Abstract
The protein K-Ras functions as a molecular switch in signaling pathways regulating cell growth. In the human mitogen-activated protein kinase (MAPK) pathway, which is implicated in many cancers, multiple K-Ras proteins are thought to assemble at the cell membrane with Ras effector proteins from the Raf family. Here we propose an atomistic structural model for such an assembly. Our starting point was an asymmetric guanosine triphosphate-mediated K-Ras dimer model, which we generated using unbiased molecular dynamics simulations and verified with mutagenesis experiments. Adding further K-Ras monomers in a head-to-tail fashion led to a compact helical assembly, a model we validated using electron microscopy and cell-based experiments. This assembly stabilizes K-Ras in its active state and presents composite interfaces to facilitate Raf binding. Guided by existing experimental data, we then positioned C-Raf, the downstream kinase MEK1 and accessory proteins (Galectin-3 and 14-3-3σ) on and around the helical assembly. The resulting Ras–Raf signalosome model offers an explanation for a large body of data on MAPK signaling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Atomic coordinates of the structural model of the eight-protomer Ras–Raf signalosome are available as Supplementary Dataset 1, and atomic coordinates of the GMA K-Ras dimer on a membrane are available as Supplementary Dataset 2. Due to the large size of the molecular dynamics trajectories (listed in Supplementary Table 1), they are available upon request (for noncommercial use) by contacting trajectories@deshawresearch.com. Source data are provided with this paper.
Code availability
The molecular dynamics simulations were performed using the Anton 2 supercomputer (the simulation code we used is specialized to Anton 2, but codes for performing molecular dynamics simulations are widely available).
References
Wennerberg, K., Rossman, K. L. & Der, C. J. The Ras superfamily at a glance. J. Cell Sci. 118, 843–846 (2005).
Hobbs, G. A., Der, C. J. & Rossman, K. L. RAS isoforms and mutations in cancer at a glance. J. Cell Sci. 129, 1287–1292 (2016).
Yuan, T. L. et al. Differential effector engagement by oncogenic KRAS. Cell Rep. 22, 1889–1902 (2018).
Tidyman, W. E. & Rauen, K. A. The RASopathies: developmental syndromes of Ras/MAPK pathway dysregulation. Curr. Opin. Genet. Dev. 19, 230–236 (2009).
Prior, I. A., Lewis, P. D. & Mattos, C. A comprehensive survey of Ras mutations in cancer. Cancer Res. 72, 2457–2467 (2012).
Simanshu, D. K., Nissley, D. V. & McCormick, F. RAS proteins and their regulators in human disease. Cell 170, 17–33 (2017).
Cherfils, J. & Zeghouf, M. Regulation of small GTPases by GEFs, GAPs, and GDIs. Physiol. Rev. 93, 269–309 (2013).
Wittinghofer, A. & Pal, E. F. The structure of Ras protein: a model for a universal molecular switch. Trends Biochem. Sci. 16, 382–387 (1991).
Wittinghofer, A. & Vetter, I. R. Structure-function relationships of the G domain, a canonical switch motif. Annu. Rev. Biochem. 80, 943–971 (2011).
Santos, E., Nebreda, A. R., Bryan, T. & Kempner, E. S. Oligomeric structure of p21 ras proteins as determined by radiation inactivation. J. Biol. Chem. 263, 9853–9858 (1988).
Dementiev, A. K-Ras4B lipoprotein synthesis: biochemical characterization, functional properties, and dimer formation. Protein Expr. Purif. 84, 86–93 (2012).
Chen, M., Peters, A., Huang, T. & Nan, X. Ras dimer formation as a new signaling mechanism and potential cancer therapeutic target. Mini-Rev. Med. Chem. 16, 391–403 (2016).
Güldenhaupt, J. et al. N-Ras forms dimers at POPC membranes. Biophys. J. 103, 1585–1593 (2012).
Werkmüller, A., Triola, G., Waldmann, H. & Winter, R. Rotational and translational dynamics of Ras proteins upon binding to model membrane systems. Chem. Phys. Chem. 14, 3698–3705 (2013).
Sarkar-Banerjee, S. et al. Spatiotemporal analysis of K-Ras plasma membrane interactions reveals multiple high order homo-oligomeric complexes. J. Am. Chem. Soc. 139, 13466–13475 (2017).
Muratcioglu, S. et al. GTP-dependent K-Ras dimerization. Structure 23, 1325–1335 (2015).
Inouye, K., Mizutani, S., Koide, H. & Kaziro, Y. Formation of the Ras dimer is essential for Raf-1 activation. J. Biol. Chem. 275, 3737–3740 (2000).
Nan, X. et al. Ras-GTP dimers activate the mitogen-activated protein kinase (MAPK) pathway. Proc. Natl Acad. Sci. USA 112, 7996–8001 (2015).
Spencer-Smith, R. et al. Inhibition of RAS function through targeting an allosteric regulatory site. Nat. Chem. Biol. 13, 62–68 (2017).
Ambrogio, C. et al. KRAS dimerization impacts MEK inhibitor sensitivity and oncogenic activity of mutant KRAS. Cell 172, 857–868 (2018).
Plowman, S. J., Muncke, C., Parton, R. G. & Hancock, J. F. H-ras, K-ras, and inner plasma membrane raft proteins operate in nanoclusters with differential dependence on the actin cytoskeleton. Proc. Natl Acad. Sci. USA 102, 15500–15505 (2005).
Tian, T. et al. Plasma membrane nanoswitches generate high-fidelity Ras signal transduction. Nat. Cell Biol. 9, 905–914 (2007).
Shalom-Feuerstein, R. et al. K-Ras nanoclustering is subverted by overexpression of the scaffold protein Galectin-3. Cancer Res. 68, 6608–6616 (2008).
Belanis, L., Plowman, S. J., Rotblat, B., Hancock, J. F. & Kloog, Y. Galectin-1 is a novel structural component and a major regulator of H-Ras nanoclusters. Mol. Biol. Cell 19, 1404–1414 (2008).
Plowman, S. J., Ariotti, N., Goodall, A., Parton, R. G. & Hancock, J. F. Electrostatic interactions positively regulate K-Ras nanocluster formation and function. Mol. Cell. Biol. 28, 4377–4385 (2008).
Sutton, M. N. et al. DIRAS3 (ARHI) blocks RAS/MAPK signaling by binding directly to RAS and disrupting RAS clusters. Cell Rep. 29, 3448–3459 (2019).
Wu, H. Higher-order assemblies in a new paradigm of signal transduction. Cell 153, 287–292 (2013).
Wright, L. P. & Philips, M. R. Thematic review series: lipid posttranslational modifications. CAAX modification and membrane targeting of Ras. J. Lipid Res. 47, 883–891 (2006).
