Abstract
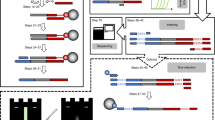
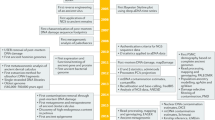
DNA preserved in ancient bones, teeth and sediments is typically highly fragmented and present only in minute amounts. Here, we provide a highly versatile silica-based DNA extraction protocol that enables the retrieval of short (≥35 bp) or even ultrashort (≥25 bp) DNA fragments from such material with minimal carryover of substances that inhibit library preparation for high-throughput sequencing. DNA extraction can be performed with either silica spin columns, which offer the most convenient choice for manual DNA extraction, or silica-coated magnetic particles. The latter allow a substantial cost reduction as well as automation on liquid-handling systems. This protocol update replaces a now-outdated version that was published 11 years ago, before high-throughput sequencing technologies became widely available. It has been thoroughly optimized to provide the highest DNA yields from highly degraded samples, as well as fast and easy handling, requiring not more than ~15 min of hands-on time per sample.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Meyer, M. et al. A mitochondrial genome sequence of a hominin from Sima de los Huesos. Nature 505, 403–406 (2014).
Orlando, L. et al. Recalibrating Equus evolution using the genome sequence of an early Middle Pleistocene horse. Nature 499, 74–78 (2013).
Willerslev, E. et al. Diverse plant and animal genetic records from Holocene and Pleistocene sediments. Science 300, 791–795 (2003).
Slon, V. et al. Neandertal and Denisovan DNA from Pleistocene sediments. Science 356, 605–608 (2017).
Höss, M. & Pääbo, S. DNA extraction from Pleistocene bones by a silica-based purification method. Nucleic Acids Res. 21, 3913–3914 (1993).
Rohland, N. & Hofreiter, M. Ancient DNA extraction from bones and teeth. Nat. Protoc. 2, 1756–1762 (2007).
Yang, D. Y., Eng, B., Waye, J. S., Dudar, J. C. & Saunders, S. R. Technical note: improved DNA extraction from ancient bones using silica-based spin columns. Am. J. Phys. Anthropol. 105, 539–543 (1998).
Pääbo, S. et al. Genetic analyses from ancient DNA. Annu. Rev. Genet. 38, 645–679 (2004).
Dabney, J. et al. Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proc. Natl. Acad. Sci. USA 110, 15758–15763 (2013).
Adler, C. J., Haak, W., Donlon, D. & Cooper, A. Survival and recovery of DNA from ancient teeth and bones. J. Archaeol. Sci. 38, 956–964 (2011).
Allentoft, M. E. et al. The half-life of DNA in bone: measuring decay kinetics in 158 dated fossils. Proc. Biol. Sci. 279, 4724–4733 (2012).
Handt, O., Höss, M., Krings, M. & Pääbo, S. Ancient DNA: methodological challenges. Experientia 50, 524–529 (1994).
Schwarz, C. et al. New insights from old bones: DNA preservation and degradation in permafrost preserved mammoth remains. Nucleic Acids Res. 37, 3215–3229 (2009).
Gansauge, M. T. & Meyer, M. Single-stranded DNA library preparation for the sequencing of ancient or damaged DNA. Nat. Protoc. 8, 737–748 (2013).
Meyer, M. et al. A high-coverage genome sequence from an Archaic Denisovan individual. Science 338, 222–226 (2012).
Meyer, M. et al. Nuclear DNA sequences from the Middle Pleistocene Sima de los Huesos hominins. Nature 531, 504–507 (2016).
Fu, Q. et al. The genetic history of Ice Age Europe. Nature 534, 200–205 (2016).
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015).
Lipson, M. et al. Parallel palaeogenomic transects reveal complex genetic history of early European farmers. Nature 551, 368–372 (2017).
Mathieson, I. et al. Genome-wide patterns of selection in 230 ancient Eurasians. Nature 528, 499–503 (2015).
Glocke, I. & Meyer, M. Extending the spectrum of DNA sequences retrieved from ancient bones and teeth. Genome Res. 27, 1230–1237 (2017).
Lipson, M. et al. Ancient genomes document multiple waves of migration in Southeast Asian prehistory. Science https://doi.org/10.1126/science.aat3188 (2018).
Korlevic, P. et al. Reducing microbial and human contamination in DNA extractions from ancient bones and teeth. Biotechniques 59, 87–93 (2015).
Gamba, C. et al. Comparing the performance of three ancient DNA extraction methods for high-throughput sequencing. Mol. Ecol. Resour. 16, 459–469 (2016).
Allentoft, M. E. et al. Population genomics of Bronze Age Eurasia. Nature 522, 167–172 (2015).
