Abstract
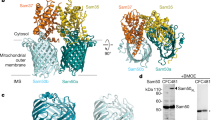
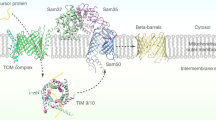
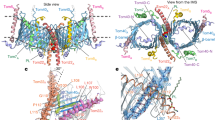
Mitochondrial β-barrel proteins are essential for the transport of metabolites, ions and proteins. The sorting and assembly machinery (SAM) mediates their folding and membrane insertion. We report the cryo-electron microscopy structure of the yeast SAM complex carrying an early eukaryotic β-barrel folding intermediate. The lateral gate of Sam50 is wide open and pairs with the last β-strand (β-signal) of the substrate—the 19-β-stranded Tom40 precursor—to form a hybrid barrel in the membrane plane. The Tom40 barrel grows and curves, guided by an extended bridge with Sam50. Tom40’s first β-segment (β1) penetrates into the nascent barrel, interacting with its inner wall. The Tom40 amino-terminal segment then displaces β1 to promote its pairing with Tom40’s last β-strand to complete barrel formation with the assistance of Sam37’s dynamic α-protrusion. Our study thus reveals a multipoint guidance mechanism for mitochondrial β-barrel folding.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The cryo-EM density map and atomic coordinates of the yeast SAMTom40-early complex have been deposited to the Electron Microscopy Data Bank and Protein Data Bank with accession numbers EMD-32019 and 7VKU, respectively. Source data are provided with this paper.
Change history
25 January 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41594-023-00926-8
References
Neupert, W. & Herrmann, J. M. Translocation of proteins into mitochondria. Ann. Rev. Biochem. 76, 723–749 (2007).
Walther, D. M. & Rapaport, D. Biogenesis of mitochondrial outer membrane proteins. Biochim. Biophys. Acta 1793, 42–51 (2009).
Walther, D. M., Rapaport, D. & Tommassen, J. Biogenesis of β-barrel membrane proteins in bacteria and eukaryotes: evolutionary conservation and divergence. Cell. Mol. Life Sci. 66, 2789–2804 (2009).
Hagan, C. L., Silhavy, T. J. & Kahne, D. β-Barrel membrane protein assembly by the Bam complex. Annu. Rev. Biochem. 80, 189–210 (2011).
Wu, R., Stephenso, R., Gichaba, A. & Noinaj, N. The big BAM theory: an open and closed case? Biochim. Biophys. Acta 1862, 183062 (2020).
Höhr, A. I. C., Straub, S. P., Warscheid, B., Becker, T. & Wiedemann, N. Assembly of β-barrel proteins in the mitochondrial outer membrane. Biochim. Biophys. Acta 1853, 75–88 (2015).
Selkrig, J. et al. Discovery of an archetypal protein transport system in bacterial outer membranes. Nat. Struct. Mol. Biol. 19, 506–510 (2012).
Paschen, S. A. et al. Evolutionary conservation of biogenesis of β-barrel membrane proteins. Nature 426, 862–866 (2003).
Wiedemann, N. et al. Machinery for protein sorting and assembly in the mitochondrial outer membrane. Nature 424, 565–571 (2003).
Gentle, I. et al. The Omp85 family of proteins is essential for outer membrane biogenesis in mitochondria and bacteria. J. Cell Biol. 164, 19–24 (2004).
Ishikawa, D., Yamamoto, H., Tamura, Y., Moritoh, K. & Endo, T. Two novel proteins in the mitochondrial outer membrane mediate β-barrel protein assembly. J. Cell Biol. 166, 621–627 (2004).
Waizenegger, T. et al. Tob38, a novel essential component in the biogenesis of β-barrel proteins of mitochondria. EMBO Rep. 5, 704–709 (2004).
Klein, A. et al. Characterization of the insertase for β-barrel proteins of the outer mitochondrial membrane. J. Cell Biol. 199, 599–611 (2012).
