Abstract
Despite the significance of N6-methyladenosine (m6A) in gene regulation, the requirement for large amounts of RNA has hindered m6A profiling in mammalian early embryos. Here we apply low-input methyl RNA immunoprecipitation and sequencing to map m6A in mouse oocytes and preimplantation embryos. We define the landscape of m6A during the maternal-to-zygotic transition, including stage-specifically expressed transcription factors essential for cell fate determination. Both the maternally inherited transcripts to be degraded post fertilization and the zygotically activated genes during zygotic genome activation are widely marked by m6A. In contrast to m6A-marked zygotic ally-activated genes, m6A-marked maternally inherited transcripts have a higher tendency to be targeted by microRNAs. Moreover, RNAs derived from retrotransposons, such as MTA that is maternally expressed and MERVL that is transcriptionally activated at the two-cell stage, are largely marked by m6A. Our results provide a foundation for future studies exploring the regulatory roles of m6A in mammalian early embryonic development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All picoMeRIP-seq data generated in this study are available in GEO with accession number GSE192440. The differentially expressed genes (Ythdf2 knockout versus wild-type GV oocytes) were obtained from GEO (GSE147849). The expression values of miRNAs in mouse MII oocyte, and one-cell (zygote), two-cell and eight-cell embryos were obtained from the Supplementary Dataset 1 of a previous study22.
Code availability
The scripts of major analysis modules in this study (including quality control, alignment, reads filtering, gene expression quantification, peak calling and motif search) are packaged into a bioinformatics pipeline called MeRipBox. MeRipBox is publicly available at GitHub, at the following address: https://github.com/Augroup/MeRipBox.
References
Zhao, B. S., Roundtree, I. A. & He, C. Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 18, 31–42 (2017).
Zaccara, S., Ries, R. J. & Jaffrey, S. R. Reading, writing and erasing mRNA methylation. Nat. Rev. Mol. Cell Biol. 20, 608–624 (2019).
Klungland, A., Dahl, J. A., Greggains, G., Fedorcsak, P. & Filipczyk, A. Reversible RNA modifications in meiosis and pluripotency. Nat. Methods 14, 18–22 (2016).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Atlasi, Y. & Stunnenberg, H. G. The interplay of epigenetic marks during stem cell differentiation and development. Nat. Rev. Genet. 18, 643–658 (2017).
Dahl, J. A. et al. Broad histone H3K4me3 domains in mouse oocytes modulate maternal-to-zygotic transition. Nature 537, 548–552 (2016).
Sui, X. et al. METTL3-mediated m6A is required for murine oocyte maturation and maternal-to-zygotic transition. Cell Cycle 19, 391–404 (2020).
Falco, G. et al. Zscan4: a novel gene expressed exclusively in late 2-cell embryos and embryonic stem cells. Dev. Biol. 307, 539–550 (2007).
Marikawa, Y. & Alarcon, V. B. Establishment of trophectoderm and inner cell mass lineages in the mouse embryo. Mol. Reprod. Dev. 76, 1019–1032 (2009).
Batista, P. J. et al. m6A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15, 707–719 (2014).
Wang, Y. et al. N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nat. Cell Biol. 16, 191–198 (2014).
Geula, S. et al. m6A mRNA methylation facilitates resolution of naive pluripotency toward differentiation. Science 347, 1002–1006 (2015).
Jin, K. X. et al. N6-methyladenosine (m6A) depletion regulates pluripotency exit by activating signaling pathways in embryonic stem cells. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2105192118 (2021).
Oron, E. & Ivanova, N. Cell fate regulation in early mammalian development. Phys. Biol. 9, 045002 (2012).
Ivanova, I. et al. The RNA m6A reader YTHDF2 is essential for the post-transcriptional regulation of the maternal transcriptome and oocyte competence. Mol. Cell 67, 1059–1067 e1054 (2017).
Li, M. et al. Ythdf2-mediated m6A mRNA clearance modulates neural development in mice. Genome Biol. 19, 69 (2018).
