Abstract
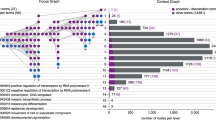
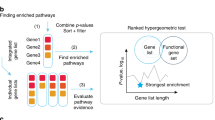
Pathway enrichment analysis helps researchers gain mechanistic insight into gene lists generated from genome-scale (omics) experiments. This method identifies biological pathways that are enriched in a gene list more than would be expected by chance. We explain the procedures of pathway enrichment analysis and present a practical step-by-step guide to help interpret gene lists resulting from RNA-seq and genome-sequencing experiments. The protocol comprises three major steps: definition of a gene list from omics data, determination of statistically enriched pathways, and visualization and interpretation of the results. We describe how to use this protocol with published examples of differentially expressed genes and mutated cancer genes; however, the principles can be applied to diverse types of omics data. The protocol describes innovative visualization techniques, provides comprehensive background and troubleshooting guidelines, and uses freely available and frequently updated software, including g:Profiler, Gene Set Enrichment Analysis (GSEA), Cytoscape and EnrichmentMap. The complete protocol can be performed in ~4.5 h and is designed for use by biologists with no prior bioinformatics training.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout













Similar content being viewed by others
Data availability
The protocol uses publicly available software packages (GSEA v.3.0 or higher, g:Profiler, Enrichment Map v.3.0 or higher, Cytoscape v.3.6.0 or higher) and custom R scripts that apply publicly available R packages (edgeR, Roast, Limma, Camera). Custom scripts are available in the Supplementary Protocols and at our GitHub web sites (https://github.com/BaderLab/Cytoscape_workflows/tree/master/EnrichmentMapPipeline and https://baderlab.github.io/Cytoscape_workflows/EnrichmentMapPipeline/index.html).
References
Lander, E. S. Initial impact of the sequencing of the human genome. Nature 470, 187–197 (2011).
Stephens, Z. D. et al. Big data: astronomical or genomical? PLoS Biol. 13, e1002195 (2015).
Mack, S. C. et al. Epigenomic alterations define lethal CIMP-positive ependymomas of infancy. Nature 506, 445–450 (2014).
Pinto, D. et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature 466, 368–372 (2010).
Pinto, D. et al. Convergence of genes and cellular pathways dysregulated in autism spectrum disorders. Am. J. Hum. Genet. 94, 677–694 (2014).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Verhaak, R. G. et al. Prognostically relevant gene signatures of high-grade serous ovarian carcinoma. J. Clin. Invest. 123, 517–525 (2013).
The Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615 (2011).
Cline, M. S. et al. Integration of biological networks and gene expression data using Cytoscape. Nat. Protoc. 2, 2366–2382 (2007).
Creixell, P. et al. Pathway and network analysis of cancer genomes. Nat Methods 12, 615–621 (2015).
Wadi, L., Meyer, M., Weiser, J., Stein, L. D. & Reimand, J. Impact of outdated gene annotations on pathway enrichment analysis. Nat. Methods 13, 705–706 (2016).
Reyna, M. A. et al. Pathway and network analysis of more than 2,500 whole cancer genomes. Preprint at https://www.biorxiv.org/content/early/2018/08/07/385294 (2018).
Reimand, J. et al. g:Profiler-a web server for functional interpretation of gene lists (2016 update). Nucleic Acids Res. 44, W83–89 (2016).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Merico, D., Isserlin, R., Stueker, O., Emili, A. & Bader, G. D. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS ONE 5, e13984 (2010).
Anders, S. et al. Count-based differential expression analysis of RNA sequencing data using R and Bioconductor. Nat. Protoc. 8, 1765–1786 (2013).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Silva, T. S. & Richard, N. Visualization and differential analysis of protein expression data using R. Methods Mol. Biol. 1362, 105–118 (2016).
Schubert, O. T., Rost, H. L., Collins, B. C., Rosenberger, G. & Aebersold, R. Quantitative proteomics: challenges and opportunities in basic and applied research. Nat. Protoc. 12, 1289–1294 (2017).
MacArthur, D. G. et al. Guidelines for investigating causality of sequence variants in human disease. Nature 508, 469–476 (2014).
Gonzalez-Perez, A. et al. Computational approaches to identify functional genetic variants in cancer genomes. Nat. Methods 10, 723–729 (2013).
Yang, H. & Wang, K. Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat. Protoc. 10, 1556–1566 (2015).
Assenov, Y. et al. Comprehensive analysis of DNA methylation data with RnBeads. Nat. Methods 11, 1138–1140 (2014).
Laird, P. W. Principles and challenges of genomewide DNA methylation analysis. Nat. Rev. Genet. 11, 191–203 (2010).