Rajakulendran, T., Sahmi, M., Lefrançois, M., Sicheri, F. & Therrien, M. A dimerization-dependent mechanism drives RAF catalytic activation. Nature 461, 542–545 (2009).
Shan, Y. et al. How does a drug molecule find its target binding site? J. Am. Chem. Soc. 133, 9181–9183 (2011).
Shan, Y. et al. Molecular basis for pseudokinase-dependent autoinhibition of JAK2 tyrosine kinase. Nat. Struct. Mol. Biol. 21, 579–584 (2014).
Plattner, N., Doerr, S., De Fabritiis, G. & Noé, F. Complete protein-protein association kinetics in atomic detail revealed by molecular dynamics simulations and Markov modelling. Nat. Chem. 9, 1005–1011 (2017).
Scheffzek, K. et al. The Ras-RasGAP complex: structural basis for GTPase activation and its loss in oncogenic Ras mutants. Science 277, 333–338 (1997).
Ahmadian, M. R., Stege, P., Scheffzek, K. & Wittinghofer, A. Confirmation of the arginine-finger hypothesis for the GAP-stimulated GTP-hydrolysis reaction of Ras. Nat. Struct. Biol. 4, 686–689 (1997).
Vetter, I. R. & Wittinghofer, A. The guanine nucleotide-binding switch in three dimensions. Science 294, 1299–1304 (2001).
Geyer, M. et al. Conformational transitions in p21ras and in its complexes with the effector protein Raf-RBD and the GTPase activating protein GAP. Biochemistry 35, 10308–10320 (1996).
Ito, Y. et al. Regional polysterism in the GTP-bound form of the human c-Ha-Ras protein. Biochemistry 36, 9109–9119 (1997).
Spoerner, M., Herrmann, C., Vetter, I. R., Kalbitzer, H. R. & Wittinghofer, A. Dynamic properties of the Ras switch I region and its importance for binding to effectors. Proc. Natl Acad. Sci. USA 98, 4944–4949 (2001).
Araki, M. et al. Solution structure of the state 1 conformer of GTP-bound H-Ras protein and distinct dynamic properties between the state 1 and state 2 conformers. J. Biol. Chem. 286, 39644–39653 (2011).
Mazhab-Jafari, M. T. et al. Oncogenic and RASopathy-associated K-RAS mutations relieve membrane-dependent occlusion of the effector-binding site. Proc. Natl Acad. Sci. USA 112, 6625–6630 (2015).
Gorfe, A. A., Hanzal-Bayer, M., Abankwa, D., Hancock, J. F. & McCammon, J. A. Structure and dynamics of the full-length lipid-modified H-Ras protein in a 1,2-dimyristoylglycero-3-phosphocholine bilayer. J. Med. Chem. 50, 674–684 (2007).
Prakash, P., Zhou, Y., Liang, H., Hancock, J. F. & Gorfe, A. A. Oncogenic K-Ras binds to an anionic membrane in two distinct orientations: a molecular dynamics analysis. Biophys. J. 110, 1125–1138 (2016).
Drosten, M. et al. Genetic analysis of Ras signalling pathways in cell proliferation, migration and survival. EMBO J. 29, 1091–1104 (2010).
Sung, Y. J., Carter, M., Zhong, J. M. & Hwang, Y. W. Mutagenesis of the H-ras p21 at glycine-60 residue disrupts GTP-induced conformational change. Biochemistry 34, 3470–3477 (1995).
Zhou, Y. & Hancock, J. F. Ras nanoclusters: versatile lipid-based signaling platforms. Biochim. Biophys. Acta 1853, 841–849 (2015).
Ostrem, J. M., Peters, U., Sos, M. L., Wells, J. A. & Shokat, K. M. K-Ras (G12C) inhibitors allosterically control GTP affinity and effector interactions. Nature 503, 548–551 (2013).
Fetics, S. K. et al. Allosteric effects of the oncogenic RasQ61L mutant on Raf-RBD. Structure 23, 505–516 (2015).
Hu, C.-D. et al. Cysteine-rich region of Raf-1 interacts with activator domain of post-translationally modified Ha-Ras. J. Biol. Chem. 270, 30274–30277 (1995).
Okada, T. et al. The strength of interaction at the Raf cysteine-rich domain is a critical determinant of response of Raf to Ras family small GTPases. Mol. Cell. Biol. 19, 6057–6064 (1999).
Drugan, J. K. et al. Ras interaction with two distinct binding domains in Raf-1 may be required for Ras transformation. J. Biol. Chem. 271, 233–237 (1996).
Williams, J. G. et al. Elucidation of binding determinants and functional consequences of Ras/Raf-cysteine-rich domain interactions. J. Biol. Chem. 275, 22172–22179 (2000).
Winkler, D. G. et al. Identification of residues in the cysteine-rich domain of Raf-1 that control Ras binding and Raf-1 activity. J. Biol. Chem. 273, 21578–21584 (1998).
Thapar, R., Williams, J. G. & Campbell, S. L. NMR characterization of full-length farnesylated and non-farnesylated H-Ras and its implications for Raf activation. J. Mol. Biol. 343, 1391–1408 (2004).
Levy, R., Biran, A., Poirier, F., Raz, A. & Kloog, Y. Galectin-3 mediates cross-talk between K-Ras and Let-7c tumor suppressor microRNA. PLoS ONE 6, e27490 (2011).
Elad-Sfadia, G., Haklai, R., Balan, E. & Kloog, Y. Galectin-3 augments K-Ras activation and triggers a Ras signal that attenuates ERK but not phosphoinositide 3-kinase activity. J. Biol. Chem. 279, 34922–34930 (2004).
Ashery, U. et al. Spatiotemporal organization of Ras signaling: rasosomes and the galectin switch. Cell. Mol. Neurobiol. 26, 469–493 (2006).
Paz, A., Haklai, R., Elad-Sfadia, G., Ballan, E. & Kloog, Y. Galectin-1 binds oncogenic H-Ras to mediate Ras membrane anchorage and cell transformation. Oncogene 20, 7486–7493 (2001).
Rotblat, B. et al. Galectin-1 (L11A) predicted from a computed galectin-1 farnesyl-binding pocket selectively inhibits Ras-GTP. Cancer Res. 64, 3112–3118 (2004).
Lukyanov, P., Furtak, V. & Ochieng, J. Galectin-3 interacts with membrane lipids and penetrates the lipid bilayer. Biochem. Biophys. Res. Commun. 338, 1031–1036 (2005).
Yang, R.-Y., Hill, P. N., Hsu, D. K. & Liu, F.-T. Role of the carboxyl-terminal lectin domain in self-association of galectin-3. Biochemistry 37, 4086–4092 (1998).
