Abstract
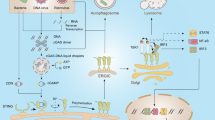
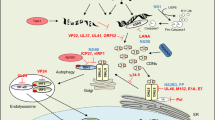
The metazoan cGAMP-activated cGAS–STING innate immunity pathway is triggered in response to genomic instability and DNA damage, thereby providing host defense against microbial pathogens. This pathway also impacts on autophagy, cellular senescence and antitumor immunity, while its overactivation triggers autoimmune and inflammatory diseases. Metazoan cGAS generates cGAMP containing distinct combinations of 3′-5′ and 2′-5′ linkages, which target the adaptor protein STING and activate the innate immune response through a signaling cascade leading to upregulation of cytokine and interferon production. This Review highlights a structure-based mechanistic perspective of recent advances in cGAMP-activated cGAS–STING innate immune signaling by focusing on the cGAS sensor, cGAMP second messenger and STING adaptor components, thereby elucidating the specificity, activation, regulation and signal transduction features of the pathway. In addition, the Review addresses progress towards identification of inhibitors and activators targeting cGAS and STING, as well as strategies developed by pathogens to evade cGAS–STING immunity. Most importantly, it highlights cyclic nucleotide second messengers as ancient signaling molecules that elicit a potent innate immune response that originated in bacteria and evolved through evolutionary adaptation to metazoans.
Similar content being viewed by others
Main
Cyclic dinucleotides (CDNs) are ancient signaling molecules that sense and respond to pathogen infection1,2,3. In the metazoan cGAS–STING pathway, aberrant or mislocalized double-stranded DNA (dsDNA) is sensed in the cytosol by the nucleotidyltransferase sensor cGAS (cyclic GMP-AMP synthase) to generate cGAMP (cyclic AMP-GMP), a diffusible cyclic dinucleotide second messenger, which in turn activates the adaptor protein STING (stimulator of interferon genes). Activation of STING triggers phosphorylation of downstream kinases and transcription factors, resulting in upregulation of cytokine and interferon production, leading to cell death4,5,6,7,8,9. Defects in cGAS and STING affect immune defense and immune adjuvant effects10, autophagy11, inflammation7 and cellular senescence7,12, while chronic activation of cGAS by self-DNA or gain-of-function mutations in the gene encoding STING can result in autoimmune and autoinflammatory diseases13,14,15. cGAS has been shown to be an innate immune sensor of retroviruses16 and is essential for the antitumor effect of immune checkpoint blockade17. The physiological and pathological relevance of the cGAS–STING pathway, and its role in mediating immunity and inflammation, has been reviewed recently18,19,20.
One advantage of second messenger signaling is that it allows for the amplification of input signals. In addition, cGAMP can be transferred to neighboring bystander cells through gap junctions21, as well as by being packaged into virions22. In addition to direct transfer, 2′3′-cGAMP can be secreted into the microenvironment by channel proteins, where patrolling immune cells can sense and import it. Thus, cGAMP can serve as an ‘immunotransmitter’ in antitumor immunity, whereas ENPP1, a membrane-bound phosphodiesterase, degrades extracellular cGAMP to attenuate cGAS–STING signaling. A non-catalytic function of nuclear cGAS involves inhibition of DNA double-strand break (DSB) repair by homologous recombination, thereby promoting tumorigenesis23,24.
There are earlier structural biology reviews on the cGAS–STING surveillance pathway8,25,26,27, so in this Review, greater emphasis has been placed on recent publications.
Sensor cGAS
Nucleotidyltransferases
Vibrio cholerae DncV and metazoan cGAS are founding members of a large family of template-independent nucleotidyltransferases that catalytically generate CDN second messengers from a pair of nucleoside triphosphates (NTPs)26,28. Their catalytic domains exhibit a common nucleotidyltransferase fold and display extreme diversity in their capacity to synthesize CDN second messengers. The catalytic domain is composed of an amino-terminal lobe containing the catalytic acidic residues, and a carboxy-terminal lobe containing a zinc-binding module that mediates DNA recognition and is involved in cGAS dimerization. The substrate-binding site is located in a cleft between these two lobes, and the CDN is generated from NTPs in two sequential reactions that involve the initial formation of a linear intermediate, which then undergoes cyclization. Notably, cGAS and DncV homologs are widespread in bacteria and are termed CD-NTases (cGAS/DncV-like nucleotidyltransferases); members of this family rapidly initiate signaling responses to changing environmental conditions through multi-turnover amplification of CDN production.
The active site of CD-NTases is composed of donor and acceptor nucleotide-binding pockets, which are lined by acidic catalytic residues, and two divalent cations that mediate SN2 nucleophilic substitution reactions. In the first step, the nucleobase and α-phosphate of the donor nucleotide is transferred to the 2′ or 3′ oxygen of the acceptor nucleotide to form a linear dinucleotide pppNpN intermediate. Cyclization occurs in the second step, following a switch of the nucleobase in the donor and acceptor positions, with the α phosphate of the intermediate transferred intramolecularly to the 2′ or 3′ oxygen of the acceptor nucleotide. Notably, both steps are catalyzed within a single binding pocket29,30.
3′3′-cGAMP produced by DncV cyclase from Vibrio cholerae
The first report on the generation of hybrid cGAMP was a demonstration of its synthesis from ATP and GTP by DncV, a dinucleotide cyclase associated with the V. cholerae strain behind the seventh cholera pandemic31. The report showed that cGAMP is a signaling molecule involved in V. cholerae virulence, and that DncV is required for the bacteria to colonize the intestine and downregulates V. cholerae chemotaxis. Follow-up studies demonstrated that cGAMP produced from DncV contained a pair of 3′-5′ linkages, and that the enzyme was constitutively active even in the absence of dsDNA.
2′3′-cGAMP produced by dsDNA-bound metazoan sensor cGAS
In a key set of seminal discoveries, cGAS was identified as a cytoplasmic DNA sensor that activates the type I interferon pathway by synthesizing the second messenger cGAMP from GTP and ATP in the presence of dsDNA32,33. Further, cGAMP bound to and activated the adaptor protein STING, resulting in the activation of the transcription factor IRF3 and subsequent induction of interferon β.
Next, the phosphodiester linkages of cGAMP were elucidated by crystallographic, enzymatic digestion, nuclear magnetic resonance, mass spectroscopy and chemical synthesis approaches29,34,35,36,37. Crystal structures of the catalytic domain of mouse cGAS (cGASCD) in the free state and bound to added ATP, GTP and both ATP and GTP have demonstrated that, upon dsDNA binding, cGASCD is activated through conformational transitions, resulting in the formation of a catalytically competent and accessible nucleotide-binding pocket for cGAMP generation29. Structural studies, complemented by chemical, biochemical and cellular assays, have established that cyclization occurs in a stepwise manner through initial generation of 5′-pppG(2′,5′)pA prior to cyclization to c[G(2′,5′)pA(3′,5′)p] (designated 2′3′-cGAMP). 2′3′-cGAMP is positioned in the catalytic pocket such that the adenosine base is stacked over a Tyr ring while the guanosine base is anchored through hydrogen bond formation29. Furthermore, mutant cGASCD dsDNA-binding residues, or mutant residues in the catalytic pocket, exhibit reduced or abrogated activity.
Three distinct dsDNA-binding sites facilitate oligomerization of cGASCD
Human cGASCD (Fig. 1a), both in the apo state and when bound to dsDNA, has been observed to form a dimer in crystal structures, resulting in cGASCD interacting with dsDNA through two juxtaposed binding sites (labeled A and B, Fig. 1b) that are associated with 2:2 cGASCD–DNA complex formation38,39. Functional studies have established that, at major site-A (Fig. 1c) and minor site-B (Fig. 1d), electrostatic interactions between the DNA phosphate backbone and basic amino acids on surface-exposed loops and structured elements of cGASCD contribute to cGAS activation, as does the protein-protein interface across the dimer. cGAS activation strongly depends on dsDNA length, giving rise to cooperative sensing by cGASCD dimers that are capable of forming ladder-like networks along longer dsDNA40.
a, Domain architecture of human c-GAS. The catalytic domain of human cGAS is labeled hcGASCD. aa, amino acids. b, Structure of the hcGASCD–DNA complex at 3.2-Å resolution, with emphasis on the 2:2 stoichiometry (PDB: 6EDB). This complex also contained a SRY protein element that targeted its sequence-specific DNA element, which has been deleted for clarity. The pairs of hcGASCD and DNA elements are labeled (1) and (2), and distinct protein-DNA interfaces are labeled site-A and site-B. c, Structural details of the major site-A interface of hcGASCD (2) bound to DNA (2). DNA-phosphate-interacting basic amino acids are labeled and shown in stick representation. d, Structural details of the minor site-B interface of hcGASCD (1) bound to DNA (2). DNA-phosphate-interacting basic amino acids are labeled and shown in stick representation. e, The structure of the complex shown in b together with DNA (3) emphasizing the major protein-DNA site-C interface. f, Structural details of the major site-C interface of hcGASCD (1) bound to DNA (3). DNA-phosphate-interacting basic amino acids are labeled and shown in stick representation. The intermolecular contacts are partitioned between the α region, KRKR loop and KKH loop segments.
