Abstract
In bacterial defense and genome editing applications, the CRISPR-associated protein Cas9 searches millions of DNA base pairs to locate a 20-nucleotide, guide RNA-complementary target sequence that abuts a protospacer-adjacent motif (PAM). Target capture requires Cas9 to unwind DNA at candidate sequences using an unknown ATP-independent mechanism. Here we show that Cas9 sharply bends and undertwists DNA on PAM binding, thereby flipping DNA nucleotides out of the duplex and toward the guide RNA for sequence interrogation. Cryogenic-electron microscopy (cryo-EM) structures of Cas9–RNA–DNA complexes trapped at different states of the interrogation pathway, together with solution conformational probing, reveal that global protein rearrangement accompanies formation of an unstacked DNA hinge. Bend-induced base flipping explains how Cas9 ‘reads’ snippets of DNA to locate target sites within a vast excess of nontarget DNA, a process crucial to both bacterial antiviral immunity and genome editing. This mechanism establishes a physical solution to the problem of complementarity-guided DNA search and shows how interrogation speed and local DNA geometry may influence genome editing efficiency.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included within this paper and its supporting information files except for the cryo-EM data/models, which can be accessed as follows: Cas9–sgRNA–DNA (S. pyogenes) with 0 RNA–DNA base pairs, open-protein/linear-DNA conformation (Protein Data Bank (PDB) 7S3H, EMD-24823); Cas9–sgRNA–DNA (S. pyogenes) with 0 RNA–DNA base pairs, closed-protein/bent-DNA conformation (PDB 7S36, EMD-24817); Cas9–sgRNA (S. pyogenes) in the open-protein conformation (PDB 7S37, EMD-24818); Cas9–sgRNA–DNA (S. pyogenes) forming a 3-base-pair R-loop (PDB 7S38, EMD-24819). The analyses and figures in this study also draw on previously determined structures with PDB codes 4ZT0, 5FQ5, 5F9R, 4CMP, 1CGP and 4UN3 as described in the figure legends and the Supplementary Information. For figures containing fluorescence images, autoradiographs or scatter plots, the original data are available as source data files. Source data are provided with this paper.
References
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Pickar-Oliver, A. & Gersbach, C. A. The next generation of CRISPR-Cas technologies and applications. Nat. Rev. Mol. Cell Biol. 20, 490–507 (2019).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Mekler, V., Minakhin, L. & Severinov, K. Mechanism of duplex DNA destabilization by RNA-guided Cas9 nuclease during target interrogation. Proc. Natl Acad. Sci. USA 114, 5443–5448 (2017).
Sternberg, S. H., Redding, S., Jinek, M., Greene, E. C. & Doudna, J. A. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature 507, 62–67 (2014).
Jiang, F. & Doudna, J. A. CRISPR-Cas9 structures and mechanisms. Annu. Rev. Biophys. 46, 505–529 (2017).
Jinek, M. et al. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science 343, 1247997 (2014).
Jiang, F., Zhou, K., Ma, L., Gressel, S. & Doudna, J. A. A Cas9-guide RNA complex preorganized for target DNA recognition. Science 348, 1477–1481 (2015).
Jiang, F. et al. Structures of a CRISPR-Cas9 R-loop complex primed for DNA cleavage. Science 351, 867–871 (2016).
Globyte, V., Lee, S. H., Bae, T., Kim, J.-S. & Joo, C. CRISPR/Cas9 searches for a protospacer adjacent motif by lateral diffusion. EMBO J. 38, e99466 (2019).
Jones, D. L. et al. Kinetics of dCas9 target search in Escherichia coli. Science 357, 1420–1424 (2017).
Verdine, G. L. & Norman, D. P. G. Covalent trapping of protein-DNA complexes. Annu. Rev. Biochem. 72, 337–366 (2003).
Anders, C., Niewoehner, O., Duerst, A. & Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513, 569–573 (2014).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Osuka, S. et al. Real-time observation of flexible domain movements in CRISPR-Cas9. EMBO J. 37, e96941 (2018).
Bruner, S. D., Norman, D. P. & Verdine, G. L. Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403, 859–866 (2000).
Fromme, J. C. & Verdine, G. L. Structure of a trapped endonuclease III-DNA covalent intermediate. EMBO J. 22, 3461–3471 (2003).
Kim, J. L., Nikolov, D. B. & Burley, S. K. Co-crystal structure of TBP recognizing the minor groove of a TATA element. Nature 365, 520–527 (1993).