Dharmaiah, S. et al. Structural basis of recognition of farnesylated and methylated KRAS4b by PDEδ. Proc. Natl Acad. Sci. USA 113, E6766–E6775 (2016).
Tzivion, G., Luo, Z. & Avruch, J. A dimeric 14-3-3 protein is an essential cofactor for Raf kinase activity. Nature 394, 88–92 (1998).
Molzan, M. et al. Stabilization of physical RAF/14-3-3 interaction by cotylenin A as treatment strategy for RAS mutant cancers. ACS Chem. Biol. 8, 1869–1875 (2013).
Hatzivassiliou, G. et al. RAF inhibitors prime wild-type RAF to activate the MAPK pathway and enhance growth. Nature 464, 431–435 (2010).
Haling, J. R. et al. Structure of the BRAF-MEK complex reveals a kinase activity independent role for BRAF in MAPK signaling. Cancer Cell 26, 402–413 (2014).
Park, E. et al. Architecture of autoinhibited and active BRAF-MEK1-14-3-3 complexes. Nature 575, 545–550 (2019).
Kondo, Y. et al. Cryo-EM structure of a dimeric B-Raf:14-3-3 complex reveals asymmetry in the active sites of B-Raf kinases. Science 366, 109–115 (2019).
Wu, H. & Fuxreiter, M. The structure and dynamics of higher-order assemblies: amyloids, signalosomes, and granules. Cell 165, 1055–1066 (2016).
Boriack-Sjodin, P. A., Margarit, S. M., Bar-Sagi, D. & Kuriyan, J. The structural basis of the activation of Ras by SOS. Nature 394, 337–343 (1998).
Margarit, S. M. et al. Structural evidence for feedback activation by Ras·GTP of the Ras-specific nucleotide exchange factor SOS. Cell 112, 685–695 (2003).
Prakash, P. et al. Computational and biochemical characterization of two partially overlapping interfaces and multiple weak-affinity K-Ras dimers. Sci. Rep. 7, 40109 (2017).
Jang, H., Muratcioglu, S., Gursoy, A., Keskin, O. & Nussinov, R. Membrane-associated Ras dimers are isoform-specific: K-Ras dimers differ from H-Ras dimers. Biochem. J. 473, 1719–1732 (2016).
Lee, K. Y. et al. Two distinct structures of membrane-associated homodimers of GTP- and GDP-bound KRAS4B revealed by paramagnetic relaxation enhancement. Angew. Chem. Int. Ed. Engl. 59, 11037–11045 (2020).
Cookis, T. & Mattos, C. Crystal structure reveals the full Ras:Raf interface and advances mechanistic understanding of Raf activation. Preprint at bioRxiv https://doi.org/10.1101/2020.07.28.225938 (2020).
Tran, T. H. et al. KRAS interaction with RAF1 RAS-binding domain and cysteine-rich domain provides insights into RAS-mediated RAF activation. Nat. Commun. 12, 1176 (2021).
Fang, Z. et al. Multivalent assembly of KRAS with the RAS-binding and cysteine-rich domains of CRAF on the membrane. Proc. Natl Acad. Sci. USA 117, 12101–12108 (2020).
Cho, K.-J. et al. Raf inhibitors target Ras spatiotemporal dynamics. Curr. Biol. 22, 945–955 (2012).
Zhang, Z. et al. Wildtype Kras2 can inhibit lung carcinogenesis in mice. Nat. Genet. 29, 25–33 (2001).
Singh, A., Sowjanya, A. P. & Ramakrishna, G. The wild-type Ras: road ahead. FASEB J. 19, 161–169 (2005).
Lavoie, H. et al. MEK drives BRAF activation through allosteric control of KSR proteins. Nature 554, 549–553 (2018).
Rodriguez-Viciana, P., Oses-Prieto, J., Burlingame, A., Fried, M. & McCormick, F. A phosphatase holoenzyme comprised of Shoc2/Sur8 and the catalytic subunit of PP1 functions as an M-Ras effector to modulate Raf activity. Mol. Cell 22, 217–230 (2006).
Li, W., Han, M. & Guan, K. L. The leucine-rich repeat protein SUR-8 enhances MAP kinase activation and forms a complex with Ras and Raf. Genes Dev. 14, 895–900 (2000).
Umstead, M., Xiong, J., Qi, Q., Du, Y. & Fu, H. Aurora kinase A interacts with H-Ras and potentiates Ras-MAPK signaling. Oncotarget 8, 28359–28372 (2017).
Dance, M., Montagner, A., Salles, J. P., Yart, A. & Raynal, P. The molecular functions of Shp2 in the Ras/Mitogen-activated protein kinase (ERK1/2) pathway. Cell. Signal. 20, 453–459 (2008).
Maurer, T. et al. Small-molecule ligands bind to a distinct pocket in Ras and inhibit SOS-mediated nucleotide exchange activity. Proc. Natl Acad. Sci. USA 109, 5299–5304 (2012).
Mott, H. R. et al. The solution structure of the Raf-1 cysteine-rich domain: a novel Ras and phospholipid binding site. Proc. Natl Acad. Sci. USA 93, 8312–8317 (1996).
Saraboji, K. et al. The carbohydrate-binding site in galectin-3 is preorganized to recognize a sugarlike framework of oxygens: ultra-high-resolution structures and water dynamics. Biochemistry 51, 296–306 (2012).
Shaw, D. E. et al. Anton 2: raising the bar for performance and programmability in a special-purpose molecular dynamics supercomputer. In Proc. International Conference for High Performance Computing, Networking, Storage and Analysis (SC14) 41–53 (IEEE, 2014).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Joung, I. S. & Cheatham, T. E. III Determination of alkali and halide monovalent ion parameters for use in explicitly solvated biomolecular simulations. J. Phys. Chem. B. 112, 9020–9041 (2008).
Wang, J., Cieplak, P. & Kollman, P. A. How well does a restrained electrostatic potential (RESP) model perform in calculating conformational energies of organic and biological molecules? J. Comput. Chem. 21, 1049–1074 (2000).
Duan, Y. et al. A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 24, 1999–2012 (2003).
Hornak, V. et al. Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins 65, 712–725 (2006).
Best, R. B. & Hummer, G. Optimized molecular dynamics force fields applied to the helix-coil transition of polypeptides. J. Phys. Chem. B. 113, 9004–9015 (2009).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78, 1950–1958 (2010).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B. 114, 7830–7843 (2010).
Khoury, G. A., Thompson, J. P., Smadbeck, J., Kieslich, C. A. & Floudas, C. A. Forcefield_PTM: ab-initio charge and AMBER force field parameters for frequently occurring post-translational modifications. J. Chem. Theory Comput. 9, 5653–5674 (2013).
Steinbrecher, T., Latzer, J. & Case, D. A. Revised AMBER parameters for bioorganic phosphates. J. Chem. Theory Comput. 8, 4405–4412 (2012).