More recently, another major dsDNA-binding site on human cGASCD, labeled site-C (Fig. 1e), has been shown to involve conserved basic residues that interact with the sugar-phosphate backbone of DNA (Fig. 1f)41. This interface increases the cGAS–DNA binding valency, thereby facilitating the formation of cluster networks for cooperative sensing of longer DNA. Studies of cGASCD site-C interfacial residue alterations support the essential contribution of site-C to the enzymatic function of cGAS.
dsDNA-mediated conformational changes in cGASCD activation loop
Initial studies focused on the major DNA-binding site on cGASCD, site-A, which resides antipodally of the catalytic pocket, where cGAMP formation occurs (Fig. 1b). Comparison of the active site in the crystal structures of mouse cGASCD in the apo state (Fig. 2a) with that bound to dsDNA and cGAMP (Fig. 2b) identified concerted conformational changes that are associated with reorganization of a pair of loops and β-strands lining the cGASCD catalytic pocket on complex formation29,37,38. These conformational transitions (Fig. 2c and its insert) occur because of electrostatic interactions between basic side chains and the phosphate backbone on complex formation, explaining sequence-independent recognition. Closer inspection revealed that a loop containing a GS (Gly-Ser) motif spanning β-strands was flexible in the apo state but adopted a well-defined conformation composed of a short helix in the dsDNA-bound state (black arrow, Fig. 2b) and was positioned in proximity to two acidic catalytic residues29,39. Such stabilization of the GS-containing loop may play a critical regulatory role among nucleotidyltransferases, given that a similar stabilization has been observed in 2′,5′-OAS (2′,5′-oligoadenylate synthetase) in complex with dsRNA and substrate analogs42,43.
a,b, Comparison of the catalytic pocket of mouse cGASCD (mcGASCD) in the 2.0-Å-resolution structure in the apo state (a; PDB: 4K8V) and in the 2.3-Å-resolution structure of cGAMP in the DNA-bound state (panel b; PDB: 4K9B). The black arrow in b highlights one of the many conformational changes on complex formation. c, Superposition of crystal structures of apo- (in blue; PDB: 4K8V) and dsDNA-bound (in magenta; PDB: 4K96) mouse cGASCD. 2′3′-cGAMP is positioned in the binding pocket in a stick representation. The boxed expanded view on right shows the superposition in the catalytic pocket, with arrows highlighting the conformational changes on complex formation. d, Domain architecture of hcGASCD, emphasizing the substitutions that are required for successful crystallization of the DNA-bound complex. e, The 2.3-Å-resolution structure of the K187N L195R dual-mutant hcGASCD–DNA complex (PDB: 6CT9), emphasizing the 2:2 complex. The dual mutant is positioned at the site-A protein-DNA interface. The structure of the K299E R300A K301E triple-mutant complex at 2.1-Å resolution (PDB: 6EDC) is not shown. The triple mutant is positioned at the site-C protein–DNA interface. f, The structure of the human cGASCD–DNA complex (PDB: 6EDB), stressing the interaction site-A between cGASCD and DNA. The amino acids mentioned in the text are labeled and shown in a space-filling representation. g–j, Chemical formulas of the hcGAS inhibitors RU-521 (g), G150 (h), PF-06928215 (i) and CU-76 (j).
Human cGASCD–dsDNA complex and DNA length, and Mn2+-dependent activation
To date, two crystal structures for modified human cGASCD (Fig. 2d) bound to 17-base-pair (bp) dsDNA have been described: one with cGASCD containing two alterations, K187N and L195R, on the site-A DNA-binding surface44, and another with cGASCD containing three alterations, K299E, K300A and K301E, on the site-C DNA-binding surface (Fig. 2d)41. The availability of this pair of structures of human cGASCD–DNA complexes (Fig. 2e) should facilitate the design of inhibitors targeting dsDNA-bound human cGASCD.
Notably, generation of cGAMP by human cGASCD is reduced compared with other mammalian analogs and is dependent on DNA length; >45 bp of dsDNA is required for efficient production of cGAMP. This regulation has been attributed to Lys187 and Leu195 in human cGASCD, which map to a long α-helical segment. These two amino acids (Fig. 2f) modify the site-A DNA-binding surface of human cGASCD, resulting in oligomerization-mediated recognition of longer DNA44.
Recent structural studies on the linear pppG(2′-5′)G intermediate in complex with cGASCD have shown that Mn2+-activated cGASCD undergoes global changes that are similar to those that DNA-activated cGASCD undergoes, except for widening of the catalytic pocket for substrate entry in 2′3′-cGAMP formation45. Unlike Mg2+, which coordinates to the acidic catalytic-triad residues of cGAS, Mn2+ instead coordinates to the triphosphate of the pppG(2′,5′)G intermediate. Importantly, the pppG(2′-5′)G intermediate adopts an inverted orientation in the active pocket in the Mn2+-activated complex.
Reprogramming of cGASCD dinucleotide linkage specificity
Comparison of the structures of DncV, which produces 3′3′-cGAMP, and cGAS, which produces 2′3′-cGAMP, highlights similarities and differences46. Despite minimal sequence identity, the overall structures and the catalytic core of DncV and cGAS exhibit remarkable similarity. Nevertheless, unlike cGAS, which adopts a dimeric fold and requires dsDNA for activation, DncV adopts a monomeric fold and an autoactivated state that is constitutively active. This unexpected behavior of DncV can be explained by its catalytic basic amino acids being positioned for competent divalent metal-ion coordination, even in the apo state, and by it not changing conformation upon binding substrates. Finally, unlike cGAS, for which regulatory loops restrict access to the substrate channel in the apo state and catalysis occurs through a pppG(2′-5′)A linear intermediate29, these loops are missing in DncV. This results in a substrate-competent, permanently open active site, and DncV cyclization proceeds through a pppA(3′-5′)G linear intermediate46. A detailed comparison of extended and recessed residues lining the catalytic pocket of DncV and cGAS established that active-site remodeling involving the size of cGAS residues Thr211 and Arg376 (Fig. 2f) controls substrate rotation and phosphodiester linkage specificity, allowing reprogramming of the cGAS active site through replacements of these residues to their DncV amino acid counterparts to produce 3′3′-cGAMP46.
Small-molecule targeting of the cGAS catalytic pocket
Current efforts to identify cGASCD-targeting inhibitors have focused on those that either target the active site or compete with DNA for binding the enzyme15,47,48,49. The most successful efforts to date have used high-throughput screens coupled with structure-guided optimization. Such efforts have identified RU.521 (Fig. 2g)50 and G150 (Fig. 2h)51, both of which bind within the cGASCD active site, with G150 exhibiting potent in vitro selectivity in humans (half-maximum inhibitory concentration (IC50)= 10.2 nM) compared with mice (IC50 > 25 μM). Other inhibitors, such as PF-06928125 (Fig. 2i), also target the cGASCD active site, while showing in vitro, but not cellular, inhibitory activity52,53. Further, other inhibitors, such as CU-76 (Fig. 2j), which was identified from a virtual screen of the cGASCD catalytic pocket, followed by synthetic optimization, have yet to be characterized by X-ray crystallography54. There has been less progress toward the identification of potent inhibitors that disrupt DNA binding by cGASCD (ref. 15).
Impact of cGAS modifications on activity
The enzymatic activity of cGAS can be modulated by post-translational modifications, including phosphorylation, ubiquitination, sumoylation, acetylation, palmitoylation and glutamylation55,56,57. Such modifications regulate the integrity of the active site, on dimer formation and on protein-DNA contact at site-A, site-B or site-C, as well as on nucleosomal interactions and liquid phase separation.
Pathogen-mediated evasion of cGAS-mediated immunity
It has been shown that herpes simplex virus-1 (HSV-1) can infect mice and humans, but not other non-human primates, by the HSV-1 UL37 tegument protein deaminating a single Asn in the activation loop of cGAS, thereby facilitating HSV-1 lytic replication58. By contrast, Asn mutants or deaminase-deficient HSV-1 robustly induced cytokines in a cGAS- and STING-dependent manner. A similar deamidation strategy is used by viruses to activate RIG-I to evade cytokine production59.
Other pathogens evade cGAS-mediated immunity using both direct and indirect strategies27,60. As an example of a direct evasion strategy, the UL31 protein of human cytomegalovirus (HCMV) directly targets cGAS, inducing by dissociating cGAS-bound DNA and inhibiting cGAS’s enzymatic function61. By contrast, in an example of an indirect strategy, Zika virus targets cGAS by inducing an inflammatory response, whereby enhanced stabilization of caspase-1 by NS1 promotes the degradation of cGAS62.
cGAS–DNA phase separation activates innate immune signaling
Recently it has been shown that dsDNA-induced liquid–liquid phase separation of cGAS promotes cGAMP production, thereby providing an additional layer in DNA-length-dependent activation of cGAS-mediated innate immune signaling63. This has been attributed to enhancement of cGAS–dsDNA liquid phase separation by the basic and disordered N-terminal segment of cGAS, Zn2+ cations and long DNAs; the cGAS enzyme is highly concentrated in liquid droplets resulting from multivalent interactions between cGAS and long DNA.
Subsequently, it was shown that the protein-DNA site-C interface also contributed to liquid phase separation, with alterations along this interface disrupting condensation41, supporting a multivalent interaction-mediated cluster model whereby both the N-terminal domain of cGAS and the site-C cGAS–DNA interface account for dsDNA-mediated phase separation.