Kim, Y., Geiger, J. H., Hahn, S. & Sigler, P. B. Crystal structure of a yeast TBP/TATA-box complex. Nature 365, 512–520 (1993).
Love, J. J. et al. Structural basis for DNA bending by the architectural transcription factor LEF-1. Nature 376, 791–795 (1995).
Slupphaug, G. et al. A nucleotide-flipping mechanism from the structure of human uracil-DNA glycosylase bound to DNA. Nature 384, 87–92 (1996).
Vassylyev, D. G. et al. Atomic model of a pyrimidine dimer excision repair enzyme complexed with a DNA substrate: structural basis for damaged DNA recognition. Cell 83, 773–782 (1995).
Werner, M. H., Huth, J. R., Gronenborn, A. M. & Clore, G. M. Molecular basis of human 46X,Y sex reversal revealed from the three-dimensional solution structure of the human SRY-DNA complex. Cell 81, 705–714 (1995).
Kahn, J. D. & Crothers, D. M. Protein-induced bending and DNA cyclization. Proc. Natl Acad. Sci. USA 89, 6343–6347 (1992).
Koo, H. S., Drak, J., Rice, J. A. & Crothers, D. M. Determination of the extent of DNA bending by an adenine-thymine tract. Biochemistry 29, 4227–4234 (1990).
Schultz, S. C., Shields, G. C. & Steitz, T. A. Crystal structure of a CAP-DNA complex: the DNA is bent by 90 degrees. Science 253, 1001–1007 (1991).
Bui, C. T., Rees, K. & Cotton, R. G. H. Permanganate oxidation reactions of DNA: perspective in biological studies. Nucleosides Nucleotides Nucleic Acids 22, 1835–1855 (2003).
Cofsky, J. C. et al. CRISPR-Cas12a exploits R-loop asymmetry to form double-strand breaks. eLife 9, e55143 (2020).
Nakane, T., Kimanius, D., Lindahl, E. & Scheres, S. H. Characterisation of molecular motions in cryo-EM single-particle data by multi-body refinement in RELION. eLife 7, e36861 (2018).
Ramstein, J. & Lavery, R. Energetic coupling between DNA bending and base pair opening. Proc. Natl Acad. Sci. USA 85, 7231–7235 (1988).
Allan, B. W. et al. DNA bending by EcoRI DNA methyltransferase accelerates base flipping but compromises specificity. J. Biol. Chem. 274, 19269–19275 (1999).
Su, T.-J., Tock, M. R., Egelhaaf, S. U., Poon, W. C. K. & Dryden, D. T. F. DNA bending by M.EcoKI methyltransferase is coupled to nucleotide flipping. Nucleic Acids Res. 33, 3235–3244 (2005).
Blosser, T. R. et al. Two distinct DNA binding modes guide dual roles of a CRISPR-Cas protein complex. Mol. Cell 58, 60–70 (2015).
Hochstrasser, M. L., Taylor, D. W., Kornfeld, J. E., Nogales, E. & Doudna, J. A. DNA targeting by a minimal CRISPR RNA-guided cascade. Mol. Cell 63, 840–851 (2016).
Westra, E. R. et al. CRISPR immunity relies on the consecutive binding and degradation of negatively supercoiled invader DNA by Cascade and Cas3. Mol. Cell 46, 595–605 (2012).
Xiao, Y. et al. Structure basis for directional r-loop formation and substrate handover mechanisms in Type I CRISPR-Cas system. Cell 170, 48–60.e11 (2017).
Ivanov, I. E. et al. Cas9 interrogates DNA in discrete steps modulated by mismatches and supercoiling. Proc. Natl Acad. Sci. USA 117, 5853–5860 (2020).
Newton, M. D. et al. DNA stretching induces Cas9 off-target activity. Nat. Struct. Mol. Biol. 26, 185–192 (2019).
Szczelkun, M. D. et al. Direct observation of R-loop formation by single RNA-guided Cas9 and Cascade effector complexes. Proc. Natl Acad. Sci. USA 111, 9798–9803 (2014).
Klimasauskas, S., Kumar, S., Roberts, R. J. & Cheng, X. HhaI methyltransferase flips its target base out of the DNA helix. Cell 76, 357–369 (1994).
Reinisch, K. M., Chen, L., Verdine, G. L. & Lipscomb, W. N. The crystal structure of HaeIII methyltransferase convalently complexed to DNA: an extrahelical cytosine and rearranged base pairing. Cell 82, 143–153 (1995).