Meagher, K. L., Redman, L. T. & Carlson, H. A. Development of polyphosphate parameters for use with the AMBER force field. J. Comput. Chem. 24, 1016–1025 (2003).
Robustelli, P., Piana, S. & Shaw, D. E. Developing a molecular dynamics force field for both folded and disordered protein states. Proc. Natl Acad. Sci. USA 115, E4578–E4766 (2018).
Abascal, J. L. F. & Vega, C. A general purpose model for the condensed phases of water: TIP4P/2005. J. Chem. Phys. 123, 234505 (2005).
Martyna, G. J., Tobias, D. J. & Klein, M. L. Constant pressure molecular dynamics algorithms. J. Chem. Phys. 101, 4177–4189 (1994).
Lippert, R. A. et al. Accurate and efficient integration for molecular dynamics simulations at constant temperature and pressure. J. Chem. Phys. 139, 164106 (2013).
Lippert, R. A. et al. A common, avoidable source of error in molecular dynamics integrators. J. Chem. Phys. 126, 046101 (2007).
Kräutler, V., Van Gunsteren, W. F. & Hünenberger, P. H. A fast SHAKE algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations. J. Comput. Chem. 22, 501–508 (2001).
Shan, Y., Klepeis, J. L., Eastwood, M. P., Dror, R. O. & Shaw, D. E. Gaussian split Ewald: a fast Ewald mesh method for molecular simulation. J. Chem. Phys. 122, 054101 (2005).
Tuckerman, M., Berne, B. J. & Martyna, G. J. Reversible multiple time scale molecular dynamics. J. Chem. Phys. 97, 1990–2001 (1992).
Zachowski, A. Phospholipids in animal eukaryotic membranes: transverse asymmetry and movement. Biochem. J. 294, 1–14 (1993).
Van Meer, G., Voelker, D. R. & Feigenson, G. W. Membrane lipids: where they are and how they behave. Nat. Rev. Mol. Cell Biol. 9, 112–124 (2008).
Lu, J. et al. KRAS G12C drug development: discrimination between switch II pocket configurations using hydrogen/deuterium-exchange mass spectrometry. Structure 25, 1442–1448 (2017).
Gureasko, J. et al. Membrane-dependent signal integration by the Ras activator Son of sevenless. Nat. Struct. Mol. Biol. 15, 452–461 (2008).
Shen, Q. T. et al. Bowl-shaped oligomeric structures on membranes as DegP’s new functional forms in protein quality control. Proc. Natl Acad. Sci. USA 106, 4858–4863 (2009).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Gasper, R., Meyer, S., Gotthardt, K., Sirajuddin, M. & Wittinghofer, A. It takes two to tango: regulation of G proteins by dimerization. Nat. Rev. Mol. Cell Biol. 10, 423–429 (2009).
Egea, P. F. et al. Substrate twinning activates the signal recognition particle and its receptor. Nature 427, 215–221 (2004).
Shima, F. et al. Structural basis for conformational dynamics of GTP-bound Ras protein. J. Biol. Chem. 285, 22696–22705 (2010).
Bum‐Erdene, K. et al. Investigation into the feasibility of thioditaloside as a novel scaffold for Galectin‐3‐specific inhibitors. ChemBioChem 14, 1331–1342 (2013).
Ismail, S. A. et al. Arl2-GTP and Arl3-GTP regulate a GDI-like transport system for farnesylated cargo. Nat. Chem. Biol. 7, 942–949 (2011).
Hoffman, G. R., Nassar, N. & Cerione, R. A. Structure of the Rho family GTP-binding protein Cdc42 in complex with the multifunctional regulator RhoGDI. Cell 100, 345–356 (2000).
Acknowledgements
We thank M. Eastwood for helpful discussions, P. Ayaz for a critical reading of the manuscript, K. Yuh for assistance with videos, J. McGillen and B. Frank for editorial assistance, and C. Scholl, J.V. Schaefer and J. Binz for their support with the BRET experiments. This work was supported in part by a Stand Up To Cancer (SU2C)–American Cancer Society Lung Cancer Dream Team Translational Research Grant (no. SU2C-AACR-DT17-15 to P.A.J.), the Gadzooks Fund (to P.A.J.), the Cammarata Family Foundation Research Fund (to P.A.J.), the Giovanni Armenise–Harvard Foundation, the Lung Cancer Research Foundation, the European Research Council under the European Union’s Horizon 2020 research and innovation program (grant agreement no. 101001288) (to C.A.), the US Department of Defense (W81XWH-16-1-0106), Hale Center and Pinard Family (DFCI), the National Institutes of Health (NIH 5R01CA244341), the Cancer Prevention and Research Institute of Texas (RP170373) (to K.D.W.) and Krebsliga Schweiz grants (nos. KFS 4147-02-2017 and KFS-5290-02-2021-R) (to A.P.). SU2C is a program of the Entertainment Industry Foundation. Research grants are administered by the American Association for Cancer Research, the scientific partner of SU2C. P.A.J. has served as a consultant for and has received sponsored research funding from AstraZeneca, Mirati Therapeutics and Araxes Pharmaceuticals.
Author information
Authors and Affiliations
Contributions
Y.S. conceived this study and, with D.E.S., oversaw molecular dynamics simulations and analysis. V.P.M., Q.W. and M.R.T. performed and analyzed molecular dynamics simulations. Y.S. and V.P.M. designed molecular dynamics simulations, built the structural model of the signalosome and, with K.D.W., C.A., P.A.J. and J.N.K., selected mutations for experimental testing. K.D.W. and Z.W.Z. designed and analyzed FRET assays. Z.W.Z. performed the FRET assays. C.A. designed and analyzed cell proliferation and western blot assays. C.A. and J.J.O. performed these assays. J.N.K. and A.P. designed and analyzed BRET assays. J.N.K and G.N.-D. performed these assays. K.D.W., X.B., L.L. and C.L. designed and analyzed negative-stain assays. L.L. and C.L. performed these assays. Q.W., Y.S. and V.P.M. designed and analyzed nucleotide-exchange assays. Y.S., V.P.M., K.D.W., C.A., P.A.J. and D.E.S. wrote the paper. Y.S., C.A., K.D.W., P.A.J., A.P. and D.E.S. supervised the overall research.