Regulation of cellular cGAS activity
Studies have also assessed the factors that contribute to cellular homeostasis by balancing innate immune activation and cytosolic DNA degradation64. In this regard, BAF (barrier-to-autointegration factor 1) has been shown to dynamically outcompete cGAS for binding to nuclear DNA, thereby preventing innate immune activation65. cGAS–DNA liquid–liquid phase separation has been shown to restrict access of the cytosolic exonuclease TREX1 to the outer periphery of cGAS liquid droplets, thereby suppressing TREX1 catalytic function and restricting DNA degradation66. In addition, a human TREX1 alteration that causes the severe autoimmune Aicardi–Goutieres syndrome altered interactions of TREX1 with cGAS–DNA droplets.
Another issue involves what regulates cGAS activity during cell cycle transitions, such as entry into mitosis, conditions under which nuclear-envelope breakdown facilitates cGAS association with chromatin. A careful study has established that hyperphosphorylation of the intrinsically disordered and highly basic N terminus of cGAS, as well as tethering of cGAS by chromatin that inhibits cGAS oligomerization, both contribute to preventing cGAS phase separation into liquid droplets, thereby silencing cGAS activity and preventing an autoimmune reaction67.
Viral tegument proteins restrict cGAS–DNA phase separation
The role of viral pathogens in manipulating cGAS–DNA liquid–liquid phase separation can result in pathogen-mediated immune evasion. To this end, the structurally related viral tegument proteins ORF52 and VP22, which contain positively charged tails, adopt a conserved mechanism that dictates pathogen-mediated regulation by outcompeting host cell cGAS–DNA liquid–liquid phase separation68,69,70.
DNA-bound cGAS at the nucleosomal level
cGAS has been identified in the cell nucleus, where it binds more tightly to nucleosomes than to naked DNA. Further, cGAMP formation by cGAS was impeded at both the mononucleosome and chromatin-fiber level, thereby implicating chromatin in suppression of cGAMP formation23,71,72. A structural explanation underlying nucleosome-mediated suppression of cGAS activity has emerged from cryo-EM studies of complexes of cGASCD bound to the nucleosome core particle (NCP), both in mice73,74,75 and humans76,77,78. The mouse cGASCD–NCP complex exhibited 1:1 stoichiometry (Fig. 3a), whereby Arg222 and Arg241, associated with loop elements projecting from cGASCD site-B, bound tightly to the acidic patch on the histone H2A-H2B component (insert, Fig. 3a), a universal recognition site for a variety of chromatin-binding proteins79. Such an alignment resulted in disruption of the dimeric state of cGASCD and also prevented dsDNA recognition of site-A and site-B, thereby providing insights into two factors that contribute to the inhibitory mechanism.
a, A structure of mouse cGASCD bound to the NCP, at 3.1-Å resolution (PDB: 7A08). The insert emphasizes protein-protein contacts involving site-B DNA interface of cGASCD. b, The structure of human cGASCD bound to the NCP, at 4.9-Å resolution (PDB: 7CCR). The insert emphasizes additional protein-DNA contacts involving α-region, KRKR loop and KKH loop lining the site-C DNA interface of cGASCD.
By contrast, in humans, the cGASCD monomer bridged adjacent NCPs (Fig. 3b), with each cGASCD monomer bound simultaneously to the acidic patch of one NCP through cGASCD site-B and to nucleosomal DNA of the adjacent NCP through cGASCD site-C (insert, Fig. 3b). In addition, a ladder-shaped higher-order assembly was also detected at lower resolution, with protein-protein contacts through site-B and protein-DNA contacts through site-C likely contributing to cGASCD-nucleosome oligomerization76.
cGAS-like receptors as innate immune sensors of dsRNA
Animal genomes contain genes encoding proteins that are homologous to human cGAS. A recent study has established that such cGAS-like receptors (cGLRs) sense distinct nucleic acid scaffolds to catalyze the synthesis of CDNs with distinct linkages that activate the innate immune response80,81. Structural studies have established that, similar to human cGAS, mammalian and insect cGLRs adopt a nucleotidyltransferase fold whose nucleic acid-binding surface is modified through insertions and deletions and is lacking a Zn module. Remarkably, Drosophila simulans cGLR1 binds dsRNA that is more than 30 bp in length, thereby avoiding the self-recognition that is associated with the 21- to 23-bp dsRNAs produced by RNA silencing80. Notably, on activation by dsRNA, Drosophila simulans cGLR1 unexpectedly synthesizes 3′2′-cGAMP.
Highlights of recent cGAS research
Overall, the above sections summarize the role of cGAS-mediated production of 2′3′- and 3′2′-cGAMP linkage isomers and compare approaches to reprogramming cGAS dinucleotide linkage specificity, and they explore the design of small molecules targeting the ligand-binding catalytic pocket. They also address the contribution of dsDNA-induced cGAS liquid–liquid phase separation in the formation of biomolecular condensates that promote cGAMP production, as well as its restriction by viral tegument proteins. In addition, they discuss the recent identification of cGAS-like receptors as innate immune sensors of dsRNA, as well as the molecular principles underlying nucleosome-mediated suppression of cGAS activity in the nucleus.
Second messenger cGAMP
Given the importance of the cGAMP-mediated immune response associated with the cGAS–STING antiviral defense pathway, attempts have been made to decipher cGAMP conformations in the free and STING-bound states, as part of ongoing efforts to design cGAMP analogs as vaccine adjuvants and immunotherapeutic agents against disease.
2′3′-cGAMP formation
The initial assumption that cGAMP produced by cGAS contained both 3′-5′ linkages, on the basis of mass spectroscopy data32,33, was updated when it was definitively demonstrated by crystallographic, spectroscopic and nuclease-cleavage approaches that the GpA adopted a 2′-5′ linkage while ApG retained a 3′-5′ linkage29,34,35,36. Nevertheless, the molecular basis underlying preferential binding of 2′3′-cGAMP to its adaptor, STING, relative to its other linkage counterparts, remained an enigma. This conundrum was resolved when it was demonstrated using spectral and computational approaches that 2′3′-cGAMP adopted an organized free-ligand conformation that resembled its STING-bound counterpart, thereby paying low enthalpy and entropy costs associated with complex formation82. By contrast, 3′3′-cGAMP and 3′2′-cGAMP adopted highly flexible, ligand-free conformations.
Phosphodiesterases that specifically cleave cGAMP 2′-5′ and 3′-5′ linkages
ENPP1 (ecto-nucleotide pyrophosphatase phosphodiesterase 1) was identified from activity-guided purification to exhibit dominant and highly specific hydrolyzing activity against 2′3′-cGAMP, but not 3′3′-cGAMP, in cultured cells83. ENPP1 is a type II transmembrane glycoprotein whose extracellular domain is composed of SMB, catalytic and nuclease-like domains. X-ray crystallographic studies have been undertaken on ENPP1 (lacking the SMB domain, Fig. 4a) in the apo form and in complex with the post-reaction intermediate of 2′3′-cGAMP84,85,86. Cleavage of the 2′-5′ linkage in the post-reaction state of the 2′3′-cGAMP complex resulted in formation of the 5′-pApG (pA(3′,5′)pG) linear intermediate (Fig. 4b), with adenosine positioned in the nucleotide-binding N pocket and guanosine positioned in the guanine-binding G pocket (Fig. 4c). The authors proposed a two-step in-line displacement mechanism, whereby Thr238 in the active site catalyzes cleavage first at the 2′-5′ linkage of 2′3′-cGAMP, generating the 5′-pApG intermediate with the contribution of two Zn2+ cations. This is followed by a flip over that positions the guanosine in the N pocket that leads to cleavage of the 3′-5′ linkage, yielding 5′-AMP and 5′-GMP.
a, Domain architecture of mouse ENPP1, emphasizing catalytic and nuclease-like domains. b, Crystal structure of catalytic and nuclease-like domains of ENPP1 in complex with pApG, a product of cGAMP hydrolysis, at 1.8-Å resolution (PBD: 6AEK). c, pApG is shown bound in the catalytic pocket of ENPP1. The G of pApG is positioned in the G pocket, and A is positioned in the N pocket. d, Domain architecture of VACV poxin, which contains N-terminal protease-like and C-terminal nuclease domains. e, The crystal structure of 2′3′-cGAMPS bound to VACV poxin in the post-reactive state, at 2.1-Å resolution (PDB: 6EA9). The protein is shown in a ribbon representation, and the 2′3′-cGAMPS analog is shown in a space-filling representation. Monomers 1 and 2 align to form a dimeric arrangement. f, Positioning of 2′3′-cGAMPS relative to residues involved in hydrolysis of the 3′-5′ phosphodiester bond. g, Topology of cGAMP when bound to VACV poxin. The position of the distorted adenine has shifted by about 60° relative to its position when bound to cGASCD. 2′-OH endo refers to the pucker of the sugar ring. h, Proposed mechanism of metal-independent hydrolysis of 2′3′-cGAMP by VACV poxin.
Studies of phosphodiesterases in V. cholerae have led to the identification of HD-GYP-containing V-cGAPs (cGAMP-specific phosphodiesterase in V. cholerae) that specifically cleaved 3′3′-cGAMP in a two-step reaction to first form the 5′-pApG intermediate, which was further hydrolyzed to 5′-ApG87. Notably, DncV activity in bacteria is also evaded by viral Acb1 enzymes that degrade cyclic dinucleotide signals88.