Dalhus, B., Laerdahl, J. K., Backe, P. H. & Bjørås, M. DNA base repair–recognition and initiation of catalysis. FEMS Microbiol. Rev. 33, 1044–1078 (2009).
Bell, J. C. & Kowalczykowski, S. C. RecA: regulation and mechanism of a molecular search engine. Trends Biochem. Sci. 41, 491–507 (2016).
Chen, Z., Yang, H. & Pavletich, N. P. Mechanism of homologous recombination from the RecA-ssDNA/dsDNA structures. Nature 453, 489–484 (2008).
Yang, D., Boyer, B., Prévost, C., Danilowicz, C. & Prentiss, M. Integrating multi-scale data on homologous recombination into a new recognition mechanism based on simulations of the RecA-ssDNA/dsDNA structure. Nucleic Acids Res. 43, 10251–10263 (2015).
Yang, H., Zhou, C., Dhar, A. & Pavletich, N. P. Mechanism of strand exchange from RecA-DNA synaptic and D-loop structures. Nature 586, 801–806 (2020).
Koo, H. S., Wu, H. M. & Crothers, D. M. DNA bending at adenine·thymine tracts. Nature 320, 501–506 (1986).
Garcia, H. G. et al. Biological consequences of tightly bent DNA: the other life of a macromolecular celebrity. Biopolymers 85, 115–130 (2007).
Cavaluzzi, M. J. & Borer, P. N. Revised UV extinction coefficients for nucleoside-5′-monophosphates and unpaired DNA and RNA. Nucleic Acids Res. 32, e13 (2004).
Acknowledgements
We thank D. Toso and J. Remis at the Cal Cryo facility for technical assistance in data collection. We thank members of the Nogales laboratory for discussions and advice on EM data processing, especially A.J. Florez Ariza and D. Herbst. We also thank J. Davis and E. Zhong for advice on EM data processing. We thank A. Chintangal for computational support. We thank N. Moriarty for assistance in modeling the thioalkane linker. We thank J. Kuriyan for scientific guidance and comments on the paper. We thank P. Pausch and H. Shi for comments on the paper. This work was supported by an National Science Foundation Graduate Research Fellowship (J.C.C.), an NHMRC Investigator grant (no. EL1, 1175568, G.J.K.), the Howard Hughes Medical Institute (J.A.D.), the National Science Foundation (award number 1817593, J.A.D.), the Centers for Excellence in Genomic Science of the National Institutes of Health (award number RM1HG009490, J.A.D.) and the Somatic Cell Genome Editing Program of the Common Fund of the National Institutes of Health (award number U01AI142817-02, J.A.D.). J.A.D. and E.N. are HHMI investigators.
Author information
Authors and Affiliations
Contributions
J.C.C. and J.A.D. conceived the study. J.C.C. produced all reagents and performed all biochemical experiments. J.C.C., K.M.S. and G.J.K. conducted structural studies including EM grid preparation, data collection and analysis, map calculation and model building and refinement. J.A.D. and E.N. provided supervision and guidance on data analysis and interpretation. J.C.C. produced the figures with assistance from K.M.S. J.C.C. wrote the manuscript with assistance from J.A.D. All authors edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The Regents of the University of California have patents issued and/or pending for CRISPR technologies on which G.J.K. and J.A.D. are inventors. J.A.D. is a cofounder of Caribou Biosciences, Editas Medicine, Scribe Therapeutics, Intellia Therapeutics and Mammoth Biosciences. J.A.D. is a scientific advisory board member of Vertex, Caribou Biosciences, Intellia Therapeutics, eFFECTOR Therapeutics, Scribe Therapeutics, Mammoth Biosciences, Algen Biotechnologies, Synthego, Algen Biotechnologies, Felix Biosciences, The Column Group, and Inari. J.A.D. is Chief Science Advisor to Sixth Street, a Director at Johnson & Johnson, Altos and Tempus, and has research projects sponsored by Biogen, Pfizer, Apple Tree Partners, and Roche. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Structural and Molecular Biology thanks Rick Russell, John van der Oost and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Sara Osman was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
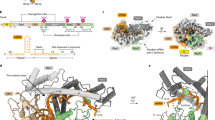
Extended Data Fig. 1 Characterization of the Cas9:DNA cross-link.
