Abstract
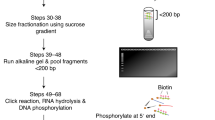
A better understanding of DNA replication initiation in human cells and how this process is altered upon DNA replication stress requires the ability to study origin firing genome wide. Previously described methods of mapping DNA replication origins in higher eukaryotes rely principally on fractionation of DNA fragments based on their size and, optionally, on the presence of ribonucleotides at their 5ʹ end. Here, we describe a protocol for EdUseq-HU, a method for mapping early S-phase replication origins. Cells, synchronized by mitotic shake-off, are released in medium containing 5-ethynyl-2′-deoxyuridine (EdU; to label nascent DNA) and hydroxyurea (HU; to limit fork progression after origin firing). After using click chemistry to tag the EdU label with a biotin conjugate that is cleavable under mild conditions, the nascent DNA is captured on streptavidin beads. One variant of EdUseq-HU allows mapping of DNA replication origins on the genome at a resolution of 10 kb, and a second variant monitors progression of replication forks. Using EdUseq-HU, the spatiotemporal program of DNA replication in human cell lines can be interrogated in <2 weeks. The protocol requires basic cell culture and molecular biology skills, as well as familiarity with the Perl programming language and the Linux operating system.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Fragkos, M., Ganier, O., Coulombe, P. & Méchali, M. DNA replication origin activation in space and time. Nat. Rev. Mol. Cell Biol. 16, 360–374 (2015).
Macheret, M. & Halazonetis, T. D. Intragenic origins due to short G1 phases underlie oncogene-induced DNA replication stress. Nature 555, 112–116 (2018).
Woodward, A. M. et al. Excess Mcm2-7 license dormant origins of replication that can be used under conditions of replicative stress. J. Cell Biol. 173, 673–683 (2006).
Ge, X. Q., Jackson, D. A. & Blow, J. J. Dormant origins licensed by excess Mcm2-7 are required for human cells to survive replicative stress. Genes Dev. 21, 3331–3341 (2007).
Anglana, M., Apiou, F., Bensimon, A. & Debatisse, M. Dynamics of DNA replication in mammalian somatic cells: nucleotide pool modulates origin choice and interorigin spacing. Cell 114, 385–394 (2003).
Rhind, N. & Gilbert, D. M. DNA replication timing. Cold Spring Harb. Perspect. Biol. 5, a010132 (2013).
Halazonetis, T. D., Gorgoulis, V. G. & Bartek, J. An oncogene-induced DNA damage model for cancer development. Science 319, 1352–1355 (2008).
Macheret, M. & Halazonetis, T. D. DNA replication stress as a hallmark of cancer. Annu. Rev. Pathol. 10, 425–448 (2015).
Zeman, M. K. & Cimprich, K. A. Causes and consequences of replication stress. Nat. Cell Biol. 16, 2–9 (2014).
Katou, Y. et al. S-phase checkpoint proteins Tof1 and Mrc1 form a stable replication-pausing complex. Nature 424, 1078–1083 (2003).
MacAlpine, D. M. Coordination of replication and transcription along a Drosophila chromosome. Genes Dev. 18, 3094–3105 (2004).
Karnani, N. & Dutta, A. The effect of the intra-S-phase checkpoint on origins of replication in human cells. Genes Dev. 25, 621–633 (2011).
Fu, H. et al. Mapping replication origin sequences in eukaryotic chromosomes. Curr. Protoc. Cell Biol. 65, 22.20.1–22.20.17 (2014). (2014).
De Brabander, M. J., Van de Veire, R. M., Aerts, F. E., Borgers, M. & Janssen, P. A. The effects of methyl (5-(2-thienyl carbonyl)-1H-benzimidazol-2-yl) carbamate, (R 17934; NSC 238159), a new synthetic antitumoral drug interfering with microtubules, on mammalian cells cultured in vitro. Cancer Res. 36, 905–916 (1976).
Meldal, M. & Tornøe, C. W. Cu-catalyzed azide-alkyne cycloaddition. Chem. Rev. 108, 2952–3015 (2008).