Lasman, L. et al. Context-dependent functional compensation between Ythdf m6A reader proteins. Genes Dev. 34, 1373–1391 (2020).
Sha, Q. Q. et al. Characterization of zygotic genome activation-dependent maternal mRNA clearance in mouse. Nucleic Acids Res. 48, 879–894 (2020).
Mu, H. et al. METTL3-mediated mRNA N6-methyladenosine is required for oocyte and follicle development in mice. Cell Death Dis. 12, 989 (2021).
Chen, T. et al. m6A RNA methylation is regulated by microRNAs and promotes reprogramming to pluripotency. Cell Stem Cell 16, 289–301 (2015).
Yang, Q. et al. Highly sensitive sequencing reveals dynamic modifications and activities of small RNAs in mouse oocytes and early embryos. Sci. Adv. 2, e1501482 (2016).
Deniz, O., Frost, J. M. & Branco, M. R. Regulation of transposable elements by DNA modifications. Nat. Rev. Genet. 20, 417–431 (2019).
Franke, V. et al. Long terminal repeats power evolution of genes and gene expression programs in mammalian oocytes and zygotes. Genome Res. 27, 1384–1394 (2017).
Chen, C. et al. Nuclear m6A reader YTHDC1 regulates the scaffold function of LINE1 RNA in mouse ESCs and early embryos. Protein Cell 12, 455–474 (2021).
Liu, J. et al. The RNA m6A reader YTHDC1 silences retrotransposons and guards ES cell identity. Nature 591, 322–326 (2021).
Chelmicki, T. et al. m6A RNA methylation regulates the fate of endogenous retroviruses. Nature 591, 312–316 (2021).
Xu, W. et al. METTL3 regulates heterochromatin in mouse embryonic stem cells. Nature 591, 317–321 (2021).
Xiong, F. et al. RNA m6A modification orchestrates a LINE-1–host interaction that facilitates retrotransposition and contributes to long gene vulnerability. Cell Res. 31, 861–885 (2021).
Lanciano, S. & Cristofari, G. Measuring and interpreting transposable element expression. Nat. Rev. Genet. 21, 721–736 (2020).
Svoboda, P. Why mouse oocytes and early embryos ignore miRNAs? RNA Biol. 7, 559–563 (2010).
DeVeale, B., Swindlehurst-Chan, J. & Blelloch, R. The roles of microRNAs in mouse development. Nat. Rev. Genet. 22, 307–323 (2021).
Marcel, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal https://doi.org/10.14806/ej.17.1.200 (2011).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Wu, Z. et al. NormExpression: an R package to normalize gene expression data using evaluated methods. Front. Genet. 10, 400 (2019).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Olarerin-George, A. O. & Jaffrey, S. R. MetaPlotR: a Perl/R pipeline for plotting metagenes of nucleotide modifications and other transcriptomic sites. Bioinformatics 33, 1563–1564 (2017).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Wu, J. et al. The landscape of accessible chromatin in mammalian preimplantation embryos. Nature 534, 652–657 (2016).
Wu, J. et al. Chromatin analysis in human early development reveals epigenetic transition during ZGA. Nature 557, 256–260 (2018).
Zhang, J. D. et al. Detect tissue heterogeneity in gene expression data with BioQC. BMC Genomics 18, 277 (2017).
Hu, H. et al. AnimalTFDB 3.0: a comprehensive resource for annotation and prediction of animal transcription factors. Nucleic Acids Res. 47, D33–D38 (2019).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Agarwal, V., Bell, G. W., Nam, J. W. & Bartel, D. P. Predicting effective microRNA target sites in mammalian mRNAs. eLife https://doi.org/10.7554/eLife.05005 (2015).
Cordaux, R. & Batzer, M. A. The impact of retrotransposons on human genome evolution. Nat. Rev. Genet. 10, 691–703 (2009).