a, Crystal structure of Cas9:sgRNA:DNA with 20-bp RNA:DNA hybrid formed (PDB 4UN3). In the inset, Arg1333 and Arg1335 recognize the two guanines of the PAM. Green, NUC lobe; blue, REC lobe; orange, guide RNA; black, DNA; yellow, PAM. b, Non-reducing SDS-PAGE (Stain-Free) analysis of cross-linking reactions and controls. Complexes were prepared identically to structural constructs but in smaller volumes and without size exclusion purification. Competitor duplex, where indicated, was added before the cross-linkable duplex at an equivalent concentration. The depicted experiment was performed once, and a prior optimization experiment yielded similar results. c, Top left, non-reducing SDS-PAGE autoradiograph to determine the fraction of DNA cross-linked to Cas9 at t = 0. The target strand is radiolabeled. Bottom, reducing urea-PAGE autoradiograph revealing target-strand cleavage kinetics; quantification depicted in top right. The depicted model is \(y = C\left( {1 - e^{ - k_1t}} \right) + (B_{\max } - C)(1 - e^{ - k_2t})\). The depicted experiments were performed once, and a prior optimization experiment yielded similar results.
Extended Data Fig. 2 Cryo-EM sample quality.
a, SDS-PAGE (Stain-Free) analysis of purified proteins and cryo-EM samples. SC, structural construct; SC 1, Cas9:sgRNA:DNA with 0 RNA:DNA matches; SC 2, Cas9:sgRNA; SC 3, Cas9:sgRNA:DNA with 3 RNA:DNA matches. The depicted experiment was performed once (on the samples used for the final cryo-EM grids). b, Reducing urea-PAGE (SYBR-Gold-stained) analysis of purified nucleic acid components and cryo-EM samples. TS, target strand; NTS, non-target strand. The depicted experiment was performed once (on the samples used for the final cryo-EM grids). c, Reducing urea-PAGE autoradiograph of radioactive mimics of structural constructs. sgRNA 1 and non-target strand 1 are those used to create SC 1. sgRNA 2 and non-target strand 2 are those used to create SC 3. sgRNA 3 bears a spacer with 20 nt of complementarity to the DNA target strand. The depicted experiment was performed once, and a prior optimization experiment yielded similar results.
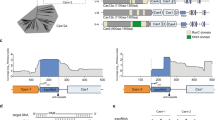
Extended Data Fig. 3 Cryo-EM analysis of Cas9:sgRNA:DNA with 0 RNA:DNA matches.
a, Classes from RELION 3D classification of closed-protein particles (threshold 6σ). The number of particles in each class is indicated next to the class number. In classes 1/2/5/6/7 the DNA is bent next to the PAM (visible for class 1 at lower contour). In classes 3/4/8 the DNA continues along a more linear trajectory for half a turn past the PAM, into the region normally occupied by REC2; in these classes, density in the region of the putative collision (black arrow) is uninterpretable as either protein or DNA, likely due to particle damage, and is thus colored gray. The class used for the final closed-protein/bent-DNA map is class 2, highlighted in yellow. Green, NUC lobe; blue, REC lobe domains 1/2; light blue, REC lobe domain 3; orange, guide RNA; magenta, DNA. b, Details of final cryo-EM maps. c, Linear DNA docked into open-protein/linear-DNA cryo-EM structure and previous Cas9 crystal structures. All structures were aligned to the C-terminal domain of PDB 5F9R; then, the linear DNA was aligned to the PAM-containing duplex of PDB 5F9R. Green, NUC lobe; blue, REC lobe; orange, guide RNA; black, docked DNA; yellow, PAM. The DNA truly belonging to each structure (if present) is depicted in gray.
Extended Data Fig. 4 Cryo-EM analysis of Cas9:sgRNA.
a, Details of cryo-EM analysis. b, Unsharpened cryo-EM map (threshold 5σ) and model of Cas9:sgRNA in open-protein conformation. Green, NUC lobe; blue, REC lobe; orange, guide RNA; gray, unattributed density (REC1 or guide RNA, see Supplementary Information).
Extended Data Fig. 5 Nucleic acid sequences used in DNA cyclization experiments.
Green star/bold T, fluorescein-conjugated dT; circled P, 5′ phosphate; CCR, candidate complementarity region.
Extended Data Fig. 6 Details of DNA cyclization experiments.
a, Fluorescence image and analysis of native PAGE gel resolving ligation products. Gel represents one replicate. Three replicates are plotted on the graphs. The polymeric/cyclized band assignments were made by reference to the relative electrophoretic mobilities observed in Kahn & Crothers, 1992. b, Comparison of Cas9:DNA cyclization data to CAP:DNA cyclization data. The depicted model is \(y = A \cdot \sin \left( {\frac{{2\pi }}{{10.45\ bp}}\left( {x + \phi _0} \right)} \right) + b\), with the following constraints: \(A > 0\), \(b > A\). The average of 224° and 212° is reported in Fig. 4c. J, J-factor (defined in Kahn & Crothers, 1992); Φ, phase difference; CBS, CAP-binding site.
