Abstract
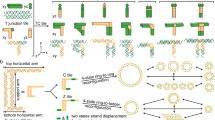
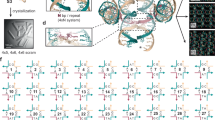
Information relay and cascaded transformation are essential in biology and engineering. Imitation of such complex behaviors via synthetic molecular self-assembly at the nanoscale remains challenging. Here we describe the use of structural DNA nanotechnology to realize prescribed, multistep, long-range information relay and cascaded transformation in rationally designed molecular arrays. The engineered arrays provide a controlled platform for studying complex dynamic behaviors of molecular arrays and have a range of potential applications, such as with reconfigurable metamaterials. A reconfigurable array consists of a prescribed number of interconnected dynamic DNA antijunctions. Each antijunction unit consists of four DNA domains of equal length with four dynamic nicking points, which are capable of switching between two stable conformations through an intermediate open conformation. By interconnecting the small DNA antijunctions, one can build custom two-dimensional (2D) molecular ‘domino’ arrays with arbitrary shapes. More important, the DNA molecular arrays are capable of undergoing programmed, multistep, long-range transformation driven by information relay between neighboring antijunction units. The information relay is initiated by the trigger strands under high temperature or formamide concentration. The array’s dynamic behavior can be regulated by external factors such as its shape and size, points of transformation initiation, and/or any engineered information propagation pathways. This protocol provides detailed strategies for designing DNA molecular arrays, as well as procedures for sample production, purification, reconfiguration, and imaging by atomic force microscopy (AFM) and transmission electron microscopy (TEM). The procedure can be completed in 4–7 d.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Park, S. H. et al. Three-helix bundle DNA tiles self-assemble into 2D lattice or 1D templates for silver nanowires. Nano Lett. 5, 693–696 (2005).
Schulman, R. & Winfree, E. Synthesis of crystals with a programmable kinetic barrier to nucleation. Proc. Natl. Acad. Sci. USA 104, 15236–15241 (2007).
Aldaye, F. A., Palmer, A. L. & Sleiman, H. F. Assembling materials with DNA as the guide. Science 321, 1795–1799 (2008).
Yan, H., Park, S. H., Finkelstein, G., Reif, J. H. & LaBean, T. H. DNA-templated self-assembly of protein arrays and highly conductive nanowires. Science 301, 1882–1884 (2003).
Liu, D., Park, S. H., Reif, J. H. & LaBean, T. H. DNA nanotubes self-assembled from triple-crossover tiles as templates for conductive nanowires. Proc. Natl. Acad. Sci. USA 101, 717–722 (2004).
Rothemund, P. W. K. et al. Design and characterization of programmable DNA nanotubes. J. Am. Chem. Soc. 126, 16344–16352 (2004).
Mathieu, F. et al. Six-helix bundles designed from DNA. Nano Lett. 5, 661–665 (2005).
Sharma, J. et al. Control of self-assembly of DNA tubules through integration of gold nanoparticles. Science 323, 112–116 (2009).
Ke, Y. et al. DNA brick crystals with prescribed depths. Nat. Chem. 6, 994–1002 (2014).
Winfree, E., Liu, F. R., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Rothemund, P. W. K., Papadakis, N. & Winfree, E. Algorithmic self-assembly of DNA Sierpinski triangles. PLoS Biol. 2, 2041–2053 (2004).
He, Y. et al. Sequence symmetry as a tool for designing DNA nanostructures. Angew. Chem. Int. Ed. 117, 6852–6854 (2005).
Gerling, T., Wagenbauer, K. F., Neuner, A. M. & Dietz, H. Dynamic DNA devices and assemblies formed by shape-complementary, non-base pairing 3D components. Science 347, 1446–1452 (2015).
Wang, P. et al. Programming self-assembly of DNA origami honeycomb two-dimensional lattices and plasmonic metamaterials. J. Am. Chem. Soc. 138, 7733–7740 (2016).
Liu, W., Halverson, J., Tian, Y., Tkachenko, A. V. & Gang, O. Self-organized architectures from assorted DNA-framed nanoparticles. Nat. Chem. 8, 867–873 (2016).
