Abstract
Hydroxyl-radical footprinting (HRF) is a powerful method for probing structures of nucleic acid–protein complexes with single-nucleotide resolution in solution. To tap the full quantitative potential of HRF, we describe a protocol, hydroxyl-radical footprinting interpretation for DNA (HYDROID), to quantify HRF data and integrate them with atomistic structural models. The stages of the HYDROID protocol are extraction of the lane profiles from gel images, quantification of the DNA cleavage frequency at each nucleotide and theoretical estimation of the DNA cleavage frequency from atomistic structural models, followed by comparison of experimental and theoretical results. Example scripts for each step of HRF data analysis and interpretation are provided for several nucleosome systems; they can be easily adapted to analyze user data. As input, HYDROID requires polyacrylamide gel electrophoresis (PAGE) images of HRF products and optionally can use a molecular model of the DNA–protein complex. The HYDROID protocol can be used to quantify HRF over DNA regions of up to 100 nucleotides per gel image. In addition, it can be applied to the analysis of RNA–protein complexes and free RNA or DNA molecules in solution. Compared with other methods reported to date, HYDROID is unique in its ability to simultaneously integrate HRF data with the analysis of atomistic structural models. HYDROID is freely available. The complete protocol takes ~3 h. Users should be familiar with the command-line interface, the Python scripting language and Protein Data Bank (PDB) file formats. A graphical user interface (GUI) with basic functionality (HYDROID_GUI) is also available.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data used in this protocol are available at the HYDROID repository at https://github.com/ncbi/HYDROID.
References
Tullius, T. D. Physical studies of protein-DNA complexes by footprinting. Annu. Rev. Biophys. Biophys. Chem. 18, 213–237 (1989).
Adilakshmi, T., Lease, R. A. & Woodson, S. A. Hydroxyl radical footprinting in vivo: mapping macromolecular structures with synchrotron radiation. Nucleic Acids Res. 34, e64 (2006).
Song, L. & Crawford, G. E. DNase-seq: a high-resolution technique for mapping active gene regulatory elements across the genome from mammalian cells. Cold Spring Harb. Protoc. https://doi.org/10.1101/pdb.prot5384 (2010).
Koohy, H., Down, T. A. & Hubbard, T. J. Chromatin accessibility data sets show bias due to sequence specificity of the DNase I enzyme. PLoS ONE 8, e69853 (2013).
Jain, S. S. & Tullius, T. D. Footprinting protein-DNA complexes using the hydroxyl radical. Nat. Protoc. 3, 1092–1100 (2008).
Armeev, G. A., Gorkovets, T. K., Efimova, D. A., Shaitan, K. V. & Shaytan, A. K. Modeling of the structure of protein–DNA complexes using the data from FRET and footprinting experiments. Moscow Univ. Biol. Sci. Bull. 71, 29–33 (2016).
Balasubramanian, B., Pogozelski, W. K. & Tullius, T. D. DNA strand breaking by the hydroxyl radical is governed by the accessible surface areas of the hydrogen atoms of the DNA backbone. Proc. Natl. Acad. Sci. USA 95, 9738–9743 (1998).
Shadle, S. E. et al. Quantitative analysis of electrophoresis data: novel curve fitting methodology and its application to the determination of a protein-DNA binding constant. Nucleic Acids Res. 25, 850–860 (1997).
Das, R., Laederach, A., Pearlman, S. M., Herschlag, D. & Altman, R. B. SAFA: semi-automated footprinting analysis software for high-throughput quantification of nucleic acid footprinting experiments. RNA 11, 344–354 (2005).
Bishop, E. P. et al. A map of minor groove shape and electrostatic potential from hydroxyl radical cleavage patterns of DNA. ACS Chem. Biol. 6, 1314–1320 (2011).
Shaytan, A. K. et al. Hydroxyl-radical footprinting combined with molecular modeling identifies unique features of DNA conformation and nucleosome positioning. Nucleic Acids Res. 45, 9229–9243 (2017).
