Abstract
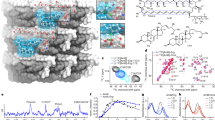
Amphotericin B (AmB) is a powerful but toxic fungicide that operates via enigmatic small molecule–small molecule interactions. This mechanism has challenged the frontiers of structural biology for half a century. We recently showed AmB primarily forms extramembranous aggregates that kill yeast by extracting ergosterol from membranes. Here, we report key structural features of these antifungal ‘sponges’ illuminated by high-resolution magic-angle spinning solid-state NMR, in concert with simulated annealing and molecular dynamics computations. The minimal unit of assembly is an asymmetric head-to-tail homodimer: one molecule adopts an all-trans C1–C13 motif, the other a C6–C7-gauche conformation. These homodimers are staggered in a clathrate-like lattice with large void volumes similar to the size of sterols. These results illuminate the atomistic interactions that underlie fungicidal assemblies of AmB and suggest this natural product may form biologically active clathrates that host sterol guests.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data that support the findings of this study are available within the paper and its Supplementary Information. Source data are provided in BMRbig, entry ID bmrbig28. Atomic coordinates are deposited in BMRB, entry ID 21097. Further requests can be directed to the corresponding authors.
References
LaPlante, S. R. et al. Compound aggregation in drug discovery: implementing a practical NMR assay for medicinal chemists. J. Med. Chem. 56, 5142–5150 (2013).
Sezgin, E., Levental, I., Mayor, S. & Eggeling, C. The mystery of membrane organization: composition, regulation and physiological relevance of lipid rafts. Nat. Rev. Mol. Cell Biol. 18, 361–374 (2017).
Lehn, J. M. Towards complex matter: supramolecular chemistry and self-organization. Eur. Rev. 17, 263–280 (2009).
Muraglia, K. A. et al. Small-molecule ion channels increase host defences in cystic fibrosis airway epithelia. Nature 567, 405–408 (2019).
Grillo, A. S. et al. Restored iron transport by a small molecule promotes absorption and hemoglobinization in animals. Science 616, 608–616 (2017).
Bongomin, F., Gago, S., Oladele, R. O. & Denning, D. W. Global and multi-national prevalence of fungal diseases—estimate precision. J. Fungi 3, 57 (2017).
Sanglard, D. Resistance of human fungal pathogens to antifungal drugs. Curr. Opin. Microbiol. 5, 379–385 (2002).
Patterson, T. F. et al. Practice guidelines for the diagnosis and management of aspergillosis: 2016 update by the Infectious Diseases Society of America. Clin. Infect. Dis. 63, e1–e60 (2016).
Fuhren, J. et al. High prevalence of azole resistance in Aspergillus fumigatus isolates from high-risk patients. J. Antimicrob. Chemother. 70, 2894–2898 (2015).
Perfect, J. R. The antifungal pipeline: a reality check. Nat. Rev. Drug Discov. 16, 603–616 (2017).
Cleveland, A. A. et al. Changes in incidence and antifungal drug resistance in candidemia: results from population-based laboratory surveillance in Atlanta and Baltimore, 2008–2011. Clin. Infect. Dis. 55, 1352–1361 (2012).
Kreusch, A. & Karstaedt, A. S. Candidemia among adults in Soweto, South Africa, 1990–2007. Int. J. Infect. Dis. 17, e621–e623 (2013).
Verweij, P. E. et al. International expert opinion on the management of infection caused by azole-resistant Aspergillus fumigatus. Drug Resist. Updat. 21–22, 30–40 (2015).
Ermishkin, L. N., Kasumov, K. M. & Potzeluyev, V. M. Single ionic channels induced in lipid bilayers by polyene antibiotics amphotericin-B and nystatine. Nature 262, 698–699 (1976).
Katzung, B. G., Masters, S. B. & Trevor, A. J. Basic & Clinical Pharmacology 12th edn (McGraw-Hill, 2012).
Umegawa, Y., Matsumori, N., Oishi, T. & Murata, M. Ergosterol increases the intermolecular distance of amphotericin B in the membrane-bound assembly as evidenced by solid-state NMR. Biochemistry 47, 13463–13469 (2008).
Khutorsky, V. E. Structures of amphotericin B–cholesterol complex. Biochim. Biophys. Acta 1108, 123–127 (1992).
Gray, K. C. et al. Amphotericin primarily kills yeast by simply binding ergosterol. Proc. Natl Acad. Sci. USA 109, 2234–2239 (2012).