Acknowledgements
We thank the members of K.F.A., J.A.D. and A.K. laboratories for their support and comments on this work. We are grateful to B. Li’s (K.F.A. laboratory) for comments on the paper. We thank the Norwegian Transgenic Center for animal housing and oocyte collection. We also thank the Norwegian Sequencing Centre (Oslo University Hospital and University of Oslo) and the Genomics Core Facility (Norwegian University of Science and Technology) for high-throughput sequencing. This work was supported by National Institutes of Health grants R01HG008759, R01HG011469 and R01GM136886 (to K.F.A., Y.W. and A.L.); Institutional fund from the Department of Biomedical Informatics, The Ohio State University (to K.F.A., Y.W. and A. L.); Institutional fund from the Department of Computational Medicine and Bioinformatics, University of Michigan (to K.F.A., Y.W. and A.L.); South-Eastern Norway Regional Health Authority Early Career Grants 2016058 and 2018063 (to J.A.D.); Research Council of Norway, FRIPRO Grant 289467 (to J.A.D); South-Eastern Norway Regional Health Authority, Grant 2018086 (to A.K.); Research Council of Norway, FRIPRO Researcher Project 275286 (to A.K.); UiO:Life Science convergence environment grant (to Y.L., T.S., A.K., G.D.G. and J.A.D.).
Author information
Authors and Affiliations
Contributions
Y.W., Y.L., A.K., J.A.D. and K.F.A. conceived the study and designed the experiments and analyses. Y.L. carried out picoMeRIP-seq experiments. Y.W. performed the data analyses with assistance from Y.L., A.L. and A.H. T.S. and Y.L. collected mouse GV oocytes. T.S. was involved in planning experiments. Y.L., I.J. and S.K. collected other stages of mouse oocyte and embryo. Mouse material collection was supervised by G.D.G., K.T.D. and J.A.D. C.Z. performed the m6A immunofluorescence staining assay. Y.W., Y.L., T.S., A.K., J.A.D. and K.F.A. prepared the paper with input from all authors. All authors read and commented on the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Beth Moorefield, Carolina Perdigoto and Dimitris Typas in collaboration with Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
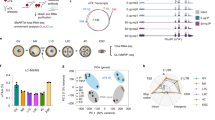
Extended Data Fig. 1 picoMeRIP-seq pipeline and m6A profiles in mouse oocytes and preimplantation embryos.
(a) Workflow of picoMeRIP-seq, created with BioRender.com. (b) Pearson correlation analyses between two biological replicates of 6 stages. (c) Overlap analysis of m6A marked genes between two biological replicates. (d) Bubble plot showing the relative enrichment in different genomic features. For each feature, the enrichment score was calculated as the log2 ratio of the observed over expected peak numbers. (e) Consensus motifs identified using the peaks called in the biological replicate 2.
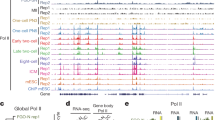
Extended Data Fig. 2 m6A profiles of repetitive stage-specifically expressed genes and key regulators for development of mouse preimplantation embryos.
(a) picoMeRIP-seq read density in exonic regions of stage-specifically expressed genes identified in Fig. 2d. (b) picoMeRIP-seq read density in exonic regions of key genes essential for mouse early embryonic development (listed in Fig. 2i). The sample-specific scale ranges are indicated in brackets with the corresponding colors.
Extended Data Fig. 3 GO analyses of m6A + and m6A − genes in Decay and ZGA genes identified in Fig. 3a, b.
Fisher’s exact test was used to calculate the one-sided P values. Only the GO terms with P value <0.05 are shown.
Extended Data Fig. 4 m6A marking and miRNA targeting profiles on imprinted genes.
Gene expression, m6A marking, miRNA targeting, number of miRNAs per gene, normalized number of miRNAs per gene in paternally- (upper panel) and maternally- (lower panel) expressed imprinted genes. For a given gene, the normalized number of miRNAs (last column, purple colored heat maps) is the sum of expression values of all miRNAs targeting this gene, where the expression value is log10 (RPM + 1). RPM, reads per million.