Yamano, K., Tanaka-Yamano, S. & Endo, T. Mdm10 as a dynamic constituent of the TOB/SAM complex directs coordinated assembly of Tom40. EMBO Rep. 11, 187–193 (2010).
Ellenrieder, L. et al. Separating mitochondrial protein assembly and endoplasmic reticulum tethering by selective coupling of Mdm10. Nat. Commun. 7, 13021 (2016).
Diederichs, K. A. et al. Structural insight into mitochondrial β-barrel outer membrane protein biogenesis. Nat. Commun. 11, 3290 (2020).
Takeda, H. et al. Mitochondrial sorting and assembly machinery operates by β-barrel switching. Nature 590, 163–169 (2021).
Wang, Q. et al. Structural insight into the SAM-mediated assembly of the mitochondrial TOM core complex. Science 373, 1377–1381 (2021).
Kutik, S. et al. Dissecting membrane insertion of mitochondrial β-barrel proteins. Cell 132, 1011–1024 (2008).
Becker, T. et al. Assembly of the mitochondrial protein import channel. Role of Tom5 in two-stage interaction of Tom40 with the SAM complex. Mol. Biol. Cell 21, 3106–3113 (2010).
Wenz, L.-S. et al. Sam37 is crucial for formation of the mitochondrial TOM–SAM supercomplex, thereby promoting β-barrel biogenesis. J. Cell Biol. 210, 1047–1054 (2015).
Stroud, D. A. et al. Biogenesis of mitochondrial β-barrel proteins: the POTRA domain is involved in precursor release from the SAM complex. Mol. Biol. Cell 22, 2823–2832 (2011).
Höhr, A. I. C. et al. Membrane protein insertion through a mitochondrial β-barrel gate. Science 359, 6834 (2018).
Araiso, Y. et al. Structure of the mitochondrial import gate reveals distinct preprotein paths. Nature 575, 395–401 (2019).
Tucker, K. & Park, E. Cryo-EM structure of the mitochondrial protein-import channel TOM complex at near-atomic resolution. Nat. Struct. Mol. Biol. 26, 1158–1166 (2019).
Qiu, J. et al. Coupling of mitochondrial import and export translocases by receptor-mediated supercomplex formation. Cell 154, 596–608 (2013).
Arnold, T., Poynor, M., Nussberger, S., Lupas, A. N. & Linke, D. Gene duplication of the eight-stranded β-barrel OmpX produces a functional pore: a scenario for the evolution of transmembrane β-barrels. J. Mol. Biol. 366, 1174–1184 (2007).
Remmert, M., Biegert, A., Linke, D., Lupas, A. N. & Söding, J. Evolution of outer membrane β-barrels from an ancestral ββ hairpin. Mol. Biol. Evol. 27, 1348–1358 (2010).
Jores, T. et al. Characterization of the targeting signal in mitochondrial β-barrel proteins. Nat. Commun. 7, 12036 (2016).
Jores, T. et al. Cytosolic Hsp70 and Hsp40 chaperones enable the biogenesis of mitochondrial β-barrel proteins. J. Cell Biol. 217, 3091–3108 (2018).
Chan, N. C. & Lithgow, T. The peripheral membrane subunits of the SAM complex function codependently in mitochondrial outer membrane biogenesis. Mol. Biol. Cell 19, 126–136 (2008).
Doyle, M. T. & Barnstein, H. D. Bacterial outer membrane proteins assemble via asymmetric interactions with the BamA β-barrel. Nat. Commun. 10, 3358 (2019).
Tomasek, D. et al. Structure of a nascent membrane protein as it folds on the BAM complex. Nature 583, 473–478 (2020).
Wu, R. et al. Plasticity within the barrel domain of BamA mediates a hybrid-barrel mechanism by BAM. Nat. Commun. 12, 7131 (2021).
Xiao, L. et al. Structures of the β-barrel assembly machine recognizing outer membrane protein substrates. FASEB J. 35, e21207 (2021).