Poxins as 2′3′-cGAMP-specific nucleases
A biochemical screen of mammalian viruses was undertaken to identify poxvirus immune nucleases (poxins) that specifically cleave the second messenger 2′3′-cGAMP, thereby allowing viruses to evade cGAS–STING immunity89. These poxins, which are composed of N-terminal protease-like and C-terminal nuclease domains (Fig. 4d), degrade 2′3′-cGAMP through metal-independent cleavage of the 3′-5′ linkage, resulting in linear 5′-Gp(2′-5′)Ap, restricting STING-dependent activation and evading cGAS–STING-mediated innate immunity. The molecular basis underlying selective cleavage was elucidated following crystal-structure determination of vaccinia virus (VACV) poxin in the apo, pre-cleavage (as a thiophosphate analog) and post-cleavage states of 2′3′-cGAMP degradation. VACV poxin adopts a V-shaped dimeric fold with the ligand-binding pocket positioned between the N- and C-terminal domains of opposing monomers (Fig. 4e). The 2′3′-cGAMP analog is positioned in a deep pocket with its 2′-5′ phosphodiester linkage buried in the interior of the fold, while its 3′-5′ linkage is presented to the active site for hydrolysis (Fig. 4f). Key catalytic residues, confirmed by impact of alterations, are positioned proximal to the 3′-5′ cleavage step (Fig. 4f), with the bound 2′3′-cGAMP adopting a distorted conformation (Fig. 4g) to facilitate metal-independent general acid- and base-mediated catalysis (Fig. 4h).
2′3′-cGAMP transport across membranes mediated by CDN carrier SLC19A1
Recently, CRISPR-based genome-wide screens have led to the identification of solute carrier SLC19A1 as a major CDN transporter across cell membranes, and such transport is required for STING activation90,91. Human SLC19A1 (hSLC19A1) is composed of 12 transmembrane segments (TMs) (Fig. 5a). Cryo-EM-derived structures of SLC19A1 in the apo state and bound to 2′3′-cGAMP, the thiophosphate analog of 2′3′-cGAMP (2′3′-cGAMPS) and 3′3′-cAA have been solved92. The 12 TMs of hSLC19A1 adopt a canonical major facilitator superfamily fold composed of TM1–TM6 and TM7–TM12 segments that together adopt an inward-open conformation, whose cryo-EM structure was solved in the apo state through utilization of a nanodisc-reconstituted hSLC19A1-antigen-binding fragment (hSLC19A1-Fab) system (Fig. 5b). In the inward-open conformation, the channel is wider on the intracellular side, narrows towards the middle of the transmembrane segment and is closed off at the extracellular side (Fig. 5c,d). This inward-open conformation is retained on binding to 2′3′-cGAMP (Figs. 5e), 2′3′-cGAMPS and 3′3′-cAA, with two molecules of tightly associated and compact CDNs bound in the inward-open cavity, as shown for the 2′3′-cGAMP complex (Fig. 5f,g). The bound CDN pairs are stacked in the dimer, and the CDN pairs are positioned such that their phosphates are directed in an upward/downward direction (Fig. 5f).
a, Domain architecture of hSLC19A1, subdivided into transmembrane TM helices TM1–TM6 and TM7–TM12. b, The cryo-EM structure of apo-hSLC19A1 in a nanodisc-reconstituted hSLC19A1-Fab system adopts an inward-open conformation, at 3.4-Å resolution (PDB: 7XPZ). hSLC19A1 is shown in blue, and the Fab is shown in gray. c, The apo conformation of hSLC19A1. TM1–TM6 are shown in yellow; TM7–TM12 are shown in magenta. d, Cut-open view of electrostatic potential surface of the inward-open structure of apo-SLC19A1. e, The cryo-EM structure of 2′,3′-cGAMP-bound hSLC19A1 complex aligned in an inward-open conformation, at 3.3-Å resolution (PDB: 7XQ2). Two molecules of interdigitated cGAMP form a compact alignment. f,g, Expanded segments showing the 2′,3′-cGAMP-bound hSLC19A1 complex in two views, highlighting the positioning of a pair of interdigitated 2′3′-cGAMP between TM1–TM6 and TM7–TM12 in an inward-open conformation.
SLC19A1 (refs. 92,93) and the related SLC46A1 (ref. 94) are major transporters for folate cofactors and antifolate therapeutics, with insights into ligand capture having emerged from cryo-EM studies of their complexes. The folate- and antifolate-bound SLC19A1 structures also adopt an inward-open conformation, except that one equivalent of ligand is bound within the upper and narrower segment of the SLC19A1 channel.
A future challenge will be to extend the structural studies of the CDN-bound SLC19A1 complex in the inward-open state (Fig. 5e) to trap and structurally characterize CDNs bound to SLC19A1 in the outward-open and occluded states.
Recently, ATP-binding cassette transporter ABCC1 has been identified as a mediator of direct ATP-dependent 2′3′-cGAMP export that modulates STING-dependent immunity95. The results of functional ABCC1 overexpression and deficiency studies highlight the regulatory role of ABCC1-mediated cGAMP export in limiting cell-intrinsic activation of STING and in ameliorating STING-dependent autoimmune Aicardi–Goutieres syndrome.
Highlights of recent cGAMP research
Overall, the above sections summarize the unexpected diversity of linkages adopted by cGAMP, the molecular insights governing SLC19A1-carrier-mediated cGAMP transport across membranes, and regulation of cGAMP levels by both phosphodiesterases and poxins.
Adaptor STING
STING is primarily composed of a four-transmembrane helical N-terminal segment that localizes to the endoplasmic reticulum, a central cytosolic CDN-binding domain and a C-terminal tail that is involved in binding to the TBK1 kinase (Fig. 6a). STING was initially shown to be a critical signaling molecule in the innate immune response pathway that senses the presence of nucleic acids in the cytosol96,97,98,99. STING, which forms a dimeric scaffold, is a known sensor of cyclic dinucleotides100, and has a critical role as an adaptor in the cGAS–STING pathway101,102. The ligand-binding pocket of metazoan STING is targeted by the 2′3′-cGAMP second messenger32,33, which induces its translocation from the endoplasmic reticulum membrane to the ER-Golgi intermediate and Golgi compartments103, where it recruits the IκB and TBK (TANK-binding) kinases. Phosphorylation events in turn result in activation of NFκB (nuclear factor κB) and IRF3 (IFN regulatory factor 3) factors, leading to the production of proinflammatory and antiviral cytokines, including IFNs (type I interferons), to inhibit pathogen infection5.
a, Domain architecture of human STING, containing N-terminal TM, central LBD and CTT domains. b,c, Crystal structures of apo-NvSTINGLBD in ‘rotated’ (b, 2.1-Å resolution, PDB: 5CFO) and ‘unrotated’ (c, 2.85-Å resolution, PDB: 5CFR) states. d,e, Crystal structures of 3′3′-cGAMP–NvSTINGLBD complex in a ‘rotated’ state (d, 2.0-Å resolution, PDB: 5CFM) and the 2′3′-cGAMP–NvSTINGLBD complex in a ‘partially rotated’ state (e, 2.1-Å resolution, PDB: 5CFQ). f,g, Crystal structure of 2′3′-cGAMP bound to hSTINGLBD, at 2.2-Å resolution (f; PDB: 4LOH) and positioning of 2′3′-cGAMP in the catalytic pocket of hSTINGLBD (g). h,i, Crystal structure of 3′2′-cGAMP bound to DrSTINGLBD, at 2.0-Å resolution (h; PDB: 7MWZ), and positioning of 3′2’-cGAMP in the catalytic pocket of DrSTINGLBD (i). j,k, Crystal structure of 3′3′-cGAMP bound to FsSTINGLBD at 1.8-Å resolution (j; PDB: 6WT4) and positioning of 3′3′-cGAMP in the catalytic pocket of FsSTINGLBD (k).
2′3′-cGAMP bound to the ligand-binding domain of human STING
Early structural studies established that the ligand-binding domain (LBD) of STING (STINGLBD) adopted a symmetrical dimeric topology in both the free and 3′3′-cGG-bound states, with the second messenger bound in a recessed conserved central binding pocket between subunits104,105,106,107,108. Bound CDNs are positioned with their sugar and phosphate moieties directed towards the bottom of the pocket, while their bases are oriented upwards in a parallel alignment. The bound CDNs are embraced through a range of intermolecular hydrogen bonding and hydrophobic contacts, and shape complementarity accounts for the affinity and specificity of recognition. Additional structural studies on the binding of distinct linkage isomers of cGAMP to mouse and human STINGLBD established that formation of a signaling-competent state was manifested in an inward shift of a pair of symmetrically related α2 helices of STINGLBD and the concomitant formation of a lid-forming four-stranded β-sheet cap36,109. Notably, 2′3′-cGAMP binds to human STINGLBD with a higher affinity than other linkage isomers35,36,109.
2′3′-cGAMP bound to sea anemone STINGLBD
Functional cGAS and STING homologs have been identified in sea anemone Nematostella vectensis, an animal divergent from humans by more than 500 million years of evolution. Structural studies on the cytoplasmic domain of N. vectensis (Nv) STINGLBD in the apo state and when bound to 2′3′-cGAMP and 3′3′-cGAMP have identified a deeply conserved STINGLBD conformational intermediate, providing insights into the ancient origin and mechanism of 2′3′-cGAMP signaling110. NvSTINGLBD in the apo state adopts two conformations, termed ‘rotated open’ wing alignment (helical tip 46-Å separation, Fig. 6b) and ‘unrotated closed’ wing alignment (30-Å separation‚ Fig. 6c). NvSTINGLBD retains the ‘rotated open’ state on binding 3′3′-cGAMP (and 3′3′-cCC) (46-Å separation, Fig. 6d), while a unique in-between conformation, reflecting a ‘partially rotated intermediate’ state, was identified on binding 2′3′-cGAMP (36-Å separation‚ Fig. 6e).