Corresponding authors
Ethics declarations
Competing interests
P.A.J. reports grants from AstraZeneca, Boehringer Ingelheim, Daiichi Sankyo, Eli-Lilly, Takeda Oncology, Astellas, PUMA and Revolution Medicines and personal fees from AstraZeneca, Boehringer Ingelheim, Pfizer, Roche/Genentech, Eli-Lilly, Chugai, Ignyta, Loxo Oncology, SFJ Pharmaceuticals, Voronoi, Daiichi Sankyo, Biocartis, Novartis, Sanofi Oncology, Takeda Oncology, Mirati Therapeutics, Trasncenta, Silicon Therapeutics, Syndax, Bayer, Esai, Allorion Therapeutics, Accutar Biotech and AbbVie outside the submitted work; P.A.J. also receives postmarketing royalties from a DFCI-owned patent on EGFR mutations issued and licensed to Lab Corp. K.D.W. receives or has received grant funding from Revolution Medicines and Astellas outside the submitted work. He also is on the scientific advisory board for Vibliome Therapeutics. The remaining authors declare no competing interests.
Additional information
Peer review information Nature Structural & Molecular Biology thanks John Hancock, Francesco Gervasio and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Anke Sparmann and Florian Ullrich were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Construction of the Ras-Raf signalosome model in 16-protomer form.
a. Top and side views of the stepwise addition (left to right) of Gal-3 (yellow), C-Raf (green), 14-3-3σ (blue), and MEK1 (violet) to the K-Ras helical assembly (red), producing the 16-protomer signalosome model. The membrane is shown as a mesh. Each C-Raf KD dimer binds to a 14-3-3σ dimer and two MEK1 kinases. b. The Cα atom RMSD (w.r.t. the starting structure) of a membrane-anchored 8-protomer Ras-Raf signalosome and its K-Ras octamer core in a 100-μs simulation. As shown, the signalosome—especially the K-Ras core—was stable in the course of the simulation. c. Similar to B, the Cα atom RMSDs of a membrane-anchored 8-protomer Ras-Raf signalosome without Gal-3 at the base tier.
Extended Data Fig. 2 The GTP-mediated K-Ras dimer model and its orientation on the membrane.
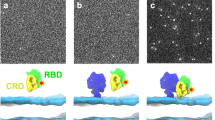
a. Left panel: A collection of snapshots of the 20 simulations starting from two separated K-Ras proteins in solvent. The snapshots are aligned to one K-Ras protein therein (cyan). The GMA dimer model is highlighted. The α4 helix is shown in cartoon representation. Snapshots 10 µs apart were taken from the 20 simulations (which have an aggregate simulation time of 680 µs). Right panel: A collection of snapshots of the 23 simulations of two GTP-bound K-Ras proteins on the membrane; snapshots 10 µs apart were taken from the 23 simulations (which have an aggregate simulation time of 210 µs). Only in one solvent and one membrane simulation was the GMA dimer reached, and in both cases the dimer remained stable. b. A stable dimer (orange) generated from one of the 20 simulations of K-Ras/K-Ras association in solvent, which is consistent with the crystal structures of Ras symmetric dimer using the α4-α5 interface (for example PDB 1QRA, cyan and green). c. Upper panel: Acceptor (yellow) interaction with the γ-phosphate of the GTP donor (cyan) in a simulation of a GMA K-Ras dimer. The Arg135 of the acceptor interacts with the γ-phosphate, while the Lys128 interacts with the Glu62 of the donor, and vice versa. Snapshots at 40 μs and 60 µs from the simulation of K-Ras dimerization on the membrane are shown; the motions of Arg135 and Lys128 are shown by dashed red arrows. Lower panel: Distances from Arg135 (black) and Lys128 (red) to the γ-phosphate. d. Comparison of GTP interactions and GMPPNP interactions with the acceptor residues at the GMA dimer interface. e. Interface residues of the GMA dimer (left) and water occupancy map (right, blue) at the interface. f. Snapshots at 10 μs from four independent runs of free GTP-bound K-Ras monomer (cyan) on the membrane. To illustrate the relationship between the membrane orientation and RBD binding of K-Ras, the C-Raf RBD (green) is positioned on each snapshot based on the Ras-RBD structure (PDB 4G0N). Shown are the distance of the β phosphorus (βP) and the amine nitrogen of the guanine ring (N2) from the plane of the membrane surface; we use these distances to describe the membrane orientation of a Ras protein (Fig. 2g). The Run 1 snapshot is compatible with RBD binding, which corresponds to the Ras membrane orientation of the lower red contours in Fig. 2g in the main text. The snapshot from Run 2 corresponds to the right center of the top red contours; this orientation of K-Ras on the membrane positions the RBD away from the membrane. In the Run 3 and Run 4 snapshots, RBD binding leads to a severe steric clash with the membrane; this corresponds to the left center of the top red contour. g. Snapshot at 100 μs from the simulation of the GMA dimer formation on the membrane, with two C-Raf RBDs (green) added based on the Ras-RBD pose; the RBDs do not clash with the membrane. The membrane orientation of the donor and acceptor Ras proteins correspond to the lower and upper black contours, respectively, in Fig. 2g in the main text. h. Examples of G nucleotide–mediated dimerization, in which an arginine interacts with a nucleotide phosphate at the dimerization interface. The Toc34 homodimer114 (PDB 3BB1), the Ffh-FtsY heterodimer115 (PDB 1RJ9), and the adenylosuccinate synthetase homodimer (PDB 4M9D) are compared to the GMA K-Ras dimer model (cyan); the host G proteins (not shown) were aligned in this comparison. i. The binding site of the synthetic monobody NS119 (red) and the donor site (cyan) of an acceptor in the GMA dimer are shown on a Ras protein (yellow). The two binding sites largely overlap. j. The electrostatic complementarity at the GMA dimer interface, with contacting areas at the interface connected by dashed lines. k. Cα RMSD of the GMA dimer of the wild type and various oncogenic mutants in simulations. l. Sequence conservation of K-Ras proteins in various species (see Materials and Methods). A graphical representation of sequence alignment of representative K-Ras proteins in evolution. The residues involved in the K-Ras/K-Ras interactions in the K-Ras helical assembly are labeled. The 32 K-Ras sequences by GenBank IDs are: OLS17184.1, OLS30914.1, OLS23071.1, XP_020603162.1, XP_023347765.1, XP_023347766.1, XP_003378992.1, XP_003377451.1, GBC14959.1, XP_027484897.1, QBM87817.1, XP_013758736.1, EHB13737.1, OQV11758.1, XP_027289840.1, NP_001356715.1, XP_027290137.1, XP_027290057.1, XP_027289839.1, XP_027289566.1, XP_029436565.1, ELK24704.1, XP_027290375.1, XP_023664743.1, XP_021777811.1, XP_021247616.1, XP_025064324.1, XP_006127724.1, NP_001243091.1, RDD38924.1, KPM02442.1, OTF69949.1.