Doyle, M. T., Jimah, J. R., Hinshaw, J. E. & Bernstein, H. D. Cryo-EM structures reveal multiple stages of bacterial outer membrane protein folding. Cell 185, 1143–1156 (2022).
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004).
Mastronarde, D. N. et al. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr. D 74, 814–840 (2018).
Kim, D. N., Gront, D. & Sanbonmatsu, K. Y. Practical considerations for atomic structure modeling with cryo-EM maps. J. Chem. Inf. Model. 60, 2436–2442 (2020).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Tama, F. et al. Normal mode based flexible fitting of high-resolution structure into low-resolution experimental data from cryo-EM. J. Struct. Biol. 147, 315–326 (2004).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinformatics 54, 5.6.1–5.6.37 (2016).
Wu, E. L. et al. CHARMM-GUI membrane builder toward realistic biological membrane simulations. J. Comput. Chem. 35, 1997–2004 (2014).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Jorgensen, W. L. et al. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Roe, D. R. & Brooks, B. R. A protocol for preparing explicitly solvated systems for stable molecular dynamics simulations. J. Chem. Phys. 153, 054123 (2020).
Balusek, C. et al. Gumbart, accelerating membrane simulations with hydrogen mass repartitioning. J. Chem. Theory Comput. 15, 4673–4686 (2019).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577 (1995).
McGibbon, R. T. et al. MDTraj: a modern open library for the analysis of molecular dynamics trajectories. Biophys. J. 109, 1528–1532 (2015).
Sastry, G. M. et al. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 27, 221–234 (2013).
Li, J. et al. The VSGB 2.0 model: a next generation energy model for high resolution protein structure modeling. Proteins 79, 2794–2812 (2011).
Lomize, M. A. et al. OPM database and PPM web server: resources for positioning of proteins in membranes. Nucleic Acids Res. 40, D370–D376 (2012).
Harder, E. et al. OPLS3: a force field providing broad coverage of drug-like small molecules and proteins. J. Chem. Theory Comput. 12, 281–296 (2016).
Kabsch, W., & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Struyvé, M., Moons, M. & Tommassen, J. Carboxy-terminal phenylalanine is essential for the correct assembly of a bacterial outer membrane protein. J. Mol. Biol. 218, 141–148 (1991).
Acknowledgements
We thank the members of the T.E. and N.P./N.W. laboratories for discussion and critical comments on the manuscript. This work was supported by JSPS KAKENHI to T.E. (15H05705 and 2222703), K.I. (16K21680, 18K11543 and 21H03551) and H.T. (18K14640), by a JST CREST grant to T.E. (JPMJCR12M1) and an AMED CREST grant to T.E. (21gm1410002h0002), by Deutsche Forschungsgemeinschaft (German Research Foundation) grants to T.B. (BE 4679/2-2; SFB1218 project ID 269925409), N.P. (PF 202/9-1; project ID 394024777) and N.W. (WI 4506/1-1 (project ID 406757425) and SFB 1381 (project ID 403222702)), Germany’s Excellence Strategy to N.P. and N.W. (CIBSS-EXC2189; Project ID 390939984) and European Research Council Consolidator grant number 648235 to N.W. The following grants are also acknowledged: a grant from the Takeda Science Foundation (to T.E.) and grants from the Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research) from AMED under grant numbers JP19am01011115 (to M.K.), JP21am0101114 (to K.I., C.M. and T.H.) and JP21am0101110 (support number 1976; to K.T). Technical assistance by J. Suzuki (T.E. laboratory) is acknowledged. H.T. was supported by a Research Fellowship for Young Scientists from the Japan Society for the Promotion of Science (18J00358) and is grateful to M. Shirouzu (RIKEN Institute) for the encouragement to H.T. I.G. was supported by an Alexander von Humboldt Foundation Research Fellowship.