Extended Data Fig. 7 Details of permanganate reactivity measurements.
a, Autoradiographs and analysis of all thymines except T(+25), which was insufficiently resolved from neighboring bands. The depicted autoradiographs are replicate 1. Due to systematic variation across replicates, individual replicates are presented on separate graphs and fitted separately. The depicted model is \(P_{ox} = \frac{{B_{\max }[Cas9:sgRNA]}}{{K_D + [Cas9:sgRNA]}}\), with KD shared across T(+1) and T(+2). CCR, candidate complementarity region; CI, confidence interval. b, Autoradiograph (same gel as ‘intact PAM’ autoradiograph in a) and analysis of experiments containing variants of Cas9:sgRNA (16 μM), with the ‘intact PAM’ DNA substrate. Graph depicts three replicates.
Extended Data Fig. 8 Structural features potentially relevant to Cas9-induced DNA bending.
a, Location of each feature in the bent-DNA structure. Green, NUC lobe; blue, REC lobe; orange, guide RNA; black, DNA; yellow, PAM. b, Comparison of phosphate lock loop (magenta) in various structures, within sharpened cryo-EM maps (row of three, threshold 8σ) or 2Fo-Fc map (upper right, threshold 1.5σ). Black arrow indicates the eponymous phosphate between nucleotides 0 and +1 of the target strand. c, Comparison of helix-rolling basic patch (magenta) in various structures, within unsharpened cryo-EM maps (first panel, threshold 5σ; second panel, threshold 6σ).
Extended Data Fig. 9 Cryo-EM analysis of Cas9:sgRNA:DNA with 3 RNA:DNA matches.
Details of cryo-EM analysis.
Supplementary information
Supplementary Information
Supplementary methods and discussion, best-fit model parameters and plasmid, protein and oligonucleotide sequences.
Supplementary Video 1
Morphing the open-protein/linear-DNA conformation to the closed-protein/bent-DNA conformation. PyMOL’s morph function was used to interpolate a physically plausible transition between the two cryo-EM structures. Only atoms shared between both structures are depicted during the transition. Green, NUC lobe; blue, REC1/2; gray, REC3; orange, RNA; black, DNA and yellow, PAM.
Supplementary Video 2
Visualizing the helix-rolling basic patch during the open-to-closed transition. Morph as described for Supplementary Video 1. Green, NUC lobe; blue, REC1/2; gray, REC3; orange, RNA; black, DNA; yellow, PAM; magenta spheres, HRBP lysines.
Supplementary Video 3
Results of multi-body refinement of the linear-DNA/open-protein particles. Five principal components of rotation/translation are shown.
Supplementary Video 4
Results of multi-body refinement of the bent-DNA/closed-protein particles. Five principal components of rotation/translation are shown.
Source data
Source Data Fig. 4
Numerical source data.
Source Data Fig. 5
Numerical source data, uncropped autoradiograph.
Source Data Extended Data Fig. 1
Numerical source data, uncropped fluorescence image, uncropped autoradiographs.
Source Data Extended Data Fig. 2
Uncropped fluorescence images, uncropped autoradiograph.
Source Data Extended Data Fig. 6
Numerical source data, uncropped fluorescence image.
Source Data Extended Data Fig. 7
Numerical source data, uncropped autoradiographs.
Rights and permissions
About this article
Cite this article
Cofsky, J.C., Soczek, K.M., Knott, G.J. et al. CRISPR–Cas9 bends and twists DNA to read its sequence. Nat Struct Mol Biol 29, 395–402 (2022). https://doi.org/10.1038/s41594-022-00756-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00756-0
This article is cited by
-
Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
Nature Communications (2024)
-
Engineering Cas9: next generation of genomic editors
Applied Microbiology and Biotechnology (2024)
-
Recent advances in therapeutic CRISPR-Cas9 genome editing: mechanisms and applications
Molecular Biomedicine (2023)
-
Targeting miRNA by CRISPR/Cas in cancer: advantages and challenges
Military Medical Research (2023)
-
Comprehensive computational analysis of epigenetic descriptors affecting CRISPR-Cas9 off-target activity
BMC Genomics (2022)