Gutaker, R. M., Reiter, E., Furtwangler, A., Schuenemann, V. J. & Burbano, H. A. Extraction of ultrashort DNA molecules from herbarium specimens. Biotechniques 62, 76–79 (2017).
de Filippo, C., Meyer, M. & Pruefer, K. Harvesting information from ultra-short ancient DNA sequences. Preprint at https://www.biorxiv.org/content/early/2018/05/10/319277 (2018).
Fu, Q. M. et al. DNA analysis of an early modern human from Tianyuan Cave, China. Proc. Natl. Acad. Sci. USA 110, 2223–2227 (2013).
Gansauge, M. T. et al. Single-stranded DNA library preparation from highly degraded DNA using T4 DNA ligase. Nucleic Acids Res. 45, e79 (2017).
Bennett, E. A. et al. Library construction for ancient genomics: single strand or double strand? Biotechniques 56, 289–300 (2014).
Wales, N. et al. New insights on single-stranded versus double-stranded DNA library preparation for ancient DNA. Biotechniques 59, 368–371 (2015).
Briggs, A. W. & Heyn, P. Preparation of next-generation sequencing libraries. in Methods in Molecular Biology: Ancient DNA, Methods and Protocols, Vol. 840 (eds. Shapiro, B. & Hofreiter, M.) 143–154 (Humana Press, Totowa, NJ, 2012).
Meyer, M. & Kircher, M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb. Protoc. 2010, pdb.prot5448 (2010).
Rohland, N., Harney, E., Mallick, S., Nordenfelt, S. & Reich, D. Partial uracil-DNA-glycosylase treatment for screening of ancient DNA. Philos. Trans. R. Soc. Lond. B Biol. Sci. 370, 20130624 (2015).
Knapp, M., Clarke, A. C., Horsburgh, K. A. & Matisoo-Smith, E. A. Setting the stage—building and working in an ancient DNA laboratory. Ann. Anat. 194, 3–6 (2012).
Kircher, M., Sawyer, S. & Meyer, M. Double indexing overcomes inaccuracies in multiplex sequencing on the Illumina platform. Nucleic Acids Res. 40, e3 (2012).
Acknowledgements
We thank S. Pääbo and D. Reich for their support; A. Weihmann and B. Schellbach for performing the sequencing runs; J. Kelso and J. Visagie for help with raw data processing; N. Broomand, M. Ferry, M. Michel, J. Oppenheimer and K. Stewardson for support in the lab; and S. Mallick for bioinformatics processing of initial experiments. We also thank P. Rudan, C. Verna, T. Kutznetsova, K. Post, G. Rabeder, M. Shunkov, R. Roberts, A. Derevianko, R. Miller, J. Stewart and M. Soressi for providing the samples. This work was funded by the Strategic Innovation Fund of the Max Planck Society and ERC grant agreement no. 694707 to S. Pääbo.
Author information
Authors and Affiliations
Contributions
N.R., I.G., A.A.-P. and M.M. designed experiments. N.R., I.G. and A.A.-P. performed experiments. N.R., I.G. and M.M. analyzed the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
1. Slon, V. et al. Science 356, 605–608 (2017): https://doi.org/10.1126/science.aam9695
2. Meyer, M. et al. Nature 505, 403–406 (2014): https://doi.org/10.1038/nature12788
3. Olalde, I. et al. Nature 555, 190–196 (2018): https://doi.org/10.1038/nature25738
This protocol is an update to: Nat. Protoc. 2, 1756–1762 (2007): https://doi.org/10.1038/nprot.2007.247
Supplementary information
Supplementary Table 1
Summary of sequencing results
Rights and permissions
About this article
Cite this article
Rohland, N., Glocke, I., Aximu-Petri, A. et al. Extraction of highly degraded DNA from ancient bones, teeth and sediments for high-throughput sequencing. Nat Protoc 13, 2447–2461 (2018). https://doi.org/10.1038/s41596-018-0050-5
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-018-0050-5
This article is cited by
-
Homo sapiens reached the higher latitudes of Europe by 45,000 years ago
Nature (2024)
-
Medieval DNA from Soqotra points to Eurasian origins of an isolated population at the crossroads of Africa and Arabia
Nature Ecology & Evolution (2024)
-
Detection of chromosomal aneuploidy in ancient genomes
Communications Biology (2024)
-
The ecology, subsistence and diet of ~45,000-year-old Homo sapiens at Ilsenhöhle in Ranis, Germany
Nature Ecology & Evolution (2024)
-
On Roth’s “human fossil” from Baradero, Buenos Aires Province, Argentina: morphological and genetic analysis
Swiss Journal of Palaeontology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.