, C. I., Fitzpatrick, A. L. & Lampe, P. D. Impact of freeze–thaw cycles and storage time on plasma samples used in mass spectrometry based biomarker discovery projects. Cancer Inform. 1, 98–104 (2005).
Glassford, S. E., Byrne, B. & Kazarian, S. G. Recent applications of ATR FTIR spectroscopy and imaging to proteins. Biochim. Biophys. Acta 1834, 2849–2858 (2013).
Kundu, J., Le, F., Nordlander, P. & Halas, N. J. Surface enhanced infrared absorption (SEIRA) spectroscopy on nanoshell aggregate substrates. Chem. Phys. Lett. 452, 115–119 (2008).
Jones, S., Carley, S. & Harrison, M. An introduction to power and sample size estimation. Emerg. Med. J. 20, 453–458 (2003).
Beebe, K. R., Pell, R. J. & Seasholtz, M. B. Chemometrics: A Practical Guide Vol. 4 (Wiley, New York,1998).
Pavia, D. L., Lampman, G. M., Kriz, G. S. & Vyvyan, J. A. Introduction to Spectroscopy (Cengage Learning, Belmont, CA, 2008).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction 2nd edn (Springer, New York, 2009).
Bro, R. & Smilde, A. K. Principal component analysis. Anal. Methods 6, 2812–2831 (2014).
Martin, F. L. et al. Identifying variables responsible for clustering in discriminant analysis of data from infrared microspectroscopy of a biological sample. J. Comput. Biol. 14, 1176–1184 (2007).
Martens, H. & Martens, M. Modified Jack-knife estimation of parameter uncertainty in bilinear modelling by partial least squares regression (PLSR). Food Qual. Prefer. 11, 5–16 (2000).
Rousseeuw, P. J. & Hubert, M. Robust statistics for outlier detection. Wiley Interdiscip. Rev. Data Min. Knowl. Discov. 1, 73–79 (2011).
Jiang, F., Liu, G., Du, J. & Sui, Y. Initialization of K-modes clustering using outlier detection techniques. Inf. Sci. 332, 167–183 (2016).
Domingues, R., Filippone, M., Michiardi, P. & Zouaoui, J. A comparative evaluation of outlier detection algorithms: experiments and analyses. Pattern Recognit. 74, 406–421 (2018).
Bakeev, K. A. Process Analytical Technology: Spectroscopic Tools and Implementation Strategies for the Chemical and Pharmaceutical Industries 2nd edn (John Wiley & Sons, Chichester, UK, 2010).
Kuligowski, J., Quintás, G., Herwig, C. & Lendl, B. A rapid method for the differentiation of yeast cells grown under carbon and nitrogen-limited conditions by means of partial least squares discriminant analysis employing infrared micro-spectroscopic data of entire yeast cells. Talanta 99, 566–573 (2012).
Morais, C. L. & Lima, K. M. Comparing unfolded and two-dimensional discriminant analysis and support vector machines for classification of EEM data. Chemom. Intell. Lab. Syst. 170, 1–2 (2017).
Seasholtz, M. B. & Kowalski, B. The parsimony principle applied to multivariate calibration. Anal. Chim. Acta 277, 165–177 (1993).
Morais, C. L. & Lima, K. M. Principal component analysis with linear and quadratic discriminant analysis for identification of cancer samples based on mass spectrometry. J. Braz. Chem. Soc. 29, 472–481 (2017).
Brereton, R. G. & Lloyd, G. R. Partial least squares discriminant analysis: taking the magic away. J. Chemom. 28, 213–225 (2014).
Hibbert, D. B. Vocabulary of concepts and terms in chemometrics (IUPAC Recommendations 2016). Pure Appl. Chem. 88, 407–443 (2016).
McCall, J. Genetic algorithms for modelling and optimisation. J. Comput. Appl. Math. 184, 205–222 (2005).
Soares, S. F. C., Gomes, A. A., Araujo, M. C. U., Galvão Filho, A. R. & Galvão, R. K. H. The successive projections algorithm. Trends Anal. Chem. 42, 84–98 (2013).
Kamandar, M. & Ghassemian, H. Maximum relevance, minimum redundancy feature extraction for hyperspectral images. in 2010 18th Iranian Conference on Electrical Engineering: Proceedings 254–259 (IEEE, Isfahan, Iran, 2010).
Sattlecker, M., Stone, N., Smith, J. & Bessant, C. Assessment of robustness and transferability of classification models built for cancer diagnostics using Raman spectroscopy. J. Raman Spectrosc. 42, 897–903 (2011).
Guo, S. et al. Towards an improvement of model transferability for Raman spectroscopy in biological applications. Vib. Spectrosc. 91, 111–118 (2017).
Luo, X. et al. Calibration transfer across near infrared spectrometers for measuring hematocrit in the blood of grazing cattle. J. Near Infrared Spectrosc. 25, 15–25 (2017).
Vaughan, A. A. et al. Liquid chromatography–mass spectrometry calibration transfer and metabolomics data fusion. Anal. Chem. 84, 9848–9857 (2012).