Indeed, this same ‘partially rotated intermediate’ conformation was also observed for 2′3′-cGAMP in complex with human STINGLBD (Fig. 6f,g)109. In addition, loop elements within the STING dimer form a four-stranded β-sheet lid over the bound cGAMP on complex formation. Structural studies led to the conclusion that the primary function of CDN second messengers is to stabilize a specific (either ‘partially rotated intermediate’ or ‘rotated open’) conformation of STINGLBD (ref. 110).
3′2′-cGAMP bound to Drosophila STINGLBD
Unexpectedly, Drosophila (Dr) STINGLBD is bound selectively by 3′2′-cGAMP, on the basis of the structure of the complex (Fig. 6h), both in vitro and in permeabilized cells, to trigger the innate immune response80. Comparison of amino acid-nucleotide contacts in the crystal structure of 2′3′-cGAMP bound to dimeric human STINGLBD (Fig. 6g) with those in the structure of 3′2′-cGAMP bound to dimeric DrSTINGLBD (Fig. 6i) explains this selectivity. Notably, the ‘partially rotated intermediate’ conformation was observed for 3′2′-cGAMP in complex with DrSTINGLBD (Fig. 6h), with 3′2′-cGAMP formation occurring notably through a pppA(2′-5′)pG linear intermediate.
3′3′-cGAMP bound to bacterial STINGLBD
In a key development, recent research has highlighted the bacterial origin of CDN recognition by STING111, through investigation of bacterial systems containing CD-NTases, wherein STING is fused to a TIR (Toll/interleukin-1 receptor) domain. Binding studies involving bacterial STINGLBD have established high affinity (nanomolar) and strict specificity for 3′-5′ linked CDNs, such as 3′3′-cGG and 3′3′-cGAMP, with a preference for the former over the latter. In its crystal structure, Flavobacteriaceae sp. (Fs) STINGLBD bound to 3′3′-cGAMP adopts a compact dimeric scaffold (Fig. 6j) when compared with its human (Fig. 6f) and Drosophila (Fig. 6h) counterparts, with the latter containing metazoan-specific insertions within the β-strand lid domain, an addition of a C-terminal α-helix and an unstructured C-terminal tail. Notably, an Asp residue of FsSTINGLBD makes a sequence-specific contact with guanine nucleobase (Fig. 6k); this interaction is specific for bacterial STINGLBD. In turn, 3′3′-CDNs targeted to bacterial STINGLBD trigger filament formation through TIR effector domain oligomerization, leading to rapid NAD+ cleavage by its hydrolase activity. Bacterial STINGLBD does not bind CDNs containing 2′-5′ linkages, and it remains to be demonstrated whether a switch in linkage specificity could be engineered through binding-pocket alterations within bacterial STINGLBD. Critically, the above results demonstrate that CDN sensing by STING originated in bacteria with evolutionary adaptation to metazoans as a consequence of metazoan-specific insertions111.
Small molecules bound to 2′3′-cGAMP pocket of mouse and human STINGLBD
Much effort has been directed towards identification of both agonists and antagonists targeted to dimeric STINGLBD in an effort to control aberrant activation of innate immune pathways15,112. The earliest efforts focused on the acridanone derivative CMA (Fig. 7a)113 and the xanthenone derivative DMXAA (Fig. 7b)114, both mouse-specific activators of STINGLBD, with crystallographic studies establishing that CMA and DMXAA bound as dimers in a head-to-tail orientation in the ligand-binding pocket of dimeric mouse STINGLBD (refs. 109,113). The crystal structure of the DMXAA-mouse STINGLBD complex (two DMXAAs bound per STINGLBD dimer) is shown in Figure 7c, and the intermolecular contacts are outlined in Figure 7d (ref. 109). Follow-up studies identified STINGLBD binding pocket and lid-region substitutions that allowed binding of DMXAA to substituted human STING, thus facilitating potential reciprocal engineering of DMXAA analogs for targeting human STINGLBD (ref. 115). Next, a nitro-furan derivative, C-178 (Fig. 7e), was identified from a cell-based screen to inhibit mouse STINGLBD, thereby reducing STING-mediated cytokine production116. C-178 was shown to covalently target Cys91 in the transmembrane domain of STING, in the process blocking activation-induced palmitoylation of STING. Recently, orally available non-nucleoside agonists, including compound SR-717 (Fig. 7f)117, amidobenzimidazole AZBI (Fig. 7g)118 and benzothiophene analog MSA-2 (Fig. 7h)119, that target human STINGLBD, and in the process exhibiting strong antitumor activity, have been identified. X-ray crystal structures of these small molecules bound to dimeric human STINGLBD established that a pair of agonists bound within the cGAMP pocket for the SR-717 complex117, the AZBI complex118 and the MSA-2 complex119 (Fig. 7i). More potent activation was observed following covalent linkage of monomers. A novel approach applied a kinase-cGAS cascade to synthesize cyclic nucleotide analog MK-1454 (Fig. 7j), a therapeutic STINGLBD activator120, with the structure of its monomeric complex shown in Figure 7k.
a,b, Chemical formulas of the inhibitors CMA (a) and DMXAA (b). c,d, Crystal structure of DMXAA bound to mouse STINGLBD, at 2.4-Å resolution (c; PDB: 4LOL) There are two DMXAAs (space-filling representation) bound per mouse STINGLBD dimer (ribbon representation). Intermolecular contacts between the pair of DMXAAs and amino acid side chains in the binding pocket of mouse STINGLBD dimer (d). e, Chemical formula of covalent binder C-178. f, Chemical formula of SR-717. g, Chemical formula of AZBI. h,i, Chemical formula of MSA-2 (h) and positioning of a pair of MSA-2s in the cGAMP-binding pocket (i; PDB: 6UKM). j,k, Chemical formulas of MK-1454 (j) and positioning of MK-1454 in the cGAMP-binding pocket (k; PDB: 7MHC). The β-sheet lid segment that caps the ligand-binding pocket is shown in a lighter representation in order to allow clearer observation of the bound drug molecules.
Full-length human STING in the apo and 2′3-cGAMP-bound states
Full-length STING (STINGFL) is composed of a four-transmembrane helical component at its N terminus, a central cytoplasmic cGAMP-binding domain and a C-terminal signaling domain (Fig. 6a). Cryo-EM structures have been reported for dimeric chicken STINGFL in both the apo and 2′3′-cGAMP-bound states121. The transmembrane (TM) helices TM2 and TM4 from both subunits form an inner layer and are surrounded by TM1 and TM3 from both subunits that form an outer layer. The ligand-binding (LBD) and transmembrane (TM) domains of dimeric STINGFL in the apo state adopt an integrated and domain-swapped assembly, linked by two connector helices that form a right-handed crossover alignment (Fig. 8a and insert). A pronounced conformational transition was observed on 2′3′-cGAMP formation that involved a 180° half-turn rotation of the LBD relative to the TBD (Fig. 8b and insert), resulting in unwinding of the transmembrane-cytoplasmic connector segment, facilitating a switch from a crossover junctional alignment in the apo state (insert, Fig. 8a) to a parallel junctional alignment in the cGAMP-bound state (insert, Fig. 8b) of dimeric STINGFL.
a, Cryo-EM structure of chicken STINGFL in the apo state at 4.0-Å resolution (PDB: 6NT6). Individual monomers shown are as sticks and are colored yellow and magenta. The insert shows the cross-over of the LBD and TM domains between monomers. b, Cryo-EM structure of chicken STINGFL bound to 2′3′-cGAMP at 4.0-Å resolution (PDB: 6NT7). The bound 2′3′-cGAMP is shown in a space-filling representation. The insert shows the parallel alignment of the LBD and TM domains between monomers. c, Chemical formula of C53, a small molecule that targets the TM domains of STINGFL. d, Cryo-EM structure of human STINGFL bound to both 2′3′-cGAMP in the LBD pocket and C53 in the TM pocket, at 3.5-Å resolution (PDB: 7SII). e, Encapsulation of C53 by the helical bundle of TM helices in the structure of C53- and 2′3′-cGAMP-bound state of chicken STINGFL. f, Conformational change in TM bundle of chicken STINGFL on proceeding from the 2′3′-cGAMP state (in silver) to the C53- and 2′3′-cGAMP-bound state (in color). g,h, Model of 2′3′-cGAMP and C53-bound dimer-of-dimers of hSTINGFL (g) and contacts between dimers of the complex (h, a zoomed-in view of the circle in g).
Oligomerization of human STINGFL
Cryo-EM studies have further established that side-by-side packing of 2′3′-cGAMP-bound STINGFL dimers arranged in a linear manner results in tetramer and higher-order oligomer formation121. Such a higher-order alignment is mediated by a loop on the side of the LBD domain adopting a less extended conformation on 2′3′-cGAMP complex formation, thereby facilitating STINGFL oligomerization. Mutational analysis supports a model wherein oligomerization of cGAMP-bound STINGFL dimers, with accompanying translocation of 2′3′-cGAMP-STINGFL from the ER (endoplasmic reticulum) to the Golgi, are both critical events for STING activation. Indeed, high-order oligomerization of both 2′3′-cGAMP-bound STINGFL and TBK1 are critical for the phosphorylation of both the C-terminal tail of STING and transcription factor IRF3, modifications required for induction of type I interferons and cytokines122. It has also been proposed that oligomerization of 2′3′-cGAMP-bound STINGFL is facilitated by C-terminal tail release on complex formation, resulting in the exposure of an oligomerization interface, with the oligomeric state stabilized through disulfide crosslinking involving Cys148 of the connector helix123,124. Notably, alterations in the linker region surrounding Cys148, encoded by three point mutations, are STINGFL hyperactive and cause severe autoimmune syndrome SAVI (STING-associated vasculopathy with onset in infancy), potentially through a STINGFL polymer stabilization mechanism124.