Xiao, H. et al. Molecular basis of CENP-C association with the CENP-A nucleosome at yeast centromeres. Genes Dev. 31, 1958–1972 (2017).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Mitternacht, S. FreeSASA: an open source C library for solvent accessible surface area calculations. F1000Res. 5, 189 (2016).
Ingle, S., Azad, R. N., Jain, S. S. & Tullius, T. D. Chemical probing of RNA with the hydroxyl radical at single-atom resolution. Nucleic Acids Res. 42, 12758–12767 (2014).
Costa, M. & Monachello, D. Probing RNA folding by hydroxyl radical footprinting. Methods Mol. Biol. 1086, 119–142 (2014).
Hampel, K. J. & Burke, J. M. Time-resolved hydroxyl-radical footprinting of RNA using Fe(II)-EDTA. Methods 23, 233–239 (2001).
Nilsen, T. W. Mapping RNA-protein interactions using hydroxyl-radical footprinting. Cold Spring Harb. Protoc. 2014, 1333–1336 (2014).
Ding, F., Lavender, C. A., Weeks, K. M. & Dokholyan, N. V. Three-dimensional RNA structure refinement by hydroxyl radical probing. Nat. Methods 9, 603–608 (2012).
Tian, S. & Das, R. RNA structure through multidimensional chemical mapping. Q. Rev. Biophys. 49, e7 (2016).
Begusova, M., Spotheim-Maurizot, M., Sy, D., Michalik, V. & Charlier, M. RADACK, a stochastic simulation of hydroxyl radical attack to DNA. J. Biomol. Struct. Dyn. 19, 141–158 (2001).
Linkert, M. et al. Metadata matters: access to image data in the real world. J. Cell Biol. 189, 777–782 (2010).
Takamoto, K., Chance, M. R. & Brenowitz, M. Semi-automated, single-band peak-fitting analysis of hydroxyl radical nucleic acid footprint autoradiograms for the quantitative analysis of transitions. Nucleic Acids Res. 32, E119 (2004).
Word, J. M., Lovell, S. C., Richardson, J. S. & Richardson, D. C. Asparagine and glutamine: using hydrogen atom contacts in the choice of side-chain amide orientation. J. Mol. Biol. 285, 1735–1747 (1999).
Case, D. A. et al. AMBER 2017 (University of California, San Francisco, San Francisco, CA, 2017).
Lee, B. & Richards, F. M. The interpretation of protein structures: estimation of static accessibility. J. Mol. Biol. 55, 379–400 (1971).
Best, R. B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone phi, psi and side-chain chi(1) and chi(2) dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Slater, G. W., Desruisseaux, C. & Hubert, S. J. DNA separation mechanisms during electrophoresis. in Capillary Electrophoresis of Nucleic Acids: Volume I: Introduction to the Capillary Electrophoresis of Nucleic Acids (eds. Keith, R. M. & Jing, C.) 27–41 (Humana Press, Totowa, NJ, 2001).
Maxam, A. M. & Gilbert, W. A new method for sequencing DNA. Proc. Natl. Acad. Sci. USA 74, 560–564 (1977).
Brahmasandra, S. N., Burke, D. T., Mastrangelo, C. H. & Burns, M. A. Mobility, diffusion and dispersion of single-stranded DNA in sequencing gels. Electrophoresis 22, 1046–1062 (2001).
Maramis, C. F. & Delopoulos, A. N. Improved modeling of lane intensity profiles on gel electrophoresis images In XII Mediterranean Conference on Medical and Biological Engineering and Computing 2010: May 27–30, 2010 Chalkidiki, Greece (eds. Panagiotis, D. B. & Nicolas, P.) 671–674 (Springer, Berlin, 2010).
Greenbaum, J. A., Pang, B. & Tullius, T. D. Construction of a genome-scale structural map at single-nucleotide resolution. Genome Res. 17, 947–953 (2007).