Rodriguez, J. D., Westenberger, B. J., Buhse, L. F. & Kauffman, J. F. Standardization of Raman spectra for transfer of spectral libraries across different instruments. Analyst 136, 4232–4240 (2011).
Yu, B., Ji, H. & Kang, Y. Standardization of near infrared spectra based on multi-task learning. Spectrosc. Lett. 49, 23–29 (2016).
Ni, L., Han, M., Luan, S. & Zhang, L. Screening wavelengths with consistent and stable signals to realize calibration model transfer of near infrared spectra. Spectrochim. Acta A 206, 350–358 (2019).
Hu, R. & Xia, J. Calibration transfer of near infrared spectroscopy based on DS algorithm. in 2011 International Conference on Electric Information and Control Engineering (ICEICE) 3062–3065 (IEEE, Wuhan, China).
Forina, M. et al. Transfer of calibration function in near-infrared spectroscopy. Chemom. Intellig. Lab. Syst. 27, 189–203 (1995).
Xiao, H. et al. Comparison of benchtop Fourier-transform (FT) and portable grating scanning spectrometers for determination of total soluble solid contents in single grape berry (Vitis vinifera L.) and calibration transfer. Sensors 17, 2693 (2017).
Yahaya, O., MatJafri, M., Aziz, A. & Omar, A. Visible spectroscopy calibration transfer model in determining pH of Sala mangoes. J. Instrum. 10, T05002 (2015).
Bin, J., Li, X., Fan, W., Zhou, J.-h & Wang, C.-w Calibration transfer of near-infrared spectroscopy by canonical correlation analysis coupled with wavelet transform. Analyst 142, 2229–2238 (2017).
Monakhova, Y. B. & Diehl, B. W. Transfer of multivariate regression models between high-resolution NMR instruments: application to authenticity control of sunflower lecithin. Magn. Reson. Chem. 54, 712–717 (2016).
Zuo, Q., Xiong, S., Chen, Z.-P., Chen, Y. & Yu, R.-Q. A novel calibration strategy based on background correction for quantitative circular dichroism spectroscopy. Talanta 174, 320–324 (2017).
Savitzky, A. & Golay, M. J. Smoothing and differentiation of data by simplified least squares procedures. Anal. Chem. 36, 1627–1639 (1964).
Geladi, P., MacDougall, D. & Martens, H. Linearization and scatter-correction for near-infrared reflectance spectra of meat. Appl. Spectrosc. 39, 491–500 (1985).
Barnes, R., Dhanoa, M. S. & Lister, S. J. Standard normal variate transformation and de-trending of near-infrared diffuse reflectance spectra. Appl. Spectrosc. 43, 772–777 (1989).
Brereton, R. G. Chemometrics: Data Analysis for the Laboratory and Chemical Plant (John Wiley & Sons, Chichester, UK, 2003).
Dixon, S. J. & Brereton, R. G. Comparison of performance of five common classifiers represented as boundary methods: Euclidean Distance to Centroids, Linear Discriminant Analysis, Quadratic Discriminant Analysis, Learning Vector Quantization and Support Vector Machines, as dependent on data structure. Chemom. Intell. Lab. Syst. 95, 1–17 (2009).
Cover, T. & Hart, P. Nearest neighbor pattern classification. IEEE Trans. Inf. Theory 13, 21–27 (1967).
Cortes, C. & Vapnik, V. Support-vector networks. Mach. Learn. 20, 273–297 (1995).
Abraham, A. Artificial neural networks. in Handbook of Measuring System Design (eds Sydenham, P. H. & Thorn, R.) (John Wiley & Sons, Chichester, UK, 2005).
Fawagreh, K., Gaber, M. M. & Elyan, E. Random forests: from early developments to recent advancements. Syst. Sci. Control Eng. 2, 602–609 (2014).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Acknowledgements
C.L.M.M. thanks Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) - Brazil (grant 88881.128982/2016-01) for financial support. The work in the laboratory of F.L.M. was supported in part by The Engineering and Physical Sciences Research Council (EPSRC; grant nos: EP/K023349/1 and EP/K023373/1). M.P. acknowledges the Rosemere Cancer Foundation for funding.
Author information
Authors and Affiliations
Contributions
F.L.M. is the principal investigator who conceived and developed the idea for the article; C.L.M.M. and M.P. wrote the manuscript. L.C., N.J.F., M.I., K.M.G.L., P.L.M.-H., H.S., J.T., M.J.W., D.Z. and Y.-G.Z. contributed recommendations and provided feedback and changes to the manuscript, and C.L.M.M., M.P. and F.L.M. brought together the text and finalized the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information: Nature Protocols thanks Åsmund Rinnan and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Martin, F. L. et al. Nat. Protoc. 5, 1748–1760 (2010): https://doi.org/10.1038/nprot.2010.133
Baker, M. J. et al. Nat. Protoc. 9, 1771–1791 (2014): https://doi.org/10.1038/nprot.2014.110
Medeiros de Morais, C. L. & de Lima, K. M. G. Anal. Methods 7, 6904–6910 (2015): https://doi.org/10.1039/C5AY01369K
Vasconcelos de Andrade, E. W. et al. Curr. Anal. Chem. 14, 488–494 (2018): https://doi.org/10.2174/1573411014666171212141909
Integrated supplementary information
Supplementary Figure 1 IR spectra of the same type of samples measured by different ATR-FIR spectrometers at the same institution.
a–d, Average (a) raw and (b) preprocessed spectra for healthy control samples, and average (c) raw and (d) preprocessed spectra for cancer samples across three different instruments (A, B and C).