Author information
Authors and Affiliations
Contributions
T.E. designed the research and wrote the paper. H.T. performed most of the experiments and wrote the paper. J.V.B., C.L., I.G. and L.-S.W. characterized yeast mutants and studied the organization of the SAM complexes and their interaction with precursor proteins. A.T. performed the cryo-EM measurements and data processing, including single-particle analyses, with supervision from M.K. Y.Y. performed the molecular dynamics simulation and molecular dynamics-assisted model building, with supervision from K.T. N.P., N.W. and T.B. designed and wrote part of the paper. K.I., T.H. and C.M. performed the model building and calculation of the subunit interaction energy. All authors contributed to the analysis and discussion of the results of the experiments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Radhakrishnan Mahalakshmi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Carolina Perdigoto, in collaboration with the Nature Structural & Molecular Biology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Analysis and purification of the SAMTom40-early intermediate.
a, Wild-type (WT) mitochondria were incubated with radiolabeled WT Tom40 and Tom40G354A mutant precursor proteins (pulse), re-isolated and incubated to follow the assembly (chase) of the precursor proteins by blue native-PAGE and radioimaging. Lane 8 shows a higher contrast image of lane 7 to visualize the assembly into the mature TOM complex. b, Digitonin lysed mitochondria isolated from WT and Tom40G354A strains grown at 23 °C in YPG were analyzed by blue native-PAGE and immunoblotting against Tom22. c, Radiolabeled wild-type (WT) Tom40 and Tom40G354A precursor proteins were imported into isolated WT mitochondria and those with Sam50 lacking the POTRA domain (ΔPOTRA), Tom5 (Δ5), Tom6 (Δ6), Tom7 (Δ7), or Sam37 (Δ37) for 30 min at 25 °C, and solubilized complexes were analyzed by blue-native PAGE and radioimaging. TOM, mature TOM complex; SAM-Tom40, SAMTom40-early and/or SAMTom40-folded, SAM-Tom40-5-6, SAMTom40-5-6, and Tom40-5-6-7, the assembly intermediate containing Tom40, Tom5, Tom6, and Tom7. d, [35S]Tom40G354A precursor was imported into wild-type (WT) and sam37Δ mitochondria for the indicated periods. Where indicated, recombinant Sam37 was imported into sam37Δ mitochondria before [35S]Tom40G354A was imported. The samples were lysed with digitonin and analyzed by native electrophoresis and radioimaging. e, The elution profile of gel filtration of the purified SAMTom40-early intermediate in 0.02% GDN. f, g, SDS-PAGE (f) and blue-native PAGE (g) gels of the SAMTom40-early intermediate stained with Coomassie Brilliant Blue (CBB). Data are representative of two (panels a-c), four (panel d), or five times (f, g) independent experiments.
Extended Data Fig. 2. Cryo-EM structural determination of the SAMTom40-early intermediate.
a, Representative cryo-EM micrograph image of the SAMTom40-early intermediate. The image is a representative of over thousand images. b, Schematic workflows of the steps followed in data processing leading to the final structures of the SAMTom40-early intermediate. c, Cryo-EM maps of the SAMTom40-early intermediate for different classes (b) filtered by different local resolutions (6σ and 9σ). Local resolution filtered maps at a lower threshold level (1σ) are overlaid in grey. Particle numbers and overall map resolutions are shown on the left. d, Upper left, local map resolutions of the final structures (class 3), with colors according to the scales. Upper right, FSC curves between the EM density maps and the atomic models of the SAMTom40-early intermediate (class 3). Lower, FSC curves between the EM density maps and the atomic models for the class 2 and 5 structures.
Extended Data Fig. 3. Cryo-EM structural determination of the SAMTom40-early intermediate.
a, The cryo-EM density maps (contoured at 7.0σ) and the atomic models for indicated segments of Sam50 and Tom40 in the SAMTom40-early intermediate. The structures of Sam35 and Sam37 are essentially the same as those reported before17. b–d, Secondary structure diagrams of Tom40 and Sam50, forming a hybrid β-barrel in the SAMTom40-early intermediate (b), Sam35 (c), and Sam37 (d) are shown. The regions invisible or unresolved are indicated in grey.