Rapaport, F. et al. Comprehensive evaluation of differential gene expression analysis methods for RNA-seq data. Genome Biol. 14, R95 (2013).
Conesa, A. et al. A survey of best practices for RNA-seq data analysis. Genome Biol. 17, 13 (2016).
Bullard, J. H., Purdom, E., Hansen, K. D. & Dudoit, S. Evaluation of statistical methods for normalization and differential expression in mRNA-Seq experiments. BMC Bioinformatics 11, 94 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Smyth, G. K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol 3, Article3 (2004).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Hochberg, Y. & Benjamini, Y. More powerful procedures for multiple significance testing. Stat. Med. 9, 811–818 (1990).
Chen, J., Bardes, E. E., Aronow, B. J. & Jegga, A. G. ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 37, W305–W311 (2009).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Mi, H., Muruganujan, A. & Thomas, P. D. PANTHER in 2013: modeling the evolution of gene function, and other gene attributes, in the context of phylogenetic trees. Nucleic Acids Res. 41, D377–D386 (2013).
Reimand, J., Kull, M., Peterson, H., Hansen, J. & Vilo, J. g:Profiler—a web-based toolset for functional profiling of gene lists from large-scale experiments. Nucleic Acids Res. 35, W193–W200 (2007).
Bindea, G. et al. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25, 1091–1093 (2009).
Maere, S., Heymans, K. & Kuiper, M. BiNGO: a Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 21, 3448–3449 (2005).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
Wang, J., Duncan, D., Shi, Z. & Zhang, B. WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): update 2013. Nucleic Acids Res. 41, W77–W83 (2013).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Fabregat, A. et al. The Reactome pathway Knowledgebase. Nucleic Acids Res. 46, D649–D655 (2018).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40, D109–D114 (2012).
Kelder, T. et al. WikiPathways: building research communities on biological pathways. Nucleic Acids Res. 40, D1301–D1307 (2012).
Kutmon, M. et al. PathVisio 3: an extendable pathway analysis toolbox. PLoS Comput. Biol. 11, e1004085 (2015).
Szklarczyk, D. et al. STRINGv10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Warde-Farley, D. et al. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 38, W214–W220 (2010).
Lechman, E. R. et al. Attenuation of miR-126 activity expands HSC in vivo without exhaustion. Cell Stem Cell 11, 799–811 (2012).
Jhas, B. et al. Metabolic adaptation to chronic inhibition of mitochondrial protein synthesis in acute myeloid leukemia cells. PLoS ONE 8, e58367 (2013).
Ballouz, S., Pavlidis, P. & Gillis, J. Using predictive specificity to determine when gene set analysis is biologically meaningful. Nucleic Acids Res. 45, e20 (2017).
Krzywinski, M. & Altman, N. Power and sample size. Nat. Methods 10, 1139–1140 (2013).
Liu, Y., Zhou, J. & White, K. P. RNA-seq differential expression studies: more sequence or more replication? Bioinformatics 30, 301–304 (2014).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Fabregat, A. et al. The Reactome pathway Knowledgebase. Nucleic Acids Res. 44, D481–D487 (2016).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 44, D471–D480 (2016).
Kandasamy, K. et al. NetPath: a public resource of curated signal transduction pathways. Genome Biol. 11, R3 (2010).
Rhee, S. Y., Wood, V., Dolinski, K. & Draghici, S. Use and misuse of the gene ontology annotations. Nat. Rev. Genet. 9, 509–515 (2008).
Skunca, N., Altenhoff, A. & Dessimoz, C. Quality of computationally inferred gene ontology annotations. PLoS Comput. Biol. 8, e1002533 (2012).
Wojtowicz, E. E. et al. Ectopic miR-125a expression induces long-term repopulating stem cell capacity in mouse and human hematopoietic progenitors. Cell Stem Cell 19, 383–396 (2016).
Tong, J. et al. Integrated analysis of proteome, phosphotyrosine-proteome, tyrosine-kinome, and tyrosine-phosphatome in acute myeloid leukemia. Proteomics 17, 1600361 (2017).
Kamdar, S. N. et al. Dynamic interplay between locus-specific DNA methylation and hydroxymethylation regulates distinct biological pathways in prostate carcinogenesis. Clin. Epigenetics 8, 32 (2016).
Liu, Y. et al. Metabolomic profiling in liver of adiponectin-knockout mice uncovers lysophospholipid metabolism as an important target of adiponectin action. Biochem. J. 469, 71–82 (2015).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Raychaudhuri, S. et al. Accurately assessing the risk of schizophrenia conferred by rare copy-number variation affecting genes with brain function. PLoS Genet. 6, e1001097 (2010).
Lee, P. H., O’Dushlaine, C., Thomas, B. & Purcell, S. M. INRICH: interval-based enrichment analysis for genome-wide association studies. Bioinformatics 28, 1797–1799 (2012).