Supplementary Figure 2 PCA scores for preprocessed spectra acquired by different ATR-FIR spectrometers at the same institution and outlier detection test.
a, PCA scores for healthy control samples according to the instrument used for spectra acquisition (A, B and C). b, PCA scores for cancer samples according to the instrument used for spectra acquisition (A, B and C). c, Hotelling’s T2 versus Q residuals test for healthy control samples according to the instrument used for spectra acquisition (A, B and C) based on a PCA using 5 PCs (94.77% cumulative variance). d, Hotelling’s T2 versus Q residuals test for cancer samples according to the instrument used for spectra acquisition (A, B and C) based on a PCA using 5 PCs (92.96% cumulative variance). Circled samples in c and d indicate outliers removed. Confidence ellipse was 95%, depicted in blue in a and b.
Supplementary Figure 3 PCA loadings for preprocessed spectra acquired by different ATR-FIR spectrometers at the same institution.
a, PCA loadings for healthy control samples measured in different instruments (A, B and C). b, PCA loadings for cancer samples measured in different instruments (A, B and C).
Supplementary Figure 4 IR spectra of healthy control samples measured by different operators at the same institution.
a,b, Average (a) raw and (b) pre-processed spectra for healthy control samples acquired with instrument A depending on the operator. c,d, Average (c) raw and (d) preprocessed spectra for healthy control samples acquired with instrument B depending on the operator. e,f, Average (e) raw and (f) preprocessed spectra for healthy control samples acquired with instrument C, varying the operator.
Supplementary Figure 5 IR spectra of ovarian cancer samples measured by different operators at the same institution.
a,b, Average (a) raw and (b) preprocessed spectra for cancer samples acquired with instrument A depending on the operator. c,d, Average (c) raw and (d) preprocessed spectra for cancer samples acquired with instrument B depending on the operator. e,f, Average (e) raw and (f) preprocessed spectra for cancer samples acquired with instrument C depending on the operator.
Supplementary Figure 6 PCA scores for preprocessed spectra acquired by different operators at the same institution.
a,b, PCA scores for (a) healthy control and (b) cancer samples acquired with instrument A depending on the operator. c,d, PCA scores for (c) healthy control and (d) cancer samples acquired with instrument B depending on the operator. e,f, PCA scores for (e) healthy control and (f) cancer samples acquired with instrument C depending on the operator. Confidence ellipse was 95%, depicted in blue.
Supplementary Figure 7 Outlier detection test for healthy controls and ovarian cancer samples.
a, Hotelling’s T2 versus Q residuals test based on a PCA using 8 PCs (99.07% cumulative variance) for healthy control samples depending on the instrument for spectra acquisition (A, B and C) used by operator 2. b, Hotelling’s T2 versus Q residuals test based on a PCA using 5 PCs (96.92% cumulative variance) for cancer samples depending on the instrument for spectra acquisition (A, B and C) used by operator 2. Circled sample in a indicates an outlier removed. The Hotelling’s T2 versus Q residuals test for operator 1 is depicted in Supplementary Fig. 2c,d.
Supplementary Figure 8 PCA scores for healthy controls (HC) and ovarian cancer (OC) samples based on the spectra acquired by both operators (1 and 2) and by all instruments (A, B and C).
Confidence ellipse at a 95% confidence level is depicted in blue.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Methods
Rights and permissions
About this article
Cite this article
Morais, C.L.M., Paraskevaidi, M., Cui, L. et al. Standardization of complex biologically derived spectrochemical datasets. Nat Protoc 14, 1546–1577 (2019). https://doi.org/10.1038/s41596-019-0150-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0150-x
This article is cited by
-
Spectrochemical analysis of blood combined with chemometric techniques for detecting osteosarcopenia
Scientific Reports (2023)
-
Rapid quantification of lignocellulose composition in rice straw varieties using artificial neural networks and FTIR spectroscopic data
Biomass Conversion and Biorefinery (2023)
-
Regional differences in clonal Japanese knotweed revealed by chemometrics-linked attenuated total reflection Fourier-transform infrared spectroscopy
BMC Plant Biology (2021)
-
Chemometric analysis in Raman spectroscopy from experimental design to machine learning–based modeling
Nature Protocols (2021)
-
Distinguishing active from quiescent disease in ANCA-associated vasculitis using attenuated total reflection Fourier-transform infrared spectroscopy
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.