Extended Data Fig. 4. Structural characterization of the Tom40-early folding intermediate on SAM.
a, Interactions between loop 1 (between β1 and β2) and the β14-17 region of Sam50 in the SAMdimer complex (PDB ID: 7BTW). b, Comparison of the structures of the Tom40 in the SAMTom40-early intermediate (red) and in the mature TOM complex (pink). c, HA-tagged cysteine-free (Cfree) Sam50 mutants with cysteine residues introduced at the indicated positions (β1 and β16) were expressed in yeast cells lacking endogenous Sam50. Mitochondria were isolated and treated with 1 mM CuSO4 (oxidant) or 1 mM DTT (reductant) and proteins were analyzed by non-reducing SDS-PAGE followed by immunoblotting with the indicated antibodies. Ox, oxidized form with disulfide-bond formation; Red, reduced form. d, Mitochondria treated with or without CuSO4 (oxidant) were solubilized with digitonin and mixed without (Mito) or with GST bound to glutathione-Sepharose beads (GST) or GST followed by the Tom40 β-signal peptide fused with GST bound to glutathione-Sepharose beads (GST-β). The GST domain contains the C-terminally attached cleavage site for 3 C protease. Then Sam50 bound to the Tom40 β-signal was eluted from the beads by 3 C protease treatment, and proteins were analyzed by non-reducing SDS-PAGE followed by immunoblotting with the anti-HA antibodies. e, Radiolabeled Por1β1-19,C276 (19 β-strands) and Por1β2-19,C276 (18 β-strands) were imported for 30 min at 25 °C into Sam50Cfree and Sam50C130 mitochondria. Subsequently, samples were oxidized with 4-DPS and analyzed on non-reducing SDS-PAGE and radioimaging. Arrowhead, disulfide-bond of Sam50 and Por1 precursor; circle, porin precursor. f–i, Cysteine-free Tom40 (CF) and the indicated double-cysteine mutants Tom40 mutants associated with the SAM complex and those integrated into the mature TOM complex were isolated by FLAG purification (g, i) and HA purification (f, h), respectively, as in Fig. 3a. Proteins were analyzed by SDS-PAGE followed by immunoblotting with the indicated antibodies. L, load (5%); E, elute (50%). Data are representative of three (panel c–i) independent experiments.
Extended Data Fig. 5 MD simulation of Sam50 and Tom40 in the SAMTom40-early intermediate.
a, Time-course changes of secondary structure for the N-terminal region of Tom40 in the SamTom40-early intermediate were calculated using DSSP63. One of the five trajectories is shown as an example. The leftmost part shows the secondary structure of the mature Tom40 (PDB ID: 6JNF). The red and blue colors indicate the residue in α-helix and β-sheet, respectively. The results showed that the residues around the β1-segment of Tom40 are stable as α-helix for a long period of time during the simulation. The tendency was similar for the other four trajectories. b, Using the trajectories after 100 ns, the root-mean-square fluctuation (RMSF) and the fractions of secondary structure were calculated for each trajectory. The RMSF of each residue of Tom40 (upper left) and Sam50 (upper right) of the trajectory corresponding to (a) was plotted. The fractions of secondary structure calculated by DSSP for the same trajectory of Tom40 (lower left) and Sam50 (lower right) were also plotted. The lower part of each panel indicates the secondary structure of mature Tom40 (PDB ID: 6JNF) and the secondary structure of the initial structure of Sam50. The red and blue colors indicate the residue in α-helix and β-sheet, respectively. c, Snapshots of the structures of the Sam50Tom40-early intermediate at 0 ns, 300 ns (as an example of one state), and 800 ns (as an example of the other state) for one of the trajectories. During the MD simulation, the initially separated Tom40 β5-7 segments develop to form a β-sheet. d, Comparison of the cryo-EM density maps and the model, for β1 and its flanking regions, derived by MD-assisted fitting. e, Stereo view of the cryo-EM density maps for β1 and the surrounding part of the hybrid barrel of Tom40 and Sam50.