Extended Data Fig. 3 Stability of the active switch I and switch II conformations.
a. Distributions of the switch I conformation of K-Ras in various simulations. Normalized histograms of the switch I backbone root-mean-square deviation (RMSD) with respect to the active switch I conformation in PDB 4DSN are shown. b. Conformations of the switch I (residues 30–38, cyan) and switch II (residue 60–76, yellow) regions in simulations of the GTP-bound K-Ras monomer. c. Conformations of the switch I and II regions in simulations of GDP-bound K-Ras monomer. d. Conformations of the switch I and II regions in simulations of GTP-bound K-Ras bound with C-Raf RBD. e. Crystal structures of Ras bound with GTP or GTP analogs, with the switch I and switch II regions highlighted. The PDB entries included are 1AGP, 1CLU, 1CTQ, 1GNP, 1GNR, 1HE8, 1JAH, 1JAI, 1K8R, 1LF0, 1LFD, 1NVU, 1NVV, 1NVW, 1NVX, 1P2S, 1P2T, 1P2U, 1P2V, 1PLJ, 1PLK, 1QRA, 1RVD, 1ZW6, 2C5L, 2RGA, 2RGB, 2RGC, 2RGD, 2RGE, 2RGG, 2UZI, 2VH5, 3DDC, 3GFT, 3I3S, 3K8Y, 3L8Y, 3L8Z, 3LBH, 3LBI, 3LBN, 3OIU, 3OIV, 3OIW, 3RRY, 3RRZ, 3RS0, 3RS2, 3RS3, 3RS4, 3RS5, 3RS7, 3RSO, 3TGP, 3V4F, 4DLR, 4DLS, 4DLT, 4DLU, 4DLV, 4DLW, 4DLX, 4DLY, 4DLZ, 4DSN, 4DSO, 4DST, 4EFL, 4EFM, 4EFN, 4G0N, 4G3X, 4K81, 4L9W, 4NMM, 4NYI, 4NYJ, 4NYM, 4RSG, 4XVQ, 4XVR, 5B2Z, 5B30, 5P21, 6Q21, 121 P, 421 P, 521 P, 621 P, 721 P, 821 P, and 221 P. f. Crystal structures of Ras loaded with GDP. The PDB entries included are 1AA9, 1CRP, 1CRQ, 1CRR, 3LO5, 1IOZ, 1LF5, 1PLL, 1Q21, 1WQ1, 1XD2, 1XJ0, 1ZVQ, 2CE2, 2CLD, 2Q21, 2QUZ, 2X1V, 3CON, 3KUD, 4DSU, 4EPR, 4EPT, 4EPV, 4EPW, 4EPX, 3EPY, 4L8G, 4L9S, 4LDJ, 4LPK, 4LRW, 4LUC, 4LV6, 4LYF, 4LYH, 4LYJ, 4M1O, 4M1S, 4M1T, 4M1W, 4M1Y, 4M21, 4M22, 4OBE, 4PZY, 4PZZ, 4Q01, 4Q02, 4Q03, 4Q21, 4QL3, 4TQ9, 4TQA, 4WA7, and 5F2E. g. Crystal structures of the T35S Ras mutant loaded with GTP or GTP analog, with the switch I and switch II regions highlighted. The PDB entries included are 1IAQ, 2LCF, 2LWI, 3KKM, and 3KKN. The hydroxyl of Thr35 in the switch I region in wild-type Ras coordinates the Mg2 + ion bound to the GTP, and the T35S mutation is known to disrupt the switch I active conformation. The switch I inactive conformations sampled by the simulations (Supplementary Fig. 3B and C) are broadly consistent with the inactive conformations in T35S structures116. h. Conformations of the switch I and II regions in simulations of 24 copies of GTP-bound K-Ras in a crystal lattice (of PDB 3GFT). i. The Cα atom RMSDs of a GMA K-Ras tetramer (w.r.t. the starting structure) in a 10-μs simulation with membrane and in a 10-μs simulation in water solvent. As shown, the membrane helps stabilize the tetramer structure.
Extended Data Fig. 4 Cellular validation of the K-Ras signalosome interfaces.
Panels A–C here refer to the experiments testing mutations at the GMA dimer interface, and panels D–F refer to the experiments testing secondary/tertiary K-Ras interfaces. a. Representative cell images at the endpoint of the experiment testing mutations at the GMA dimer interface (Fig. 3f); scale bar: 50 µm. b. Growth rates of K-Raslox/K-RASMUT cells expressing the indicated mutations in cis with either G12C or G12D mutations in 10%, 1%, and 0.5% FBS medium, represented by the confluence value assessed by IncuCyte. Representative pictures at the endpoint are shown in the bottom panels (scale bar: 50 µm). Data are shown as mean +/− standard deviation (n = 3 biologically independent experiments). c. Phosphorylation of ERK and AKT in K-Raslox/K-RASMUT cells expressing the indicated mutations in cis with either G12C or G12D mutations. Cells were lysed after 48 hours incubation in 0.5% or 10% FBS, as indicated, and analyzed by Western blot. These results are representative of three independent experiments with similar results. d. Representative cell images at the endpoint of the experiment testing mutations at the secondary and tertiary K-Ras/K-Ras interfaces (Fig. 6b); scale bar: 50 µm. e. Phosphorylation of ERK and AKT in K-Raslox/K-RASMUT cells expressing the indicated mutations in cis with either G12C or G12D mutations. Cells were lysed after 48 hours incubation in 0.5% or 10% FBS as indicated and analyzed by Western blot. These results are representative of three independent experiments with similar results. f. Growth rates of K-Raslox/K-RASMUT cells expressing the indicated mutations in cis with either G12C or G12D mutations to test the secondary and tertiary K-Ras/K-Ras interfaces in the background of either G12C or G12D, shown as confluence values measured by IncuCyte. Representative images at the end point are also shown. Cells were kept in 0.5% or 10% FBS. Data are shown as mean +/− standard deviation (n = 3 biologically independent experiments).
Extended Data Fig. 5 FRET and BRET validation of K-Ras signalosome interfaces.
