Abstract
The spike (S) protein of SARS-CoV-2 mediates receptor binding and cell entry and is the dominant target of the immune system. It exhibits substantial conformational flexibility. It transitions from closed to open conformations to expose its receptor-binding site and, subsequently, from prefusion to postfusion conformations to mediate fusion of viral and cellular membranes. S-protein derivatives are components of vaccine candidates and diagnostic assays, as well as tools for research into the biology and immunology of SARS-CoV-2. Here we have designed mutations in S that allow the production of thermostable, disulfide-bonded S-protein trimers that are trapped in the closed, prefusion state. Structures of the disulfide-stabilized and non-disulfide-stabilized proteins reveal distinct closed and locked conformations of the S trimer. We demonstrate that the designed, thermostable, closed S trimer can be used in serological assays. This protein has potential applications as a reagent for serology, virology and as an immunogen.
Similar content being viewed by others
Main
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the virus responsible for the 2019 coronavirus disease (COVID-19) pandemic1,2. SARS-CoV-2 belongs to the betacoronavirus family and is closely related to SARS-CoV-1, which was responsible for the 2003 SARS epidemic3. Similar to other coronaviruses, the spike (S) protein is the major glycoprotein on the SARS-CoV-2 virus surface. S protein is responsible for receptor binding and subsequent virus uptake and fusion with the target cell, and is a dominant target of the immune system4. S protein is trimeric and has two distinct structural states—prefusion and postfusion4. Recognition by the immune system of the prefusion state displayed on the virus surface is crucial to mount an effective immune response5.
S-protein trimers of SARS-CoV-1 and Middle East respiratory syndrome (MERS)-CoV are unstable in their metastable prefusion state6,7. Stimuli including protease digestion8,9, receptor binding and antibody binding8 can promote the structural transition of S into the postfusion state. The SARS-CoV-2 S protein exhibits similar instability, and S trimers on the surface of the SARS-CoV-2 virion or solubilized by detergent undergo a structural transition into the postfusion state during purification10,11. Successful immunization strategies require stable antigens, and instability of the prefusion state of enveloped virus fusion proteins presents a challenge for vaccine development. For human immunodeficiency virus-1 (HIV-1), introduction of the SOSIP mutations (a disulfide bond and a stabilizing proline) into HIV-1 glycoprotein GP16012, and for paramyxoviruses, introduction of disulfide bonds and cavity-filling mutations into the fusion protein F, were required to stabilize these proteins in their prefusion state for structural or vaccinology studies5,13. Attempts have been made to stabilize SARS-CoV-1 and MERS-CoV S proteins in their prefusion state through the introduction of two prolines within S2 to prevent central α-helix extension, a structural change needed for transition into the postfusion state6,7. Here, we have found that such stabilization does not maintain coronavirus S proteins in the prefusion form for a prolonged period of time.
Cryo-electron microscopy (cryo-EM) studies have shown that the fusion core of the S proteins of SARS-CoV-1 and MERS-CoV is crowned by the receptor binding domain (RBD), surrounded by the N-terminal domains (NTDs) to suppress fusion5. Both NTD and RBD exhibit some flexibility, but the major conformational dynamics are between a closed and an open conformation. In the closed conformation, the RBD is bound in trans into a pocket formed by the NTD and RBD of the neighboring monomer, and the receptor binding site is largely occluded. In the open conformation, the RBD points upwards and the receptor binding site is exposed14,15. Similar RBD conformational dynamics has been observed for the SARS-CoV-2 S protein16,17. It has been suggested that such conformational dynamics is related to both receptor binding and subsequent fusogenic activity14,15,18. Binding of receptor or antibody to the open conformation appears to stimulate the transition of S into the postfusion state8,19. In addition, RBD dynamics is modulated by protease digestion, where S proteins are cleaved into S1 and S2 subunits; this also appears to stimulate the transition to the postfusion state8,9.
Based on this understanding of the S-protein fusion mechanism, we sought to stabilize the S protein in the prefusion state by trapping it in the closed form using engineered disulfide bonds. Our motivation for doing so was fourfold: (1) for use in vaccinology—a stabilized trimer may elicit an improved immune response as part of a vaccine, and stabilized vaccines may be more easily distributed; (2) to induce different immune responses—a closed trimer can be expected to lead to production of an antibody repertoire different from that raised against open S proteins (a successful SARS-CoV-2 vaccine will probably need to induce a broad immune response, and a successful immunotherapy may also require a combination of antibodies); (3) for diagnostics—a more thermostable trimer could enable better diagnostics in regions with poor infrastructure, and a closed trimer can be used to distinguish different immune responses in diagnostic assays (understanding the functional relevance of different antibody responses is critical to establish meaningful assays for protective immunity); (4) to support basic research, including structural biology. Here we have designed an exclusively closed trimeric S protein that is stable to incubation at 60 °C and to prolonged storage.