Wei, B., Dai, M. & Yin, P. Complex shapes self-assembled from single-stranded DNA tiles. Nature 485, 623–627 (2012).
Chen, J. H. & Seeman, N. C. Synthesis from DNA of a molecule with the connectivity of a cube. Nature 350, 631–633 (1991).
Shih, W. M., Quispe, J. D. & Joyce, G. F. A 1.7-kilobase single-stranded DNA that folds into a nanoscale octahedron. Nature 427, 618–621 (2004).
Goodman, R. P. et al. Rapid chiral assembly of rigid DNA building blocks for molecular nanofabrication. Science 310, 1661–1665 (2005).
He, Y. et al. Hierarchical self-assembly of DNA into symmetric supramolecular polyhedra. Nature 452, 198–201 (2008).
Andersen, E. S. et al. Self-assembly of a nanoscale DNA box with a controllable lid. Nature 459, 73–76 (2009).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Dietz, H., Douglas, S. M. & Shih, W. M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725–730 (2009).
Han, D. et al. DNA origami with complex curvatures in three-dimensional space. Science 332, 342–346 (2011).
Ke, Y., Ong, L. L., Shih, W. M. & Yin, P. Three-dimensional structures self-assembled from DNA bricks. Science 338, 1177–1183 (2012).
Iinuma, R. et al. Polyhedra self-assembled from DNA tripods and characterized with 3D DNA-PAINT. Science 344, 65–69 (2014).
Benson, E. et al. DNA rendering of polyhedral meshes at the nanoscale. Nature 523, 441–444 (2015).
Veneziano, R. et al. DNA nanotechnology designer nanoscale DNA assemblies programmed from the top down. Science 352, aaf4388 (2016).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Ke, Y., Castro, C. & Choi, J. H. Structural DNA nanotechnology: artificial nanostructures for biomedical research. Annu. Rev. Biomed. Eng. 20, 375–401 (2018).
Wang, P. et al. Practical aspects of structural and dynamic DNA nanotechnology. MRS Bull. 42, 889–896 (2017).
Castro, C. E. et al. A primer to scaffolded DNA origami. Nat. Methods 8, 221 (2011).
Wang, P., Meyer, T. A., Pan, V., Dutta, P. K. & Ke, Y. The beauty and utility of DNA origami. Chem 2, 359–382 (2017).
Yurke, B., Turberfield, A. J., Mills, A. P., Simmel, F. C. & Neumann, J. L. A DNA-fuelled molecular machine made of DNA. Nature 406, 605–608 (2000).
Yan, H., Zhang, X. P., Shen, Z. Y. & Seeman, N. C. A robust DNA mechanical device controlled by hybridization topology. Nature 415, 62–65 (2002).
Liu, M. et al. A DNA tweezer-actuated enzyme nanoreactor. Nat. Commun. 4, 2127 (2013).
Ke, Y. et al. Regulation at a distance of biomolecular interactions using a DNA origami nanoactuator. Nat. Commun. 7, 10935 (2016).
Kuzyk, A. et al. Reconfigurable 3D plasmonic metamolecules. Nat. Mater. 13, 862 (2014).
Kuzyk, A. et al. A light-driven three-dimensional plasmonic nanosystem that translates molecular motion into reversible chiroptical function. Nat. Commun. 7, 10591 (2016).
Sherman, W. B. & Seeman, N. C. A precisely controlled DNA biped walking device. Nano Lett. 4, 1203–1207 (2004).
Yin, P., Yan, H., Daniell, X. G., Turberfield, A. J. & Reif, J. H. A unidirectional DNA walker that moves autonomously along a track. Angew. Chem. Int. Ed. 43, 4906–4911 (2004).
Omabegho, T., Sha, R. & Seeman, N. C. A bipedal DNA brownian motor with coordinated legs. Science 324, 67–71 (2009).
Yin, P. et al. Programming biomolecular self-assembly pathways. Nature 451, 318–322 (2008).
Lund, K. et al. Molecular robots guided by prescriptive landscapes. Nature 465, 206–210 (2010).
Wickham, S. F. J. et al. Direct observation of stepwise movement of a synthetic molecular transporter. Nat. Nanotechnol. 6, 166–169 (2011).