a. CFP emission of wild-type K-Ras and various K-Ras mutants to test the K-Ras/K-Ras GMA dimer. HEK293T cells were co-transfected with CFP-K-Ras and YFP-K-Ras, serum starved, and stimulated with 10% FBS. Each K-Ras construct is labeled under its respective plot. Assays were repeated three times. Error bars represent mean ± S.E.M. WT: n = 42; D154Q: n = 33; G12C/D154Q: n = 35; G12D/D154Q: n = 20; D30R: n = 32; G12C/D30R: n = 30; G12D/D30R: n = 28; E31R: n = 42; G12C/E31R: n = 26; G12D/E31R: n = 28; E62R: n = 33; G12C/E62R: n = 32; G12D/E62R: n = 22; K147D: n = 36; G12C/K147D: n = 34; G12D/K147D: n = 38; K147D/D154Q: n = 29. (**** denotes P < 0.0001 by two-way ANOVA; ns stands for not significant). b. CFP emission of wild-type K-Ras and T35 and G60 mutants to test K-Ras association. Assays were repeated three times. Error bars represent mean ± S.E.M. WT: n = 14; T35A: n = 18; T35S: n = 11; G60A: n = 17. (P value was calculated by two-way ANOVA; ns stands for not significant). c. CFP emission of wild-type K-Ras and T35 and G60 mutants with C-Raf to monitor Ras-Raf binding at 10% FBS or 10 ng mL−1 EGF. Assays were repeated three times. Error bars represent mean ± S.E.M. T35A (FBS): n = 36; T35A (No FBS): n = 27; T35A (No FBS + EGF): n = 21; G60A (FBS): n = 55; G60A (No FBS): n = 38; G60A (No FBS + EGF): n = 37; WT (FBS): n = 12; WT (No FBS): n = 37; WT (No FBS + EGF): n = 11. (**** denotes P < 0.0001 by two-way ANOVA; ns stands for not significant). d. Nucleotide exchange assay of wild-type K-Ras with (orange) and without (gray) SOS1, and R135A without SOS1 (blue). e. CFP emission of wild-type K-Ras and R135A mutants. Assays were repeated three times. Error bars represent mean ± S.E.M. In 10% FBS condition, WT: n = 44; R135A: n = 82; G12C/R135A: n = 50; G12D/R135A: n = 34; D154Q/R135A: n = 51; D154Q: n = 33. In 0.5% FBS condition, WT: n = 25; R135A: n = 47; G12C/R135A: n = 44; G12D/R135A: n = 34; D154Q/R135A: n = 42; D154Q: n = 26. (* denotes P < 0.05, ** denotes P < 0.01, *** denotes P < 0.001, **** denotes P < 0.0001 by two-way ANOVA; ns stands for not significant). f. BRET signal as an indicator of K-Ras assembly for G12C and G13D mutants, and for the K-Ras construct lacking the membrane-anchoring HVR tail (residues 1–166). Co-transfection of increasing ratios of donor and acceptor plasmids enables discrimination between specific and non-specific (random collision) protein-protein interactions. G. CFP emission of wild-type K-Ras and K88D, Q129L, and R149D mutants. Assays were repeated three times. Error bars represent mean ± S.E.M. In 0.5% FBS condition, WT: n = 43; K88D: n = 38; Q129L: n = 34; R149D: n = 37; G12C: n = 22; G12C/K88D: n = 29; G12C/Q129L: n = 34; G12C/R149D: n = 23; G12D: n = 37; G12D/K88D: n = 34; G12D/Q129L: n = 44; G12D/R149D: n = 38. In 10% FBS condition, WT: n = 40; K88D: n = 35; Q129L: n = 40; R149D: n = 42; G12C: n = 27; G12C/K88D: n = 41; G12C/Q129L: n = 46; G12C/R149D: n = 57; G12D: n = 39; G12D/K88D: n = 67; G12D/Q129L: n = 49; G12D/R149D: n = 42. (* denotes P < 0.05, ** denotes P < 0.01, *** denotes P < 0.001, **** denotes P < 0.0001 by two-way ANOVA; ns stands for not significant).
Extended Data Fig. 6 Structural details of secondary and tertiary Ras-Ras and Ras-RBD interactions.
a. Key contacts of the secondary (stacking) Ras-Ras interaction of K-Ras n (blue) and K-Ras n + 4 (green). A series of salt bridges form intermittently in MD simulations between the Lys88, Glu91, Glu98, Arg102, and Asp105 residues on the α3 helix of K-Ras n and the Asp154, Arg161, Lys165, Glu168, and Lys172 residues, respectively, on the α5 helix of K-Ras n + 4. b. Key contacts at the interface of K-Ras n + 3 (purple) and K-Ras n (blue). c. Key contacts of the secondary Ras-RBD interaction of C-Raf RBD n (yellow) with K-Ras n + 1 (green), which is the GTP acceptor of K-Ras n (purple), the primary binding partner of the RBD. The RBD also interacts with the HVR of K-Ras n + 1. Residues 1–53 of C-Raf are structurally unresolved but are likely to make additional interactions at this secondary Ras-RBD interface. d. Key contacts of the tertiary Ras-RBD interaction of Raf RBD n + 3 (yellow) and K-Ras n (blue). e. Unoccupied Switch-II pocket in a crystal structure of K-Ras (PDB 4LDJ) compared with the Switch-II pocket of K-Ras n in the stacking interaction. f. Superposition of stacking (n and n + 4) K-Ras proteins from the signalosome model with a crystal dimer of K-Ras (PDB 5UQW); in both cases the dimer interaction is mediated by a β2- β3 turn inserting into a Switch-II pocket. G. Comparison of D154Q/R161E with the WT in K-Ras oligomer simulations. As shown, the double mutation disrupts the D154-K147 salt bridge at the GMA dimer (n/n-1), and as compensation, allows an E161-R102 salt bridge in stacking (n/n + 4) by shifting the relative position of the α3 and α5 helices.
Extended Data Fig. 7 Structural details of interactions of K-Ras, Gal-3 and the C-Raf CRD.
a. Gal-3 n (olive) binds to the HVR fCys185 of Ras n (blue) and to C-Raf RBD n − 1 (orange). The thiodigalactoside inhibitor binding site of Gal-3117 (PDB 4JC1) is shown in red. b. Gal-3 stacking; Gal-3 n (green) and Gal-3 n + 4 (yellow) are stacked in parallel with K-Ras stacking. c. Position of Gal-3 (olive) with respect to K-Ras (blue), in comparison with that of PDE6δ (orange) with respect to farnesylated K-Ras61 (PDB 5TAR), PDE6δ (yellow) with respect to farnesylated Rheb118 (PDB 3T5G), and RhoGDI (cyan) with respect to geranylgeranylated CDC42119 (PDB 1DOA). The Gal-3 position somewhat resembles the PDE6δ complex in the overall pose. The insert shows the simulation-generated Gal-3 binding pose of fCys with the Gal-3 crystal structure (PDB 3ZSM) overlaid for comparison. d. Left: snapshots of the simulation in which fCys enters the Gal-3 (yellow) pocket, with color coding on fCys indicating simulation time; right: the final Gal-3/fCys model superimposed on the Gal-3 structure used to initiate the simulation (PDB 3ZSM). e. Final snapshots from 55 simulations of the three-domain system of the CRD (light orange) tethered to a K-Ras-bound RBD (orange). The snapshots were aligned with respect to the K-Ras protein. Of the 55 simulations, 24 were in solvent (80 µs in aggregate), and the other 31 were with the RBD and the membrane (117 μs in aggregate). The simulation-generated CRD poses are grouped into three clusters: 1) clashing with the GMA K-Ras dimer; 2) clashing with the membrane if applied to the GMA K-Ras dimer on the membrane (Fig. 1a); and 3) near the C-terminal of the α5 helix and the β1, β2, and β3 strands of K-Ras. Only cluster 3 is compatible with the helical assembly of K-Ras; cluster 2 is similar to recently reported structures74,75,76. f. Details of the CRD-Ras interface of a CRD pose selected from the third cluster. NMR-identified Ras residues involved in Ras-CRD interaction are shown. The Glu3 residue on the β1 strand of the K-Ras protein interacts stably with a CRD-bound Zn2 + ion (lower right panel). The Lys157(CRD)/Glu76(K-Ras) distance in simulations is shown to indicate the stability of the Ras-CRD pose (lower left panel). The upper right panel shows the relative positions of K-Ras (blue), CRD (orange), Gal-3 (olive), and GTP (red) in a top view. g. CRD residues identified by an NMR study51 as part of the K-Ras interface. K-Ras M170, fCys185, and CRD Zn2 + ions are also shown. h. The surfaces of RBD-bound K-Ras and CRD colored by their electrostatic potential (calculated using PyMOL). A cartoon image is used to show the orientation of the Ras-CRD complex. The CRD-bound Zn2 + ions are shown in purple. I. A C-Raf CRD at the base tier with RBD and K-Ras. Arg143, Lys144, Lys148, and the two coordinating Zn2 + ions of the CRD are shown, together with the occupancy map of POPS lipids in contact with the three amino acids shown in red mesh.