Extended Data Fig. 6 EM density map of the folding intermediates of Tom40G354A,Δ2-92 on the SAM complex.
Cryo-EM density maps of SAM-Tom40(G354A) (upper) and SAM-Tom40(G354A,Δ2-92) (lower) intermediates contoured at 6.0s, 3.0s, 2.5s, and 2.0s above average are shown. Loop 4, not β1, is seen for SAM-Tom40(G354A,D2-92).
Extended Data Fig. 7 Modeling of Tom5 and Tom6 bound to the SAMTom40-early intermediate.
a, Structures of Tom5 and Tom6 bound to the SAMTom40-early intermediate (SAMTom40-5-6) modeled by fitting simulation analysis. b, c, Close-up views of Tom6 (b) and Tom5 (c) bound to the SAMTom40-early intermediate. d, Structure of the TOM complex (PDB ID: 6JNF). e, f, Close-up views of Tom6 (e) and Tom5 (f) in the TOM complex. g, Interaction energy of Tom40 with Tom5 and Tom6 in the TOM complex (shown by bars in black). Gray zones represent the residues of Tom40 that are not modeled in the EM structure of the SAMTom40-early intermediate. Purple and blue bars in the upper part of the panel indicate the residues interacting with Tom5 and Tom6, respectively. The estimated total subunit interaction free energy is −139.1 kcal/mol for the SAMTom40-early complex and −197.6 kcal/mol for the TOM complex.
Extended Data Fig. 8 Role of the α-protrusion of Sam37 in Tom40 folding on the SAM complex.
a, b, Behaviour of the variant precursor Tom40M94L (used in Fig. 5e). Radiolabeled Tom40 constructs were analyzed by SDS-PAGE and radioimaging (a). The variant precursor Tom40M94L (shown in lane 1) prevents the generation of a truncated Tom40 via translation initiation from the internal methionine M94. Radiolabelled wild-type (WT) Tom40 and Tom40M94L precursor proteins were imported into isolated WT mitochondria for the indicated periods (min) at 25 °C, and solubilized complexes were analysed by blue-native PAGE and radioimaging (b). TOM, mature TOM complex; SAM-Tom40, SAMTom40-early and/or SAMTom40-folded, SAM-Tom40-5-6, SAMTom40-5-6, and Tom40-5-6-7, the assembly intermediate containing Tom40, Tom5, Tom6, and Tom7. Tom40M94L assembled like WT Tom40. c–i, Analysis of the α-protrusion of Sam37. c, Growth of YPH499 wild type, Sam37Δ186-209 and sam37Δ yeast cells on fermentable (YPD) and non-fermentable (YPG) medium at 19 °C, 24 °C, 30 °C and 36 °C °C (Sam37Δ186-209 is chromosomally expressed and contains a GAG linker at the site of truncation). d, Steady-state protein levels of isolated mitochondria from YPH499 wild type and Sam37Δ186-209 cells, as analyzed by Tris/Tricine gel electrophoresis and immunoblotting. Mitochondria were isolated on non-fermentable (YPG) medium at 23 °C. e, f, Radiolabeled Tom22 (e) and Tom20 (f) precursor proteins were imported into isolated YPH499 wild type, Sam37Δ186-209, and sam37Δ mitochondria for the indicated periods, and the digitonin-solubilized complexes were analyzed by blue native PAGE and autoradiography. TOM, the TOM complex. g–i, Radiolabeled Tom5, Tom6, and Tom7 precursor proteins were imported into isolated YPH499 wild type and Sam37Δ186-209 mitochondria for the indicated periods at 25 °C, and the digitonin-solubilized complexes were analyzed by blue native PAGE and radioimaging. TOM, the TOM complex; TOM40-5-6-7, the assembly intermediate consisting of Tom40, Tom5, Tom6, and Tom7. Data are representative of two (panels a, b, d, g–i) or three (panels c, e, f) independent experiments.