Palacios, D. S., Anderson, T. M. & Burke, M. D. A post-PKS oxidation of the amphotericin B skeleton predicted to be critical for channel formation is not required for potent antifungal activity. J. Am. Chem. Soc. 129, 13804–13805 (2007).
Palacios, D. S., Dailey, I., Siebert, D. M., Wilcock, B. C. & Burke, M. D. Synthesis-enabled functional group deletions reveal key underpinnings of amphotericin B ion channel and antifungal activities. Proc. Natl Acad. Sci. USA 108, 6733–6738 (2011).
Anderson, T. M. et al. Amphotericin forms an extramembranous and fungicidal sterol sponge. Nat. Chem. Biol. 10, 400–406 (2014).
Guo, X. R. et al. Sterol sponge mechanism is conserved for glycosylated polyene macrolides. ACS Cent. Sci. 7, 781–791 (2021).
Sun, D., Rosokha, S. V., Lindeman, S. V. & Kochi, J. K. Intervalence (charge-resonance) transitions in organic mixed-valence systems. Through-space versus through-bond electron transfer between bridged aromatic (redox) centers. J. Am. Chem. Soc. 125, 15950–15963 (2003).
Wylie, B. J., Franks, W. T., Graesser, D. T. & Rienstra, C. M. Site-specific 13C chemical shift anisotropy measurements in a uniformly 15N,13C-labeled microcrystalline protein by 3D magic-angle spinning NMR spectroscopy. J. Am. Chem. Soc. 127, 11946–11947 (2005).
Schwieters, C. D., Kuszewski, J. J. & Marius Clore, G. Using Xplor-NIH for NMR molecular structure determination. Prog. Nucl. Magn. Reson. Spectrosc. 48, 47–62 (2006).
Ulrich, E. L. et al. BioMagResBank. Nucleic Acids Res. 36, D402–D408 (2008).
Shen, Y., Delaglio, F., Cornilescu, G. & Bax, A. TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J. Biomol. NMR 44, 213–223 (2009).
Caulkins, B. G. et al. NMR crystallography of a carbanionic intermediate in tryptophan synthase: chemical structure, tautomerization, and reaction specificity. J. Am. Chem. Soc. 138, 15214–15226 (2016).
Wylie, B. J., Schwieters, C. D., Oldfield, E. & Rienstra, C. M. Protein structure refinement using 13C alpha chemical shift tensors. J. Am. Chem. Soc. 131, 985–992 (2009).
Wylie, B. J. et al. Ultrahigh resolution protein structures using NMR chemical shift tensors. Proc. Natl Acad. Sci. USA 108, 16974–16979 (2011).
Dalling, D. K. & Grant, D. M. Carbon-13 magnetic resonance. IX. The methylcyclohexanes. J. Am. Chem. Soc. 760, 6612–6622 (1967).
Hohwy, M., Rienstra, C. M., Jaroniec, C. P. & Griffin, R. G. Fivefold symmetric homonuclear dipolar recoupling in rotating solids: application to double quantum spectroscopy. J. Chem. Phys. 110, 7983–7992 (1999).
Reif, B., Jaroniec, C. P., Rienstra, C. M., Hohwy, M. & Griffin, R. G. 1H–1H MAS correlation spectroscopy and distance measurements in a deuterated peptide. J. Magn. Reson. 151, 320–327 (2001).
Takegoshi, K., Nakamura, S. & Terao, T. 13C–1H dipolar-assisted rotational resonance in magic-angle spinning NMR. Chem. Phys. Lett. 344, 631–637 (2001).
Adamo, C. & Barone, V. Exchange functionals with improved long-range behavior and adiabatic connection methods without adjustable parameters: the mPW and mPW1PW models. J. Chem. Phys. 108, 664–675 (1998).
Krishnan, R., Binkley, J. S., Seeger, R. & Pople, J. A. Self-consistent molecular orbital methods. XX. A basis set for correlated wave functions. J. Chem. Phys. 72, 650–654 (1980).
Paëpe, G. D. E., Lewandowski, J. R., Loquet, A., Böckmann, A. & Griffin, R. G. Proton assisted recoupling and protein structure determination. J. Chem. Phys. 129, 245101 (2008).
Nimerovsky, E. et al. Phase-modulated LA-REDOR: a robust, accurate and efficient solid-state NMR technique for distance measurements between a spin-1/2 and a quadrupole spin. J. Magn. Reson. 244, 107–113 (2014).