Berg, J. M. APPENDIX: Lorentzian lineshapes are intrinsic to autoradiographic detection. Nucleic Acids Res. 25, 850–860 (1997).
Spotheim-Maurizot, M. & Davidkova, M. Radiation damage toDNA in DNA-protein complexes. Mutat. Res. 711, 41–48 (2011).
Morozov, A. V. et al. Using DNA mechanics to predict in vitro nucleosome positions and formation energies. Nucleic Acids Res. 37, 4707–4722 (2009).
Cloutier, T. E. & Widom, J. DNA twisting flexibility and the formation of sharply looped protein-DNA complexes. Proc. Natl. Acad. Sci. USA 102, 3645–3650 (2005).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Lu, X. J. & Olson, W. K. 3DNA: a versatile, integrated software system for the analysis, rebuilding and visualization of three-dimensional nucleic-acid structures. Nat. Protoc. 3, 1213–1227 (2008).
Draizen, E. J. et al. HistoneDB 2.0: a histone database with variants—an integrated resource to explore histones and their variants. Database 2016, baw014 (2016).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Lowary, P. T. & Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 276, 19–42 (1998).
Rychkov, G. N. et al. Partially assembled nucleosome structures at atomic detail. Biophys. J. 112, 460–472 (2017).
Eslami-Mossallam, B., Schiessel, H. & van Noort, J. Nucleosome dynamics: sequence matters. Adv. Colloid Interface Sci. 232, 101–113 (2016).
Kappen, L. S. & Goldberg, I. H. Deoxyribonucleic acid damage by neocarzinostatin chromophore: strand breaks generated by selective oxidation of C-5ʹ of deoxyribose. Biochemistry 22, 4872–4878 (1983).
Acknowledgements
This work was supported by the Intramural Research Programs of the National Library of Medicine (A.K.S., D.L. and A.R.P.) and the National Cancer Institute (H.X.), National Institutes of Health; the Russian Science Foundation (grant no. 14-24-00031; agreement no. 14-24-00031-p; development of gel simulation algorithms and HYDROID_GUI package; G.A.A. and A.K.S.); the Howard Hughes Medical Institute Janelia Research Campus and a Johns Hopkins University Bloomberg Distinguished Professorship (C.W.); the US–Russia Collaboration in the Biomedical Sciences, National Institutes of Health Visiting Fellows program (A.K.S.); NIH grants R01GM119398 and R21CA220151 (V.M.S.); and NIH grants R21DE025398 and P50DE019032 (D.A.G.). This work utilized the computational resources of the NIH HPC Biowulf cluster (http://hpc.nih.gov) and the shared research facilities of HPC computing resources at Lomonosov Moscow State University. We thank T. Tullius for valuable suggestions that helped to formulate the initial idea of this study, L. Zaslavsky for discussions on optimal data-fitting algorithms, and S. Mitternacht for modifications to the FreeSASA program.
Author information
Authors and Affiliations
Contributions
A.K.S. designed and developed the HYDROID package. A.R.P. supervised the project. A.K.S and A.R.P. tested the protocol and wrote the paper. H.X. performed experimental studies on 3′-labeled nucleosomes. D.A.G. performed experimental studies of 5′-labeled nucleosomes. G.A.A. developed gel simulation algorithms and the HYDROID_GUI package and also contributed to software packaging. G.A.K. tested the software and contributed to video tutorials. D.L., V.M.S. and C.W. contributed to the discussions and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
1. Shaytan, A. K. et al. Nucleic Acids Res. 45, 9229–9243 (2017): https://doi.org/10.1093/nar/gkx616
2. Xiao, H. et al. Genes Dev. 31, 1958–1972 (2017): http://www.genesdev.org/cgi/doi/10.1101/gad.304782.117
Integrated supplementary information
Supplementary Figure 1 Comparison of various fitting methods and band-shape models applied to the analysis of an HRF gel lane profile.