Extended Data Fig. 9 Mitochondrial β-signal versus bacterial β-signal interactions with the β-barrel assembly machineries.
a, Alignment of mitochondrial and bacterial β-barrel protein sequences (left) containing the C-terminal β-signal strand (green lettering)19,26,33,36,64. The β-signal consensus of mitochondria and bacteria is highlighted19. Alignment of Sam50 and BamA sequences (right) containing β1 (underlined in blue). The consensus with the highly conserved glycine on the C-terminal side of β1 allows a direct comparison between bacteria and mitochondria. C.c., Caulobacter crescentus; C.g., Candida glabrata; D.m., Drosophila melanogaster; E.c., Escherichia coli; H.s., Homo sapiens; N.c., Neurospora crassa; N.m., Neisseria meningitidis; R.p., Rickettsia prowazekii; S.c. Saccharomyces cerevisiae; V.c., Vibrio cholerae. b, Cartoon representation of the hybrid interaction between the C-terminal β-signal (green lettering)-containing strand according to structural or crosslinking data (shaded green) of the β-barrel substrates with β1 of Sam50 or BamA, respectively (shaded blue). For the bacterial BAM machinery, the β-signal-strand of several substrates was found to associate with its N-terminl part with BamA β1 to form an asymmetric intermediate (register shift) creating a C-terminal overhang (shaded red) of the β-signal of the substrate (lower panel)33,35. It was proposed that β1 of these substrates can begin to form hydrogen bonds with the overhang of its own C-terminal β-signal-strand to start the barrel closure followed by sequential disruption of the hydrogen bonds between the β-signal-strand and BamA33. In contrast for the mitochondrial SAM machinery, the β-signal-strands of the Tom40 and Por1 substrates are specifically associated with Sam50 β1 without a register shift (upper panel)23, as also observed for the bacterial substrate EspP34,36. Thus, all analyzed mitochondrial β-signals (and one bacterial substrate) associate with Sam50/BamA β1 without a C-terminal overhang, implying a direct β-strand exchange mechanism for barrel closure and release23. c, Interactions between BamA (PDB ID: 6V05) and its substrate (second BamA) with its close-up view with C-terminal overhang. Hydrophilic and hydrophobic interactions are shown with brown dotted lines and black broken lines, respectively. Residues facing the barrel lumen and lipid phase are shown in black and gray, respectively.
Supplementary information
Supplementary Information
Supplementary Tables 1–3.
Supplementary Video 1
Five trajectories of molecular dynamics simulation of Tom40 in the SAMTom40-early intermediate. The five 1-μs trajectories of molecular dynamics simulations of the SAMTom40-early intermediate were packed into the continuous animation. The N-terminal region (residues 76–100) of Tom40 is highlighted in red. To fix the point of view, each trajectory was superimposed on the top view of the complex. Animation of the trajectories was done using VMD (Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graphics Modell. 14, 33–38 (1996)).
Source data
Source Data Fig. 1
Uncropped gels and plates.
Source Data Fig. 2
Uncropped gels and plates.
Source Data Fig. 3
Uncropped blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Uncropped blots.
Source Data Fig. 5
Uncropped gels.
Source Data Fig. 6
Uncropped gels.
Source Data Extended Data Fig. 1
Uncropped gels.
Source Data Extended Data Fig. 4
Uncropped gels and blots.
Source Data Extended Data Fig. 8
Uncropped gels, blots and plates.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Takeda, H., Busto, J.V., Lindau, C. et al. A multipoint guidance mechanism for β-barrel folding on the SAM complex. Nat Struct Mol Biol 30, 176–187 (2023). https://doi.org/10.1038/s41594-022-00897-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00897-2