It has also been proposed that palmitoylation of cysteines Cys88 and Cys91, adjacent to the oligomerization interface, could contribute to both transport and oligomerization of STINGFL (ref. 125). Similarly, STING oligomerization and TBK1 activation are boosted on binding linear sulfated acidic polysaccharides (sGAGs) synthesized in Golgi103.
STINGFL oligomer formation required for transport and pathway activation
The C-terminal tail (CTT) domain of STINGFL (Fig. 6a) is sequestered in the apo state and released on binding of second messenger, as the CTT domain is required for TBK1 and IRF3 activation122,126,127. A cryo-EM structure of cGAMP-bound dimeric chicken STINGFL in complex with dimeric human TBK1, established insertion of an induced β-strand of the CTT of STING into a groove between the kinase domain of one TBK1 subunit and the SDD (scaffold and dimerization domain) of the second subunit122. Given that the Ser366 phosphorylation site on the CTT domain of STING is far from the active site of TBK1, a model involving oligomerization of dimers of both proteins was proposed to facilitate trans phosphorylation of Ser366. The phosphorylated Ser within the pLXIS motif of the CTT of STING constitutes an IRF3 binding motif, thereby in turn recruiting IRF3 for phosphorylation by TBK1. IRF3 then dimerizes and translocates to the nucleus, where it induces the expression of type I interferons to restrict viral replication and spread. Notably, a recent study discovered ER-resident STING forms spherical ER membranous puzzle-like structures of biocondensate to prevent innate immunity from overactivation, thereby providing an additional example of the phase separation-mediated regulation of cGAS-STING immune signaling128.
Small molecule targeted to hSTINGFL
Recent efforts to discover additional potent small-molecule agonists of human STINGFL have yielded C53 (compound 53) (Fig. 8c), which exhibits robust on-target functional activation129. The molecular basis underlying C53 recognition by human STINGFL has emerged from a cryo-EM structure determination of both 2′3′-cGAMP- and C53-bound simultaneously to a dimer-of-dimers (tetramer) of hSTINGFL (ref. 130). The 2′3′-cGAMP and C53-bound STINGFL complex retains the parallel alignment linking the LBD and TM domains (Fig. 8d). C53, a relatively hydrophobic small molecule, adopts a C-shaped conformation and inserts into a deep pocket formed by transmembrane helices from both subunits of the STINGFL dimer (Fig. 8e). There is an outward shift of transmembrane helices of the STINGFL dimer to accommodate the bound C53 (Fig. 8f), which in turn induces interdimer contacts between TM helices of adjacent dimers, with such side-by-side packing resulting in higher-order oligomer formation.
In the presence of both cGAMP and C53, human STINGFL dimer formed both tetramers (Fig. 8g) and higher oligomeric states, which adopt a curved shape reflective of the positive membrane curvature associated with the rim of the Golgi and ER, thereby facilitating anterograde transport130. Current efforts aim to identify a cellular endogenous ligand that like, C53, could bind within the TMD pocket and facilitate cGAMP-induced STINGFL activation.
Highlights of recent STING research
Overall, the above sections highlight the conformational changes in both STINGLBD and STINGFL on cGAMP complex formation, on the contribution of STINGFL oligomerization towards its activation and transport from the ER to the Golgi, as well as identification of small molecules targeting both STING’s cytoplasmic and membrane-spanning domains to regulate its activity.
Future challenges
The most important take-home message is that CDN second messengers are ancient signaling molecules eliciting a potent innate immune response that originated in bacteria and evolved through evolutionary adaptation to metazoans. Structural studies have established that the central components of both bacterial and metazoan signaling pathways adopt a common set of principles enabling signal activation at the nucleotidyltransferase-sensor, CDN-second-messenger and STING-adaptor levels. Thus, metazoan-specific insertions within the core STING scaffold facilitate a switch from direct effector function in bacteria to regulation of antiviral transcription in metazoans.
Ongoing challenges in the immunity and inflammation fields require an improved understanding from a structural perspective of factors involved in the tightly regulated cGAMP-mediated cGAS–STING pathway, including formation, stability, trafficking and degradation of key components. Thus, further studies are required to molecularly elucidate how post-translational modifications regulate cGAS and STING activity, and how viral and bacterial factors are involved in pathogen evasion of this innate immune pathway. Furthermore, developing small molecules targeting cGAS or STING agonists as therapeutics for cancer immunotherapy, antagonists for treatment of autoimmune diseases, such as systemic lupus erythematosus and Sjogren’s syndrome, and adjuvant inducers for vaccine development against pathogens, is sure to be the focus of future studies. A remaining challenge relates to improving the understanding of how the cGAS–STING pathway distinguishes invading foreign from abundant host DNA, as well as how this pathway cross-talks with other pathways.
References
Athukoralage, J. S. & White, M. F. Cyclic nucleotide signaling in phage defense and counter-defense. Annu. Rev. Virol. 9, 451–468 (2022).
Zaver, S. A. & Woodward, J. J. Cyclic dinucleotides at the forefront of innate immunity. Curr. Opin. Cell Biol. 63, 49–56 (2020).
Patel, D. J., Yu, Y. & Jia, N. Bacterial origins of cyclic nucleotide-activated antiviral immune signaling. Mol. Cell 82, 4591–4610 (2022).
Chen, Q., Sun, L. & Chen, Z. J. Regulation and function of the cGAS–STING pathway of cytosolic DNA sensing. Nat. Immunol. 17, 1142–1149 (2016).
Cai, X., Chiu, Y. H. & Chen, Z. J. The cGAS–cGAMP–STING pathway of cytosolic DNA sensing and signaling. Mol. Cell 54, 289–296 (2014).
Hopfner, K.-P. P. & Hornung, V. Molecular mechanisms and cellular functions of cGAS–STING signalling. Nat. Rev. Mol. Cell Biol. 21, 501–521 (2020).
Li, T. & Chen, Z. J. The cGAS–cGAMP–STING pathway connects DNA damage to inflammation, senescence, and cancer. J. Exp. Med. 215, 1287–1299 (2018).
Zhang, X., Bai, X. & Chen, Z. J. Structures and mechanisms in the cGAS–STING innate immune pathway. Immunity 53, 43–53 (2020).
Fang, R., Jiang, Q., Yu, X., Zhao, Z. & Jiang, Z. Recent advances in the activation and regulation of the cGAS–STING pathway. Adv. Immunol. 156, 55–102 (2022).
Li, X.-D. et al. Pivotal roles of cGAS–cGAMP signaling in antiviral defense and immune adjuvant effects. Science 341, 1390–1394 (2013).
Gui, X. et al. Autophagy induction via STING trafficking is a primordial function of the cGAS pathway. Nature 567, 262–266 (2019).
Yang, H., Wang, H., Ren, J., Chen, Q. & Chen, Z. J. cGAS is essential for cellular senescence. Proc. Natl Acad. Sci. USA 114, E4612–E4620 (2017).
Gao, D. et al. Activation of cyclic GMP-AMP synthase by self-DNA causes autoimmune diseases. Proc. Natl Acad. Sci. USA 112, E5699–E5705 (2015).
Hu, M.-M. & Shu, H.-B. Innate immune response to cytoplasmic DNA: mechanisms and diseases. Annu. Rev. Immunol. 38, 79–98 (2020).
Decout, A., Katz, J. D., Venkatraman, S. & Ablasser, A. The cGAS–STING pathway as a therapeutic target in inflammatory diseases. Nat. Rev. Immunol. 21, 548–569 (2021).
Gao, D. et al. Cyclic GMP-AMP synthase is an innate immune sensor of HIV and other retroviruses. Science 341, 903–906 (2013).
Wang, H. et al. cGAS is essential for the antitumor effect of immune checkpoint blockade. Proc. Natl Acad. Sci. USA 114, 1637–1642 (2017).
Ablasser, A. & Chen, Z. J. cGAS in action: expanding roles in immunity and inflammation. Science 363, eaat8657 (2019).
Motwani, M., Pesiridis, S. & Fitzgerald, K. A. DNA sensing by the cGAS–STING pathway in health and disease. Nat. Rev. Genet. 20, 657–674 (2019).
Ablasser, A. & Hur, S. Regulation of cGAS-and RLR-mediated immunity to nucleic acids. Nat. Immunol. 21, 17–29 (2020).
Ablasser, A. et al. Cell intrinsic immunity spreads to bystander cells via the intercellular transfer of cGAMP. Nature 503, 530–534 (2013).
Gentili, M. et al. Transmission of innate immune signaling by packaging of cGAMP in viral particles. Science 349, 1232–1236 (2015).
Liu, H. et al. Nuclear cGAS suppresses DNA repair and promotes tumorigenesis. Nature 563, 131–136 (2018).
Jiang, H. et al. Chromatin-bound cGAS is an inhibitor of DNA repair and hence accelerates genome destabilization and cell death. EMBO J. 38, e102718 (2019).
Danilchanka, O. & Mekalanos, J. J. Cyclic dinucleotides and the innate immune response. Cell 154, 962–970 (2013).