Results
Construct design
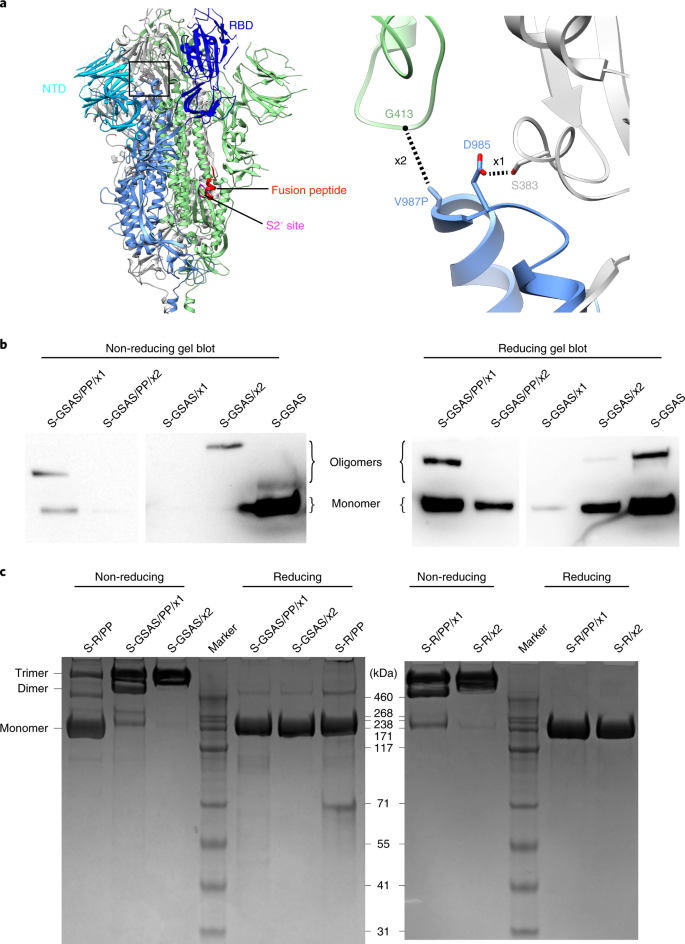
Based on published structures of the trimeric S protein17, we identified two amino acid pairs at an appropriate distance for disulfide bond formation, and in positions where we expected insertion of a covalent bond to stabilize the closed form of the trimer. These pairs, residues 383 and 985 (disulfide 1, x1) and residues 413 and 987 (disulfide 2, x2), are illustrated in Fig. 1a. Starting with the previously published stabilized trimeric ectodomain constructs in which the S1/S2 cleavage site (residues 682–685) was replaced by GSAS and two stabilizing prolines were inserted at positions 986 and 987 (S-GSAS/PP)16, we inserted cysteines at the identified positions to generate the constructs S-GSAS/PP/x1 and S-GSAS/PP/x2 (note that in S-GSAS/PP/x2, the second proline at position 987 was replaced by cysteine, Supplementary Fig. 1). Two preprints have reported constructs similar or equivalent to S-GSAS/PP/x1 (refs. 20,21). In addition, we inserted cysteines at the same positions in a related construct in which the residues PRRA had been deleted at the S1/S2 cleavage site, to leave a single arginine resembling the cleavage site in SARS-CoV-1 and SARS-CoV-2 potential progenitor RaTG131 spikes (R), generating S-R/PP/x1 and S-R/PP/x2 (Supplementary Fig. 1). Finally, anticipating that the proximity of the stabilizing prolines at positions 986 and 987 to the closed RBD might affect the RBD conformation, we additionally engineered x1 and x2 into the GSAS and R constructs without the stabilizing proline mutations, generating S-GSAS/x1, S-GSAS/x2, S-R/x1 and S-R/x2 (Supplementary Fig. 1).