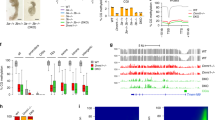
Extended Data Fig. 5 Abundant enrichment of m6A on retrotransposon-derived RNAs in mouse oocytes and early embryos.
(a) Percentage of m6A peaks overlapping with the non-exonic regions of annotated genes in GENCODE vM23 (left panel), including both intronic and intergenic regions (right panel). (b) Percentage of non-exonic m6A peaks overlapping with retrotransposons, considering both intronic and intergenic m6A peaks (left panel), intronic only (middle panel) and intergenic only (right panel). (c) Bubble plot showing the significant enrichment of m6A peaks in LTR (for GV, MII, zygote and 2-cell) and LINE (for 8-cell and blastocyst). For each of three types of retrotransposons, the enrichment score is calculated as the log2 ratio of the observed over expected peak numbers. Only the biological replicate 2 for each stage was plotted. (d) The same as in panel c, but for 8 major retrotransposon families. Only the biological replicate 2 for each stage was plotted. (e) The same as in panel c, but for representative retrotransposon subfamilies. (f) Heatmap plots showing the ratios of genomic copies/loci with >0 m6A signal score (MeRIP vs Input) and >0 expression value (RPKM), respectively (calculated using biological replicate 2), for representative retrotransposon subfamilies.
Extended Data Fig. 6 m6A profiles on ORR1A0 and ORR1A1 derived RNAs.
(a) m6A marking status, and m6A signal and expression profiles across the copies of ORR1A0 and ORR1A1. Only the internal sequences were considered here. The upper line plots show the percentage of copies/loci with m6A marking, >0 m6A signal value, >0 expression value. Color range of m6A signal: ORR1A0, −1.6 to 4.9, ORR1A1, −3.9 to 4.6. (b) m6A signal density pileups along the full-length ORR1A0 and ORR1A1 sequences. The lower bar plots show the frequency (bin size = 10 bp) of GGACU motif across all copies/loci along the full-length structure. See ‘Methods’ for more details.
Extended Data Fig. 7 m6A signal density and motif distributions along the full-length retrotransposon subfamilies.
For each of 5 subfamilies, the upper panels show m6A signal density pileups generated using the uniquely-aligned reads together with the randomly-assigned multiply-aligned reads. See ‘Methods’ for more details about the random assignment of multiply-aligned reads. The lower bar plots show the frequency (bin size = 10 bp) of RRACH motifs across all copies/loci along the full-length structure.
Extended Data Fig. 8 Length distribution of genomic copies of 5 subfamilies.
Statistics of lengths of all genomic copies for each subfamily. The length distributions were used as the cutoff to define the full-length MTA, MERVL and L1Md_T (Fig. 4d; Extended Data Fig. 7), as well as ORR1A0 and ORR1A1 (Extended Data Figs. 6b and 7) for plotting the m6A signal density pileups. In brief, for a given locus/copy, we defined it as full-length if the lengths of each part (5′ LTR, internal and 3′ LTR) were in the ranges indicated by the red colored balls which represent the most frequent lengths. See ‘Methods’ for more details.
Supplementary information
Supplementary Table 1
Expression level and m6A marking status of genes in GENCODE annotation library (vM23).
Supplementary Table 2
Developmental stage specifically expressed genes.
Supplementary Table 3
Decay and ZGA genes.
Supplementary Table 4
miRNA targeting status of genes in GENCODE annotation library (vM23).
Supplementary Table 5
Expression level, m6A marking status and m6A signal of retrotransposon loci.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Y., Li, Y., Skuland, T. et al. The RNA m6A landscape of mouse oocytes and preimplantation embryos. Nat Struct Mol Biol 30, 703–709 (2023). https://doi.org/10.1038/s41594-023-00969-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-00969-x
This article is cited by
-
ALKBH5 controls the meiosis-coupled mRNA clearance in oocytes by removing the N 6-methyladenosine methylation
Nature Communications (2023)
-
Single-cell m6A mapping in vivo using picoMeRIP–seq
Nature Biotechnology (2023)