Zhou, C., Duan, X. & Liu, N. A plasmonic nanorod that walks on DNA origami. Nat. Commun. 6, 8102 (2015).
Marras, A. E., Zhou, L., Su, H.-J. & Castro, C. E. Programmable motion of DNA origami mechanisms. Proc. Natl. Acad. Sci. USA 112, 713 (2015).
Endo, M. et al. Helical DNA origami tubular structures with various sizes and arrangements. Angew. Chem, Int. Ed. 53, 7484–7490 (2014).
Song, J. et al. Reconfiguration of DNA molecular arrays driven by information relay. Science 357, eaan3377 (2017).
Du, S. M., Zhang, S. W. & Seeman, N. C. DNA junctions, antijunctions, and mesojunctions. Biochemistry 31, 10955–10963 (1992).
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414 (2009).
Dietz, H., Douglas, S. M. & Shih, W. M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725 (2009).
Thubagere, A. J. et al. A cargo-sorting DNA robot Science 357, eaan6558 (2017).
Choi, Y., Choi, H., Lee, A. C., Lee, H. & Kwon, S. A reconfigurable DNA accordion rack. Angew. Chem. Int. Ed. 57, 2811–2815 (2018).
Sobczak, J.-P. J., Martin, T. G., Gerling, T. & Dietz, H. Rapid folding of DNA into nanoscale shapes at constant temperature. Science 338, 1458–1461 (2012).
Bellot, G., McClintock, M. A., Chou, J. J. & Shih, W. M. DNA nanotubes for NMR structure determination of membrane proteins. Nat. Protoc. 8, 755–770 (2013).
Stahl, E., Martin, T. G., Praetorius, F. & Dietz, H. Facile and scalable preparation of pure and dense DNA origami solutions. Angew. Chem. Int. Ed. 53, 12735–12740 (2014).
Blake, R. D. & Delcourt, S. G. Thermodynamic effects of formamide on DNA stability. Nucleic Acids Res. 24, 2095–2103 (1996).
Fischer, S. G. & Lerman, L. S. DNA fragments differing by single base-pair substitutions are separated in denaturing gradient gels: correspondence with melting theory. Proc. Natl. Acad. Sci. USA 80, 1579–1583 (1983).
McConaughy, B. L., Laird, C. D. & McCarthy, B. J. Nucleic acid reassociation in formamide. Biochemistry 8, 3289–3295 (1969).
Acknowledgements
This work was supported by the NSF (CAREER Award DMR-1654485), the Wallace H. Coulter Department of Biomedical Engineering Startup Fund, a Billi and Bernie Marcus Research Award (to Y.K.), the National Natural Scientific Foundation of China (grants 11761141006 and 21605102 to J.S.), and the National Key Research and Development Program of China (grant 2017FYA0205301 to D.C.).
Author information
Authors and Affiliations
Contributions
J.S. and Y.K. conceived and led the project. J.S., P.W., and Y.K. designed and conducted the experiments. D.W., J.S., P.W., V.P., Y.Z., D.C., and Y.K. contributed to the writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A provisional US patent application based on the work described in this paper has been filed.
Additional information
Related links
Article describing the development of the approach
1. Song, J. et al. Science 357, eaan3377 (2017): https://doi.org/10.1126/science.aan3377
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1
Design diagram of the 11 × 4 52-bp DNA origami array.
Supplementary Figure 2
Design diagram of the 20 × 8 42-bp DNA brick array.
Supplementary information
Supplementary Data 1
Python code for sequence generation
Supplementary Data 2
caDNAno files for the 11 × 4 52-bp DNA origami array and the 20 × 8 42-bp DNA brick array
Rights and permissions
About this article
Cite this article
Wang, D., Song, J., Wang, P. et al. Design and operation of reconfigurable two-dimensional DNA molecular arrays. Nat Protoc 13, 2312–2329 (2018). https://doi.org/10.1038/s41596-018-0039-0
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-018-0039-0
This article is cited by
-
A Triode-like Enzyme-free Catalytic Circuit with Junction Fuel
Chemical Research in Chinese Universities (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.