a, Schematic of the structure of trimeric S in its closed, prefusion form (PDB 6VXX), illustrating the location of the RBD and the NTD. The box indicates the region shown in the right-hand panel, which illustrates the positions of disulfide bonds x1 and x2. Note that, in PDB 6VXX and other published SARS-CoV-2 S-protein structures, residues K986 and V987 have been mutated to proline. b, Western blots using antibodies against the C-terminal 6×His-tagged S-protein constructs in small-scale expression supernatants under non-reducing and reducing conditions. S-GSAS/PP/x2 and S-GSAS/x1 showed very low expression levels and were not considered further. c, Comparison of non-reducing and reducing Coomassie-stained gels from medium-scale purifications of S-protein constructs. S-GSAS/PP/x1 and S-R/PP/x1 show a mixture of trimers and lower-order oligomers under non-reducing conditions, whereas S-GSAS/x2 and S-R/x2 show trimers under these conditions.
Protein expression and disulfide bond formation
Expression of all GSAS constructs in mammalian Expi293 cells was assessed by western blotting (Fig. 1b). We were able to express S-GSAS/PP/x1 and S-GSAS/x2, but not S-GSAS/PP/x2 or S-GSAS/x1. Expression levels of the cysteine mutants are somewhat lower than those of the initial construct, possibly due to the effect of engineered cysteines on protein folding.
Based on the western blot results, we performed scaled-up expression and purification of S-GSAS/PP/x1 and S-GSAS/x2, as well as S-R/PP/x1 and S-R/x2. Purified proteins were assessed for the presence of the engineered disulfide bond on non-reducing gels. S-GSAS/PP/x1 and S-R/PP/x1 have bands corresponding to trimers, dimers and monomers, whereas S-GSAS/x2 and S-R/x2 run exclusively as trimers (Fig. 1c). Under reducing conditions, only monomers were observed, confirming that the oligomeric states present in the non-reducing gel are due to disulfide bonds.
Stability of disulfide bond mutants
We imaged the proteins by negative-stain EM to assess their oligomeric states (Fig. 2, left column). Freshly purified non-disulfide-stabilized variants, as well as the x1 disulfide bond mutants, show a majority of trimers with some disintegrated oligomers. The x2 disulfide bond mutants, which migrate exclusively as trimers in gels, show almost exclusively trimers by negative-stain EM.
Freshly purified proteins were imaged by negative-stain EM (left column). Non-disulfide-stabilized S proteins appear predominantly as trimers with some smaller material, while S-R/x2 appears almost exclusively as trimers. Following prolonged incubation at 4 °C or heat treatment, non-disulfide-stabilized S proteins disintegrate, while disulfide-stabilized S proteins remain trimeric. Scale bar, 100 nm.
To assess the stability of the proteins, we repeated the negative-stain EM after prolonged storage at 4 °C (Fig. 2, middle columns). Elimination of the furin cleavage site and insertion of the stabilizing prolines are both likely to stabilize the trimers. Nevertheless, non-disulfide-stabilized variants started to disintegrate into disordered structures after one day of storage, and disintegration is mostly complete after three days of storage. We suggest that the disintegration of S trimers is facilitated by RBD opening. In contrast, the disulfide-bonded variants could be stored at 4 °C for at least one month without substantial disintegration of trimers being observed by negative-stain EM.
We further tested the stability of the disulfide bond variants by subjecting them to incubation at 60 °C for 30 min before imaging by negative-stain EM (Fig. 2, right column). We found that non-disulfide-stabilized variants denatured upon exposure to 60 °C, while disulfide-stabilized mutants showed minimal changes, remaining in their trimeric state.
These results indicate that stabilizing S in a closed conformation by insertion of disulfide bonds leads to a substantial increase in protein stability. S-R/x2 was arrested in the prefusion state by insertion of a single disulfide bond to prevent opening, without the insertion of any other point mutations, suggesting that in the absence of a disulfide bond, S-protein denaturation proceeds via transition into the open form.