Extended Data Fig. 8 Membrane interface of K-Ras, Gal-3, and C-Raf.
a. Positively charged residues of a base-tier K-Ras protein proximal to the membrane in the signalosome model. The inset shows the direct interaction between the HVR lysines of a K-Ras protein at the base tier and the phosphotidylserine lipids. b. Positively charged residues and prolines of Gal-3 proximal to the membrane at the base tier. A Val126 residue that has been suggested to be involved in Gal-3 membrane interaction23 is also shown. c. Positively charged residues of the C-Raf RBD and CRD and the two CRD-bound Zn2 + ions proximal to the membrane.
Extended Data Fig. 9 Additional interactions between Raf, 14-3-3σ, and MEK1.
a. Superposition of the structural model (gray) of the Raf/MEK/14-3-3 complex with the separately resolved cryo-EM structure (PDB 6Q0J). b. Interaction between the Raf kinase domain dimer (green) and 14-3-3σ dimer (cyan). The linker residues pSer338 and pTyr341 are located between the two αC helices of the kinase dimer, and pSer621 is bound to a 14-3-3σ protein. c. Electrostatic potential surface of the C-Raf kinase domain dimer; the region of the two αC helices, which is where pSer339 and pTyr341 (shown) are located, is highly positively charged. d. Interaction of phosphorylated residues pSer338 and pTyr341 with the positively charged residues of the two αC helices of the C-Raf KD dimer. e. Samples of conformations of the C-Raf linker (rainbow colors) in the Ras-Raf signalosome model. The C-Raf KD (gray) is attached to the linker for reference. f. Top and side views of an alternative eight-protomer signalosome model, wherein C-Raf n and C-Raf n + 1 form a dimer, as opposed to the model shown in Fig. 1d and e, wherein C-Raf n and C-Raf n + 4 form a dimer. The color scheme and protein representation are similar to those in Fig. 1d. The inset shows that in this alternative model a C-Raf linker (linker 2) has to circumvent a Gal-3 protein (Gal-3 2) to engage the other C-Raf linker in a C-Raf dimer. G. An illustration of how the GEF domain of SOS1 can reach the K-Ras 8. The modeling was based on PDB entry 1NVU for the K-Ras/SOS1 pose, and PDB entry 1XD4 for the structures of the PH domain and the PH-GEF linker; the linker orientation with respect to the PH and the GEF domains was manually adjusted and energetically minimized in the modeling. The helical hairpin of SOS1 is shown in yellow.
Supplementary information
Supplementary Information
Supplementary Note, Analyses and Table 1.
Supplementary Video 1
The architecture of the Ras–Raf signalosome model. Illustration of the 16-protomer signalosome, in which a 16-member helical assembly of K-Ras on the membrane was assembled, followed by sequential addition of Gal-3, the RBD, CRD, linker and kinase domain of C-Raf, the 14-3-3σ dimer, and finally MEK1 kinase. Colors are consistent with those in Fig. 1d.
Supplementary Video 2
Simulation of K-Ras dimer formation in solvent. An unbiased simulation starting with two spatially separated GTP-bound K-Ras proteins in solvent, in which the GMA dimer formed spontaneously at approximately 16 µs. The simulation time is marked. The GTP donor is colored in cyan and the GTP acceptor is colored in pink. At the end of the video, this dimer is superimposed with the GMA dimer that was generated by simulating K-Ras dimerization on a membrane.
Supplementary Video 3
Simulation of K-Ras dimer formation on a membrane. An unbiased simulation starting with two spatially separated GTP-bound K-Ras proteins on a membrane, in which the GMA dimer formed spontaneously at approximately 6.5 µs. The simulation time is marked. The GTP donor is colored in cyan and the GTP acceptor is colored in pink. The membrane is shown in gray in the background; GTP molecules are also shown.
Supplementary Dataset 1
Atomic coordinates of the structural model of the eight-protomer Ras–Raf signalosome with K-Ras (chains A, B, P, Q, W, Y, E, G), C-Raf (chains C, D, R, S, X, Z, F, H), Gal-3 (chains N, O, V, U, I, J, K, T), 14-3-3σ (chains n, o, v, u, i, j, k, t) and MEK1 (chains c, d, r, s, x, z, f, h).
Supplementary Dataset 2
Atomic coordinates of the GMA K-Ras dimer on a membrane.
Source data
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Unprocessed western blots and micrographs.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Rights and permissions
About this article
Cite this article
Mysore, V.P., Zhou, ZW., Ambrogio, C. et al. A structural model of a Ras–Raf signalosome. Nat Struct Mol Biol 28, 847–857 (2021). https://doi.org/10.1038/s41594-021-00667-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-021-00667-6
This article is cited by
-
Genetic variants of SOS2, MAP2K1 and RASGRF2 in the RAS pathway genes predict survival of HBV-related hepatocellular carcinoma patients
Archives of Toxicology (2023)
-
SHOCing RAF into action
Nature Structural & Molecular Biology (2022)
-
Identification of a BRAF/PA28γ/MEK1 signaling axis and its role in epithelial-mesenchymal transition in oral submucous fibrosis
Cell Death & Disease (2022)
-
Structure–function analysis of the SHOC2–MRAS–PP1C holophosphatase complex
Nature (2022)
-
Building insights into KRAS signaling complexes
Nature Structural & Molecular Biology (2021)