Khatri, P., Sirota, M. & Butte, A. J. Ten years of pathway analysis: current approaches and outstanding challenges. PLoS Comput. Biol. 8, e1002375 (2012).
Wu, D. & Smyth, G. K. Camera: a competitive gene set test accounting for inter-gene correlation. Nucleic Acids Res. 40, e133 (2012).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, R14 (2010).
Gu, Z. & Wang, J. CePa: an R package for finding significant pathways weighted by multiple network centralities. Bioinformatics 29, 658–660 (2013).
Fang, Z., Tian, W. & Ji, H. A network-based gene-weighting approach for pathway analysis. Cell Res. 22, 565–580 (2012).
Farfan, F., Ma, J., Sartor, M. A., Michailidis, G. & Jagadish, H. V. THINK Back: KNowledge-based Interpretation of High Throughput data. BMC Bioinformatics 13(Suppl. 2), S4 (2012).
Tarca, A. L. et al. A novel signaling pathway impact analysis. Bioinformatics 25, 75–82 (2009).
Draghici, S. et al. A systems biology approach for pathway level analysis. Genome Res. 17, 1537–1545 (2007).
Glaab, E., Baudot, A., Krasnogor, N., Schneider, R. & Valencia, A. EnrichNet: network-based gene set enrichment analysis. Bioinformatics 28, i451–i457 (2012).
Schaefer, C. F. et al. PID: the Pathway Interaction Database. Nucleic Acids Res. 37, D674–D679 (2009).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Bader, G. D., Cary, M. P. & Sander, C. Pathguide: a pathway resource list. Nucleic Acids Res. 34, D504–D506 (2006).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Tavazoie, S., Hughes, J. D., Campbell, M. J., Cho, R. J. & Church, G. M. Systematic determination of genetic network architecture. Nat. Genet. 22, 281–285 (1999).
Goeman, J. J. & Bühlmann, P. Analyzing gene expression data in terms of gene sets: methodological issues. Bioinformatics 23, 980–987 (2007).
Bansal, V., Libiger, O., Torkamani, A. & Schork, N. J. Statistical analysis strategies for association studies involving rare variants. Nat. Rev. Genet. 11, 773–785 (2010).
Acknowledgements
The authors are grateful to J. Mesirov for comments on the manuscript. This project was supported by an Investigator Award to J.R. from the Ontario Institute for Cancer Research through funding from the Government of Ontario and by a Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery Grant to J.R. (RGPIN-2016-06485). This work was supported by US National Institutes of Health grants P41 GM103504, R01 GM070743, U41 HG006623 and R01 CA121941 to G.D.B.
Author information
Authors and Affiliations
Contributions
J.R., R.I., V.V., A.R., D.M. and G.D.B. wrote the manuscript. R.I. created the step-by-step protocols, figures, R scripts and R notebooks, except for g:Profiler (J.R.). M.K. and C.T.-L. developed EnrichmentMap 3.0 and AutoAnnotate Cytoscape applications. L.W., M.M., J.W., C.X. and V.V. tested the protocol. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Pinto, D. et al. Nature 466, 368–372 (2010): https://doi.org/10.1038/nature09146
Pajtler, K. W. et al. Cancer Cell 27, P728–P743 (2015): https://doi.org/10.1016/j.ccell.2015.04.002
Cavalli, F. M. G. et al. Cancer Cell 31, P737–P754 (2017): https://doi.org/10.1016/j.ccell.2017.05.005
Supplementary information
Supplementary Tables and Methods
Supplementary Tables 1–13 and Supplementary Protocols 1–4
Rights and permissions
About this article
Cite this article
Reimand, J., Isserlin, R., Voisin, V. et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat Protoc 14, 482–517 (2019). https://doi.org/10.1038/s41596-018-0103-9
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-018-0103-9
This article is cited by
-
Large-scale bioreactor production of extracellular vesicles from mesenchymal stromal cells for treatment of acute radiation syndrome
Stem Cell Research & Therapy (2024)
-
Decreased TLR7 expression was associated with airway eosinophilic inflammation and lung function in asthma: evidence from machine learning approaches and experimental validation
European Journal of Medical Research (2024)
-
Cord blood-derived biologics lead to robust axonal regeneration in benzalkonium chloride-injured mouse corneas by modulating the Il-17 pathway and neuropeptide Y
Molecular Medicine (2024)
-
Link between organic nanovescicles from vegetable kingdom and human cell physiology: intracellular calcium signalling
Journal of Nanobiotechnology (2024)
-
vissE: a versatile tool to identify and visualise higher-order molecular phenotypes from functional enrichment analysis
BMC Bioinformatics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.