Structural features of disulfide-stabilized mutants
To understand the increased stability, we imaged S-R/PP/x1 and S-R/x2 by cryo-EM. For comparison we also performed cryo-EM on S-R/PP and S-R expressed and purified under the same conditions. Classification of the cryo-EM data showed that S-R/PP/x1 and S-R/x2 exclusively form trimers with three closed RBDs, suggesting that the designed disulfides have successfully trapped the RBDs in the closed state (Extended Data Fig. 1). As expected, the disulfide bond is clearly resolved in the fully disulfide-bonded S-R/x2, and visible, but less well resolved in the incompletely disulfide-bonded S-R/PP/x1 (Fig. 3a).
a, Cryo-EM densities for S-R/x2 in the closed conformation and for S-R/PP/x1 in both closed and locked conformations. Boxes mark the regions in the right-hand panels, which show the resolved density for the inserted disulfide bonds at the expected positions. b, Comparison of the RBD and NTD conformations in S trimers of locked and closed conformations as viewed from the top of the trimer at the three-fold axis after alignment based on the central S2 helices. Between the locked and closed conformations, the S1 domains show a twist motion (right panel, and compare the left and middle panels). In the locked conformation, the RBD moves closer to the three-fold axis. The position of the NTD is variable. c, Folding of 833–855 into a helix-turn-helix motif locks the RBD in a locked conformation. Left panel: locations of the RBD, 833–855 motif (orange), fusion peptide and S2′ site are colored and indicated. Middle panel: detailed density for the 833–855 motif, viewed from near the fusion peptide towards the RBD (indicated by the eye icon in the left panel). The density shows well-resolved density that was disordered in previous SARS-CoV-1 and SARS-CoV-2 S structures. Right panel: the position of the RBD in the open conformation is aligned onto the locked monomer. In the open conformation, the RBD would clash into the folded 833–855 motif (orange).
Both non-disulfide-stabilized constructs contain closed trimers and trimers with one RBD in the open state (Extended Data Fig. 1). We observed that ~20% of S-R/PP trimers are in the closed state and ~80% have one open RBD (Extended Data Fig. 1). In contrast we found that ~80% of S-R trimers are in the closed conformation and only ~20% have one open RBD. In the closed S-R trimer, negatively charged residues D427 and D428 in the RBD are in proximity to positively charged K986 (Fig. 3a), and may electrostatically stabilize the closed form. This electrostatic attraction is lost by substituting the basic K986 with neutral proline in S-R/PP. We suggest that the widely used PP mutation favors the transition into the open state by abolishing electrostatic interactions between K986 and negatively charged residues in the RBD. The higher fraction of closed trimers present in S-R when compared to S-R/PP provides an explanation for the higher efficiency of disulfide formation seen in S-R/x2 when compared to S-R/PP/x1.
Further sub-classification of the closed conformations revealed that S-R and S-R/x2 adopt a single closed conformation similar to previously reported closed SARS-CoV-2 S structures, with some flexibility in the position of the NTD (residue 1–305) indicated by the lower resolution of this region in the EM density (Extended Data Fig. 2). By contrast, S-R/PP and S-R/PP/x1 variants exist in two distinct closed states—although the majority of trimers are in the previously described closed conformation, some of the closed trimers are in a more tightly closed form (henceforth called the locked conformation) in which the RBDs have twisted to move ~2 Å closer to the three-fold symmetry axis at the tip of the spike (Fig. 3b). The NTD and RBD are better resolved in the cryo-EM density in the locked conformation than in the closed conformation (Extended Data Fig. 2a), suggesting they are less mobile in this conformation.
Intriguingly, the locked and closed forms show two other structural differences. First, the S-R/PP locked conformation has a lipid bound into a recently described lipid-binding pocket in the RBD22 while in the closed conformation the lipid is absent. Second, the region N-terminal to the fusion peptide and S2′ site, residues 833–855, is disordered in the closed conformation (Fig. 3c) and in all SARS-CoV-1 and SARS-CoV-2 S-protein structures published so far. In the locked conformation it has folded into two helices connected by a loop (Fig. 3c). Structural comparison to an open monomer suggests folding of this motif would prevent the motion of the RBD during opening, effectively locking the RBD in a closed conformation (Fig. 3c). Consistent with this hypothesis, a folded motif is present at this position in the structures of S from murine hepatitis virus (MHV)23, a betacoronavirus, and from alpha- (NL63-CoV24), gamma- (IBV, infectious bronchitis virus25) and delta- (porcine deltacoronavirus26) coronaviruses—for these S proteins no open conformation was observed (Extended Data Fig. 3a). We only observed the locked conformation in variants stabilized with PP, but a recent report of full-length, wild-type S trimers purified in detergent micelles11 also described folding of the 833–855 region. This observation suggests that folded 833–855 is a relevant state that is selectively stabilized by the PP mutation. A SARS-CoV-2 variant that has now become the prevalent form globally contains a D614G mutation27. A recent report suggests that the D614G mutation reduces S1 shedding28. Our data suggest that, by abolishing a salt bridge between D614 and K854 within the folded 833–855 motif (Extended Data Fig. 3b), this change could perturb the locked conformation to affect RBD opening and potentially modulate virus receptor binding and membrane fusion activities. We speculate that the locked conformation represents an early intermediate state before RBD can open to bind receptor, but further research will be required to elucidate possible links between RBD position, lipid binding, folding of the 833–855 motif and the 614–854 salt bridge.