Hornung, V., Hartmann, R., Ablasser, A. & Hopfner, K.-P. OAS proteins and cGAS: unifying concepts in sensing and responding to cytosolic nucleic acids. Nat. Rev. Immunol. 14, 521–528 (2014).
Eaglesham, J. B. & Kranzusch, P. J. Conserved strategies for pathogen evasion of cGAS–STING immunity. Curr. Opin. Immunol. 66, 27–34 (2020).
Kranzusch, P. J. cGAS and CD-NTase enzymes: structure, mechanism, and evolution. Curr. Opin. Struct. Biol. 59, 178–187 (2019).
Gao, P. et al. Cyclic [G(2′,5′)pA(3′,5′)p] is the metazoan second messenger produced by DNA-activated cyclic GMP-AMP synthase. Cell 153, 1094–1107 (2013).
Kato, K. et al. Cyclic GMP-AMP as an endogenous second messenger in innate immune signaling by cytosolic DNA. Annu. Rev. Biochem. 339, 826–830 (2017).
Davies, B. W., Bogard, R. W., Young, T. S. & Mekalanos, J. J. Coordinated regulation of accessory genetic elements produces cyclic di-nucleotides for V. cholerae virulence. Cell 149, 358–370 (2012).
Sun, L. et al. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Wu, J. et al. Cyclic GMP-AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339, 826–830 (2013).
Diner, E. J. J. et al. The innate immune DNA sensor cGAS produces a noncanonical cyclic dinucleotide that activates human STING. Cell Rep. 3, 1355–1361 (2013).
Ablasser, A. et al. cGAS produces a 2′-5′-linked cyclic dinucleotide second messenger that activates STING. Nature 498, 380–384 (2013).
Zhang, X. et al. Cyclic GMP-AMP containing mixed phosphodiester linkages is an endogenous high-affinity ligand for STING. Mol. Cell 51, 226–235 (2013).
Civril, F. et al. Structural mechanism of cytosolic DNA sensing by cGAS. Nature 498, 332–337 (2013).
Li, X. et al. Cyclic GMP-AMP synthase is activated by double-stranded DNA-induced oligomerization. Immunity 39, 1019–1031 (2013).
Zhang, X. et al. The cytosolic DNA sensor cGAS forms an oligomeric complex with DNA and undergoes switch-like conformational changes in the activation loop. Cell Rep. 6, 421–430 (2014).
Andreeva, L. et al. cGAS senses long and HMGB/TFAM-bound U-turn DNA by forming protein–DNA ladders. Nature 549, 394–398 (2017).
Xie, W. et al. Human cGAS catalytic domain has an additional DNA-binding interface that enhances enzymatic activity and liquid-phase condensation. Proc. Natl Acad. Sci. USA 116, 11946–11955 (2019).
Donovan, J., Dufner, M. & Korennykh, A. Structural basis for cytosolic double-stranded RNA surveillance by human oligoadenylate synthetase 1. Proc. Natl Acad. Sci. USA 110, 1652–1657 (2013).
Lohöfener, J. et al. The activation mechanism of 2′-5′-oligoadenylate synthetase gives new insights into OAS/cGAS triggers of innate immunity. Structure 23, 851–862 (2015).
Zhou, W. et al. Structure of the human cGAS–DNA complex reveals enhanced control of immune surveillance. Cell 174, 300–311 (2018).
Zhao, Z. et al. Mn2+ directly activates cGAS and structural analysis suggests Mn2+ induces a noncanonical catalytic synthesis of 2′3′-cGAMP. Cell Rep. 32, 108053 (2020).
Kranzusch, P. J. et al. Structure-guided reprogramming of human cGAS dinucleotide linkage specificity. Cell 158, 1011–1021 (2014).
Ding, C., Song, Z., Shen, A., Chen, T. & Zhang, A. Small molecules targeting the innate immune cGAS‒STING‒TBK1 signaling pathway. Acta Pharm. Sin. B 10, 2272–2298 (2020).
Hertzog, J. & Rehwinkel, J. Regulation and inhibition of the DNA sensor cGAS. EMBO Rep. 21, e51345 (2020).
Tian, X. et al. Medicinal chemistry perspective on cGAS–STING signaling pathway with small molecule inhibitors. Eur. J. Med. Chem. 244, 114791 (2022).
Vincent, J. et al. Small molecule inhibition of cGAS reduces interferon expression in primary macrophages from autoimmune mice. Nat. Commun. 8, 750 (2017).
Lama, L. et al. Development of human cGAS-specific small-molecule inhibitors for repression of dsDNA-triggered interferon expression. Nat. Commun. 10, 2261 (2019).
Hall, J. et al. Discovery of PF-06928215 as a high affinity inhibitor of cGAS enabled by a novel fluorescence polarization assay. PLoS ONE 12, e0184843 (2017).
Hall, J. et al. The catalytic mechanism of cyclic GMP-AMP synthase (cGAS) and implications for innate immunity and inhibition. Protein Sci. 26, 2367–2380 (2017).
Padilla-Salinas, R. et al. Discovery of small-molecule cyclic GMP-AMP synthase inhibitors. J. Org. Chem. 85, 1579–1600 (2020).
Wu, Y. & Li, S. Role of post-translational modifications of cGAS in innate immunity. Int. J. Mol. Sci. 21, 7842 (2020).
Liu, J. et al. The cGAS–STING pathway: post-translational modifications and functional implications in diseases. Cytokine Growth Factor Rev. 68, 69–80 (2022).
Yu, Y. et al. Post-translational modifications of cGAS–STING: a critical switch for immune regulation. Cells 11, 3043 (2022).
Zhang, J. et al. Species-specific deamidation of cGAS by herpes simplex virus UL37 protein facilitates viral replication. Cell Host Microbe 24, 234–248 (2018).
He, S. et al. Viral pseudo-enzymes activate RIG-I via deamidation to evade cytokine production. Mol. Cell 58, 134–146 (2015).
Meade, N. et al. Poxviruses evade cytosolic sensing through disruption of an mTORC1–mTORC2 regulatory circuit. Cell 174, 1143–1157.e17 (2018).
Huang, Z.-F. et al. Human cytomegalovirus protein UL31 inhibits DNA sensing of cGAS to mediate immune evasion. Cell Host Microbe 24, 69–80 (2018).
Zheng, Y. et al. Zika virus elicits inflammation to evade antiviral response by cleaving cGAS via NS1–caspase-1 axis. EMBO J. 37, e99347 (2018).
Du, M. & Chen, Z. J. DNA-induced liquid phase condensation of cGAS activates innate immune signaling. Science 361, 704–709 (2018).
Guey, B. & Ablasser, A. Emerging dimensions of cellular cGAS–STING signaling. Curr. Opin. Immunol. 74, 164–171 (2022).
Guey, B. et al. BAF restricts cGAS on nuclear DNA to prevent innate immune activation. Science 369, 823–828 (2020).
Zhou, W., Mohr, L., Maciejowski, J. & Kranzusch, P. J. cGAS phase separation inhibits TREX1-mediated DNA degradation and enhances cytosolic DNA sensing. Mol. Cell 81, 739–755 (2021).
Li, T. et al. Phosphorylation and chromatin tethering prevent cGAS activation during mitosis. Science 371, abc5386 (2021).
Xu, G. et al. Viral tegument proteins restrict cGAS–DNA phase separation to mediate immune evasion. Mol. Cell 81, 2823–2837 (2021).
Bhowmik, D. et al. Cooperative DNA binding mediated by KicGAS/ORF52 oligomerization allows inhibition of DNA-induced phase separation and activation of cGAS. Nucleic Acids Res. 49, 9389–9403 (2021).
Bhowmik, D., Tian, Y., Wang, B., Zhu, F. & Yin, Q. Structural basis of higher order oligomerization of KSHV inhibitor of cGAS. Proc. Natl Acad. Sci. USA 119, e2200285119 (2022).
Volkman, H. E., Cambier, S., Gray, E. E. & Stetson, D. B. Tight nuclear tethering of cGAS is essential for preventing autoreactivity. eLife 8, e47491 (2019).
Zierhut, C. et al. The cytoplasmic DNA sensor cGAS promotes mitotic cell death. Cell 178, 302–315 (2019).
Boyer, J. A. et al. Structural basis of nucleosome-dependent cGAS inhibition. Science 370, 450–454 (2020).
Michalski, S. et al. Structural basis for sequestration and autoinhibition of cGAS by chromatin. Nature 587, 678–682 (2020).
Zhao, B. et al. The molecular basis of tight nuclear tethering and inactivation of cGAS. Nature 587, 673–677 (2020).
Cao, D., Han, X., Fan, X., Xu, R.-M. & Zhang, X. Structural basis for nucleosome-mediated inhibition of cGAS activity. Cell Res. 30, 1088–1097 (2020).
Kujirai, T. et al. Structural basis for the inhibition of cGAS by nucleosomes. Science 370, 455–458 (2020).
Pathare, G. R. et al. Structural mechanism of cGAS inhibition by the nucleosome. Nature 587, 668–672 (2020).
Zhou, K., Gaullier, G. & Luger, K. Nucleosome structure and dynamics are coming of age. Nat. Struct. Mol. Biol. 26, 3–13 (2019).
Slavik, K. M. et al. cGAS-like receptors sense RNA and control 3′2′-cGAMP signalling in Drosophila. Nature 597, 109–113 (2021).