Results of HYDROIDexp Stage 4 are presented for different combinations of band shape models (Gaussian and Lorentzian) and constrained fitting methods. Data are taken from an HRF experiment on S. cerevisiae centromeric nucleosome reconstituted on 601TA sequence (Example 1). Application of constrained fitting methods dramatically improves the quality of the fitted solution. A combination of Gaussian band shapes and “dSIGMA>= 0” constraint options provides the best fitting quality while presenting a physically plausible solution. The x-axis corresponds to the distance along the gel lane in pixels. The y-axis shows the intensity of the pixels along the gel lane profile.
Supplementary Figure 2 Comparison of extracted band intensities from Supplementary Fig. 1.
Solutions with no constraint options employed during fitting manifest unphysical fluctuations due to overfitting (nearby bands should have similar widths) in the region where the bands are located close together. The use of Lorentzians versus Gaussians provides slightly altered results in the low and high regions of the cleavage frequency values. Zero on the x-axis marks the nucleotide positioned on the central dyad symmetry axis of nucleosome.
Supplementary Figure 3 Example of HRF profile deconvolution for a profile with a certain level of noise and interpolated band locations.
Our data fitting method works well even if individual bands lack discernible local maxima at their respective positions as long as their approximate positions are interpolated from the spacing of the neighboring bands. Data from the HRF gel lane profile for nucleosome of G. gallus reconstituted on 601 sequence was used (Example 2).
Supplementary Figure 4 Reproducibility of the HRF quantification results from independently repeated experiments.
Left: PAGE gel of HRF experiments for G. gallus nucleosome reconstituted on a 601 sequence. Experimental details can be found in ref. 36. Briefly, histones were extracted from G. gallus erythrocytes and nucleosomes were reconstitutes on a 601 Widom positioning sequence with a radioactive label on the 5ʹ end. After hydroxyl-radical footprinting of nucleosomes, DNA was extracted and subjected to denaturing PAGE followed by radioactive signal acquisition using phosphorimaging. The lanes on the gel correspond to the following reaction products: TS - experiments for the top DNA strand, BS – experiments for the bottom DNA strand. N represents HRF reaction products for the nucleosomal DNA, and A, G and T are the sequencing lanes and mark the products of primer extension using Taq DNA polymerase and terminating dideoxynucleotide triphosphates (ddATP (A), ddTTP (T), and ddGTP (G), respectively). Experiment A and Experiment B labels mark the lanes resulting from two independent HRF experiments performed on the same sample of nucleosomes. The blue lines indicate the area of the gel used to extract gel lane profiles. Right: Comparison of profiles extracted by HYDROIDexp from independently repeated HRF experiments. The y-axis represents the normalized values of DNA cleavage frequency. When plotting two profiles on one plot the “every” method from HYDORID package was used, which simply normalizes every profile to its maximum value. Zero on the x-axis marks the nucleotide positioned on the central dyad symmetry axis of nucleosome.
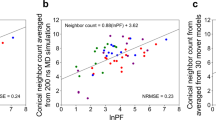
Supplementary Figure 5 Comparison between theoretically estimated DNA cleavage profiles generated from X-ray-derived structure and MD simulations.
Theoretically estimated DNA cleavage profiles (H-SASA profiles) for S. cerevisiae centromeric nucleosome reconstituted on 601TA sequence based on all deoxyribose hydrogen atoms were obtained for two cases: from an X-ray-derived homology model (DNA geometry strictly corresponds to the X-ray structure) and averaged over frames of MD simulation. Differences are attributed to the relaxation of small DNA structure irregularities and averaging over DNA backbone dynamics.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5
Rights and permissions
About this article
Cite this article
Shaytan, A.K., Xiao, H., Armeev, G.A. et al. Structural interpretation of DNA–protein hydroxyl-radical footprinting experiments with high resolution using HYDROID. Nat Protoc 13, 2535–2556 (2018). https://doi.org/10.1038/s41596-018-0048-z
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-018-0048-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.