Makrinich, M., Nimerovsky, E. & Goldbourt, A. Pushing the limit of NMR-based distance measurements – retrieving dipolar couplings to spins with extensively large quadrupolar frequencies. Solid State Nucl. Magn. Reson. 92, 19–24 (2018).
Schwieters, C. D., Bermejo, G. A. & Clore, G. M. Xplor-NIH for molecular structure determination from NMR and other data sources. Protein Sci. 27, 26–40 (2018).
Rousseeuw, P. et al. cluster: 'Finding Groups in Data’: Cluster Analysis Extended Rousseeuw et al. R package version 2.1.2 https://rdrr.io/cran/cluster/ (2019).
Chorghade, R. S. et al. Amphotericin B induces epithelial voltage responses in people with cystic fibrosis. J. Cyst. Fibros. 20, 540–550 (2021).
Richardson, D., Vriend, G. & Allison, J. Tools 2020: a compilation of tools for protein science. Protein Sci. 29, 5–7 (2020).
Tuttle, M. D. Solid-state NMR structure of a pathogenic fibril of full-length human α-synuclein. Nat. Struct. Mol. Bio. 23, 409–415 (2016).
Nishimura, S. & Matsumori, N. Chemical diversity and mode of action of natural products targeting lipids in the eukaryotic cell membrane. Nat. Prod. Rep. 37, 677–702 (2020).
Matsumori, N., Sawada, Y. & Murata, M. Large molecular assembly of amphotericin B formed in ergosterol-containing membrane evidenced by solid-state NMR of intramolecular bridged derivative. J. Am. Chem. Soc. 128, 11977–11984 (2006).
Baginski, M., Resat, H. & Borowski, E. Comparative molecular dynamics simulations of amphotericin B–cholesterol/ergosterol membrane channels. Biochim. Biophys. Acta - Biomembr. 1567, 63–78 (2002).
Ganis, P., Avitabil, G., Mechlins, W. & Schaffne, C. Polyene macrolide antibiotic amphotericin-B – crystal structure of N-iodoacetyl derivative. J. Am. Chem. Soc. 93, 4560–4564 (1971).
Jarzembska, K. N. et al. Controlled crystallization, structure, and molecular properties of iodoacetylamphotericin B. Cryst. Growth Des. 12, 2336–2345 (2012).
Matsumori, N., Sawada, Y. & Murata, M. Mycosamine orientation of amphotericin B controlling interaction with ergosterol: sterol-dependent activity of conformation-restricted derivatives with an amino-carbonyl bridge. J. Am. Chem. Soc. 127, 10667–10675 (2005).
Wilcock, B. C., Endo, M. M., Uno, B. E. & Burke, M. D. C2′-OH of amphotericin B plays an important role in binding the primary sterol of human cells but not yeast cells. J. Am. Chem. Soc. 135, 8488–8491 (2013).
Knight, Z. A. & Shokat, K. M. Features of selective kinase inhibitors. Chem. Biol. 12, 621–637 (2005).
Duggan, K. C. et al. (R)-Profens are substrate-selective inhibitors of endocannabinoid oxygenation by COX-2. Nat. Chem. Biol. 7, 803–809 (2011).
Hisao, G. S. et al. An efficient method and device for transfer of semisolid materials into solid-state NMR spectroscopy rotors. J. Magn. Reson. 265, 172–176 (2016).
Sebaugh, J. L. Guidelines for accurate EC50/IC50 estimation. Pharm. Stat. 10, 128–134 (2011).
MATLAB v.R2015a. (MathWorks, 2015).
Hanwell, M. D. et al. Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J. Cheminform. 4, 17 (2012).
Frisch, M. J. et al. Gaussian v.9 (Gaussian Inc., 2009).
Stephens, P. J., Devlin, F. J., Chabalowski, C. F. & Frisch, M. J. Ab initio calculation of vibrational absorption and circular dichroism spectra using density functional force fields. J. Phys. Chem. 98, 11623–11627 (1994).
Hariharan, P. C. & Pople, J. A. The influence of polarization functions on molecular orbital hydrogenation energies. Theor. Chim. Acta 28, 213–222 (1973).
Tomasi, J., Mennucci, B. & Cammi, R. Quantum mechanical continuum solvation models. Chem. Rev. 105, 2999–3093 (2005).
Morcombe, C. R. & Zilm, K. W. Chemical shift referencing in MAS solid state NMR. J. Magn. Reson. 162, 479–486 (2003).