Holleufer, A. et al. Two cGAS-like receptors induce antiviral immunity in Drosophila. Nature 597, 114–118 (2021).
Shi, H., Wu, J., Chen, Z. J. & Chen, C. Molecular basis for the specific recognition of the metazoan cyclic GMP-AMP by the innate immune adaptor protein STING. Proc. Natl Acad. Sci. USA 112, 8947–8952 (2015).
Li, L. et al. Hydrolysis of 2′3′-cGAMP by ENPP1 and design of nonhydrolyzable analogs. Nat. Chem. Biol. 10, 1043–1048 (2014).
Kato, K. et al. Crystal structure of Enpp1, an extracellular glycoprotein involved in bone mineralization and insulin signaling. Proc. Natl Acad. Sci. USA 109, 16876–16881 (2012).
Kato, K. et al. Structural insights into cGAMP degradation by Ecto-nucleotide pyrophosphatase phosphodiesterase 1. Nat. Commun. 9, 4424 (2018).
Dennis, M. L. et al. Crystal structures of human ENPP1 in apo and bound forms. Acta Crystallogr. Sect. D. Struct. Biol. 76, 889–898 (2020).
Gao, J. et al. Identification and characterization of phosphodiesterases that specifically degrade 3′3′-cyclic GMP-AMP. Cell Res. 25, 539–550 (2015).
Hobbs, S. J. et al. Phage anti-CBASS and anti-Pycsar nucleases subvert bacterial immunity. Nature 605, 522–526 (2022).
Eaglesham, J. B., Pan, Y., Kupper, T. S. & Kranzusch, P. J. Viral and metazoan poxins are cGAMP-specific nucleases that restrict cGAS–STING signalling. Nature 566, 259–263 (2019).
Luteijn, R. D. et al. SLC19A1 transports immunoreactive cyclic dinucleotides. Nature 573, 434–438 (2019).
Ritchie, C., Cordova, A. F., Hess, G. T., Bassik, M. C. & Li, L. SLC19A1 is an importer of the immunotransmitter cGAMP. Mol. Cell 75, 372–381 (2019).
Zhang, Q. et al. Recognition of cyclic dinucleotides and folates by human SLC19A1. Nature 617, 170–176 (2022).
Wright, N. J. et al. Methotrexate recognition by the human reduced folate carrier SLC19A1. Nature 609, 1056–1062 (2022).
Parker, J. L. et al. Structural basis of antifolate recognition and transport by PCFT. Nature 595, 130–134 (2021).
Maltbaek, J. H., Cambier, S., Snyder, J. M. & Stetson, D. B. ABCC1 transporter exports the immunostimulatory cyclic dinucleotide cGAMP. Immunity 55, 1799–1812 (2022).
Zhong, B. et al. The adaptor protein MITA links virus-sensing receptors to IRF3 transcription factor activation. Immunity 29, 538–550 (2008).
Sun, W. et al. ERIS, an endoplasmic reticulum IFN stimulator, activates innate immune signaling through dimerization. Proc. Natl Acad. Sci. USA 106, 8653–8658 (2009).
Jin, L. et al. MPYS, a novel membrane tetraspanner, is associated with major histocompatibility complex class II and mediates transduction of apoptotic signals. Mol. Cell. Biol. 28, 5014–5026 (2008).
Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 674–678 (2008).
Burdette, D. L. et al. STING is a direct innate immune sensor of cyclic-di-GMP. Nature 478, 515–518 (2011).
Burdette, D. L. & Vance, R. E. STING and the innate immune response to nucleic acids in the cytosol. Nat. Immunol. 14, 19–26 (2013).
Barber, G. N. STING: infection, inflammation and cancer. Nat. Rev. Immunol. 15, 760–770 (2015).
Fang, R. et al. Golgi apparatus-synthesized sulfated glycosaminoglycans mediate polymerization and activation of the cGAMP sensor STING. Immunity 54, 962–975 (2021).
Huang, Y.-H., Liu, X.-Y., Du, X.-X., Jiang, Z.-F. & Su, X.-D. The structural basis for the sensing and binding of cyclic di-GMP by STING. Nat. Struct. Mol. Biol. 19, 728–730 (2012).
Ouyang, S. et al. Structural analysis of the STING adaptor protein reveals a hydrophobic dimer interface and mode of cyclic di-GMP binding. Immunity 36, 1073–1086 (2012).
Shang, G. et al. Crystal structures of STING protein reveal basis for recognition of cyclic di-GMP. Nat. Struct. Mol. Biol. 19, 725–727 (2012).
Shu, C., Yi, G., Watts, T., Kao, C. C. & Li, P. Structure of STING bound to cyclic di-GMP reveals the mechanism of cyclic dinucleotide recognition by the immune system. Nat. Struct. Mol. Biol. 19, 722–724 (2012).
Yin, Q. et al. Cyclic di-GMP sensing via the innate immune signaling protein STING. Mol. Cell 46, 735–745 (2012).
Gao, P. et al. Structure-function analysis of STING activation by c[G(2′,5′) pA(3′,5′)p] and targeting by antiviral DMXAA. Cell 154, 748–762 (2013).
Kranzusch, P. J. et al. Ancient origin of cGAS–STING reveals mechanism of universal 2′,3′ cGAMP signaling. Mol. Cell 59, 891–903 (2015).
Morehouse, B. R. et al. STING cyclic dinucleotide sensing originated in bacteria. Nature 586, 429–433 (2020).
Guerini, D. STING agonists/antagonists: their potential as therapeutics and future developments. Cells 11, 1159 (2022).
Cavlar, T., Deimling, T., Ablasser, A., Hopfner, K.-P. & Hornung, V. Species-specific detection of the antiviral small-molecule compound CMA by STING. EMBO J. 32, 1440–1450 (2013).
Conlon, J. et al. Mouse, but not human STING, binds and signals in response to the vascular disrupting agent 5,6-dimethylxanthenone-4-acetic acid. J. Immunol. 190, 5216–5225 (2013).
Gao, P. et al. Binding-pocket and lid-region substitutions render human STING sensitive to the species-specific drug DMXAA. Cell Rep. 8, 1668–1676 (2014).
Haag, S. M. et al. Targeting STING with covalent small-molecule inhibitors. Nature 559, 269–273 (2018).
Chin, E. N. et al. Antitumor activity of a systemic STING-activating non-nucleotide cGAMP mimetic. Science 369, 993–999 (2020).
Ramanjulu, J. M. et al. Design of amidobenzimidazole STING receptor agonists with systemic activity. Nature 564, 439–443 (2018).
Pan, B. S. et al. An orally available non-nucleotide STING agonist with antitumor activity. Science 369, eaba6098 (2020).
McIntosh, J. A. et al. A kinase-cGAS cascade to synthesize a therapeutic STING activator. Nature 603, 439–444 (2022).
Shang, G., Zhang, C., Chen, Z. J., Bai, X. & Zhang, X. Cryo-EM structures of STING reveal its mechanism of activation by cyclic GMP–AMP. Nature 567, 389–393 (2019).
Zhang, C. et al. Structural basis of STING binding with and phosphorylation by TBK1. Nature 567, 394–398 (2019).
Ergun, S. L., Fernandez, D., Weiss, T. M. & Li, L. STING polymer structure reveals mechanisms for activation, hyperactivation, and inhibition. Cell 178, 290–301 (2019).
Ergun, S. L. & Li, L. Structural insights into STING signaling. Trends Cell Biol. 30, 399–407 (2020).
Mukai, K. et al. Activation of STING requires palmitoylation at the Golgi. Nat. Commun. 7, 11932 (2016).
Zhao, B. et al. Structural basis for concerted recruitment and activation of IRF-3 by innate immune adaptor proteins. Proc. Natl Acad. Sci. USA 113, E3403–E3412 (2016).
Zhao, B. et al. A conserved PLPLRT/SD motif of STING mediates the recruitment and activation of TBK1. Nature 569, 718–722 (2019).
Yu, X. et al. The STING phase-separator suppresses innate immune signalling. Nat. Cell Biol. 23, 330–340 (2021).
Pryde, D. C. et al. The discovery of potent small molecule activators of human STING. Eur. J. Med. Chem. 209, 112869 (2021).
Lu, D. et al. Activation of STING by targeting a pocket in the transmembrane domain. Nature 604, 557–562 (2022).
Acknowledgements
D.J.P. was supported by funds from NIH (GM129430 and AI141507), the Maloris Foundation and Memorial Sloan-Kettering Cancer Center Core grant P30 CA008748. W.X. was supported by funds from the National Natural Science Foundation of China Grant 82151216. We thank P. Gao for providing coordinates of the structures of major CDN transporter SLC19A1 in the apo and cGAMP-bound forms prior to publication92.
Author information
Authors and Affiliations
Contributions
D.J.P. wrote the first draft of the Review, and Y.Y. and W.X. provided input to help complete it.
Corresponding author
Ethics declarations
Competing interests
D.J.P. is a paid consultant for Ventus Therapeutics. Y.Y. and W.X. declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Philip Kranzusch and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Carolina Perdigoto and Dimitris Typas, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Patel, D.J., Yu, Y. & Xie, W. cGAMP-activated cGAS–STING signaling: its bacterial origins and evolutionary adaptation by metazoans. Nat Struct Mol Biol 30, 245–260 (2023). https://doi.org/10.1038/s41594-023-00933-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-023-00933-9
This article is cited by
-
STING pathway as a cancer immunotherapy: Progress and challenges in activating anti-tumor immunity
Molecular Biology Reports (2024)