Salic, A. & Mitchison, T. J. A chemical method for fast and sensitive detection of DNA synthesis in vivo. Proc. Natl. Acad. Sci. USA 105, 2415–2420 (2008).
Koç, A., Wheeler, L. J., Mathews, C. K. & Merrill, G. F. Hydroxyurea arrests DNA replication by a mechanism that preserves basal dNTP pools. J. Biol. Chem. 279, 223–230 (2004).
Dellino, G. I. et al. Genome-wide mapping of human DNA-replication origins: Levels of transcription at ORC1 sites regulate origin selection and replication timing. Genome Res. 23, 1–11 (2013).
Mesner, L. D., Crawford, E. L. & Hamlin, J. L. Isolating apparently pure libraries of replication origins from complex genomes. Mol. Cell 21, 719–726 (2006).
Martin, M. M. et al. Genome-wide depletion of replication initiation events in highly transcribed regions. Genome Res. 21, 1822–1832 (2011).
Besnard, E. et al. Unraveling cell type-specific and reprogrammable human replication origin signatures associated with G-quadruplex consensus motifs. Nat. Struct. Mol. Biol. 19, 837–844 (2012).
Petryk, N. et al. Replication landscape of the human genome. Nat. Commun. 7, 10208 (2016).
Urban, J. M., Foulk, M. S., Casella, C. & Gerbi, S. A. The hunt for origins of DNA replication in multicellular eukaryotes. F1000Prime Rep. 7, 30 (2015).
Vashee, S. et al. Sequence-independent DNA binding and replication initiation by the human origin recognition complex. Genes Dev. 17, 1894–1908 (2003).
Foulk, M. S., Urban, J. M., Casella, C. & Gerbi, S. A. Characterizing and controlling intrinsic biases of lambda exonuclease in nascent strand sequencing reveals phasing between nucleosomes and G-quadruplex motifs around a subset of human replication origins. Genome Res. 25, 725–735 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Acknowledgements
We thank present and past laboratory members for helpful discussions, N. Roggli for help with the graphics scripts and M. Docquier and the Genomics Platform of the University of Geneva for scientific input and for performing the high-throughput sequencing. This work was supported by grants from the European Commission (ONIDDAC) and the Swiss Science National Foundation.
Author information
Authors and Affiliations
Contributions
T.D.H. and M.M. designed the experiments and wrote the paper; M.M. performed the experiments; T.D.H. wrote the computer scripts with contributions from M.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related link
Key reference using this protocol
Macheret, M. & Halazonetis, T. D. Nature 555, 112–116 (2018): https://doi.org/10.1038/nature25507
Integrated supplementary information
Supplementary Figure 1 Gating strategy.
Typical flow cytometry profiles of U2OS cells (with normal levels of cyclin E) that were released into medium containing EdU (25 µM) for 14 h after mitosis exit. The strategy used to analyse the flow cytometry profiles consists of first gating the cells based on their distribution in the forward scatter/ side scatter plot (gate A, left panel); then, a second gate is applied to eliminate cell doublets on the propidium iodide (PI) peak height versus integral plot (gate B, center-left panel). After gating, the DNA content histograms and the EdU versus DNA content density scatter plots of the cells are plotted.
Supplementary information
Rights and permissions
About this article
Cite this article
Macheret, M., Halazonetis, T.D. Monitoring early S-phase origin firing and replication fork movement by sequencing nascent DNA from synchronized cells. Nat Protoc 14, 51–67 (2019). https://doi.org/10.1038/s41596-018-0081-y
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-018-0081-y
This article is cited by
-
RECQL4 is not critical for firing of human DNA replication origins
Scientific Reports (2024)
-
Monitoring genome-wide replication fork directionality by Okazaki fragment sequencing in mammalian cells
Nature Protocols (2021)
-
A transcription-based mechanism for oncogenic β-catenin-induced lethality in BRCA1/2-deficient cells
Nature Communications (2021)
-
DNA copy-number measurement of genome replication dynamics by high-throughput sequencing: the sort-seq, sync-seq and MFA-seq family
Nature Protocols (2020)
-
Sonic hedgehog accelerates DNA replication to cause replication stress promoting cancer initiation in medulloblastoma
Nature Cancer (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.