Gullion, T. Measurement of dipolar interactions between spin-12 and quadrupolar nuclei by rotational-echo, adiabatic-passage, double-resonance NMR. Chem. Phys. Lett. 246, 325–330 (1995).
Chen, L. et al. Distance measurement between a spin-1/2 and a half-integer quadrupolar nuclei by solid-state NMR using exact analytical expressions. J. Magn. Reson. 206, 269–273 (2010).
Schwieters, C. D. & Clore, G. M. A pseudopotential for improving the packing of ellipsoidal protein structures determined from NMR data. J. Phys. Chem. B 112, 6070–6073 (2008).
Acknowledgements
This work was supported by the US National Institutes of Health (NIH) R01-GM112845 and R01-GM123455 to C.M.R. and R35-GM118185 to M.D.B. This study made use of the National Magnetic Resonance Facility at Madison, including the technology development program which is supported by NIH grant P41GM136463. C.D.S. was supported by the National Institutes of Health Intramural Research Program of the National Institute of Diabetes and Digestive and Kidney Diseases. We would like to dedicate this paper to the memory of our coauthor, A. Khandelwal, who passed away during the course of this investigation.
Author information
Authors and Affiliations
Contributions
A.L., A.I.G., C.M.R. and M.D.B. designed the research. J.T.H. and G.S.H. prepared and purified [U-13C]AmB. J.T.H. prepared and purified [13C] skip-labeled AmB. A.L. and C.P.S. prepared samples for SSNMR. A.L., G.S.H., C.P.S. and C.M.R. acquired SSNMR data. A.L. and C.P.S. performed UV spectroscopy measurements. A.L, C.P.S. and G.S.H. analyzed SSNMR data. E.N. performed and processed the PM-RESPDOR experiments. A.L., C.P.S. and A.I.G analyzed UV data. A.I.G. designed and together with A.L. and C.P.S. performed PCA analysis. A.K. and J.Z. and A.M.S. performed cell-based assays. A.M.S. synthesized AmB-I. J.Z. and A.M.S. purified AmB-I. T.V.P. performed structural calculations in MOE. A.I.G. and A.M.D.L. performed and analyzed DFT structural calculations. A.L., C.P.S., C.D.S and C.M.R. performed XPLOR-NIH structure calculations. A.M.D.L. and T.V.P. designed and performed clustering analyses. A.L., C.P.S., A.M.D.L., T.V.P, M.D.B and C.M.R. prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Structural & Molecular Biology thanks Hartmut Oschkinat and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Florian Ullrich was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 AmB homogenization validation.
(a) UV-Vis spectra of AmB aggregates before (black) and after (red) homogenization. Grey UV-Vis spectra correspond to representative samples from the middle of the PC1 range (Fig. 1e). (b) LC-MS spectra of AmB before and after homogenization. (c) 1H NMR spectra of AmB before (black line) and after (red line) homogenization.
Extended Data Fig. 2 AmB cluster analysis and Erg docking.
(a) Principal component analysis (PCA) of the 9-dimensional data used for dimer structure clustering analysis for all 11 cluster ensembles. The inset shows the lowest clusters’ energies overlaid on the total energy histogram. (b) Alignment of medoid dimer structures of clusters 1–3 and lattice structures of medoid structures from the lowest energy cluster; states A and B of AmB are marked. (c) Medoid lattices of clusters 1–3 showing representative void volumes in green. Approximate volumes for each void in clusters 1-3 are 390 Å3, 830 Å3, and 530 Å3, respectively. The volume of ergosterol is estimated to be 427 Å3 21. (d) Snapshots of all four void pockets found in cluster 1 lattice - front view (left) and side view (right). The volume of each pocket is approximately 390 Å3. The number density of this lattice is 6.36 molecules per 10,000 Å3. (e) Snapshot of an ergosterol molecule (cpk-blue) docked to the cluster 2 medoid lattice – front view. (f) Close-up snapshot of AmB (white) – ergosterol (blue) docked interaction as seen in (e). Select atom numbers near interaction site are labeled for both AmB and ergosterol molecules.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7, Discussion and Tables 1–6.
Rights and permissions
About this article
Cite this article
Lewandowska, A., Soutar, C.P., Greenwood, A.I. et al. Fungicidal amphotericin B sponges are assemblies of staggered asymmetric homodimers encasing large void volumes. Nat Struct Mol Biol 28, 972–981 (2021). https://doi.org/10.1038/s41594-021-00685-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-021-00685-4