Disulfide-stabilized proteins in diagnostic assays
We next compared the performance of S-R/x2 and non-disulfide-stabilized constructs in diagnostic assays. First, we performed ELISA titrations against S-R/x2 for sera from 48 individuals who were seropositive against S-R/PP. A positive response against S-R/x2 was seen for 42 of these sera. There was a clear correlation between the strength of response against S-R/PP and S-R/x2 (Fig. 4a), with the response against S-R/x2 reduced by approximately threefold (Fig. 4b and Extended Data Fig. 4). This reduction presumably reflects the reduced exposure of RBD epitopes in the closed conformation.
a,b, Comparison of immune responses to open and closed conformations of the SARS-CoV-2 spike protein. Serum samples from polymerase chain reaction (PCR)-positive individuals were screened for reactivity against S-R/PP, and any samples scored as positive (n = 48) were subsequently selected for end-point titrations against both S-R/PP and S-R/x2. a, The area under the curve (AUC) was calculated for each seropositive sample using end-point titrations. Paired AUC values for reactivity against S-R/PP and S-R/x2 were plotted for each positive sample. Pearson’s correlation analysis was calculated as r = 0.9599 (95% confidence interval (CI) 0.929–0.977) and R2 of 0.921. The dashed line represents the theoretical line of equality, whereas the solid line represents the best fit using linear regression. Where possible, samples were used for subsequent half-maximum effective concentration (EC50) determination (Extended Data Fig. 4). Data points in gray are those where an EC50 value for both S-R/PP and S-R/x2 could not be determined as one or both did not reach saturation. Colored data points highlight the three representative samples shown in b. b, End-point titration curves against S-R/PP and S-R/x2. In each case, the dashed line represents reactivity against S-R/PP and the solid line represent the reactivity against the closed conformation S-R/x2. OD450, optical density at 450 nm. c,d, Summed mean fluorescence intensity (MFI) in a multiplexed flow cytometry (Luminex) assay to measure binding of immunoglobulin-G (IgG) in sera to S-protein variants and RBD (c). Sera from seven individuals seropositive for SARS-CoV-2 are shown (as different symbols), as well as two negative control sera (points along the x axis). In d, S proteins were incubated at the indicated temperatures before use in the assay.
Second, we performed Luminex-based assessment of the binding of S-GSAS/PP, S-R/PP, S-R, S-R/x2 and isolated RBD to immunoglobulin-G (IgG) in sera from seven other individuals who were seropositive against SARS-CoV-2 antigens (four patients, and three individuals with a history of mild or no symptoms). A strong response was seen for all five constructs against all seven sera (Fig. 4c). S-GSAS/PP and S-R/PP showed the strongest and almost identical responses, S-R showed a slightly lower response and S-R/x2 a reduced response; in the latter two cases, this was presumably due to the reduced exposure of epitopes within the angiotensin-converting enzyme 2 (ACE2)-binding region of the RBD in the closed spike conformation. The response to S-R/x2 is considerably stronger than that to the RBD, suggesting that the closed S trimer displays considerably more epitopes than the RBD alone.
We used the Luminex-based assay to assess the impact of the S-protein conformational state on interactions with antibodies in sera. We incubated S-R/PP and S-R/x2 at 37 °C for 36 h or subjected them to 60 °C for 30 min, before coupling to Luminex beads. We then compared their binding to the same seven sera used above (Fig. 4d). In the case of S-R/PP, we found that exposure to temperature led to a reduction in IgG response for five sera, consistent with the loss of trimeric structure observed by negative-stain EM (Fig. 2). By contrast, S-R/x2, which maintains its structure upon heat treatment, also maintained its antibody interactions with sera for these five sera. Two sera did not show a loss of binding following heat treatment of S-R/PP, and showed increased binding against 60 °C heat-treated S-R/x2. Notably these two sera also had the lowest binding against RBD. We cannot draw conclusions on the serological differences between the individuals based on these data, but speculate that they may contain antibodies against peripheral linear epitopes that become increasingly exposed upon heat treatment.
The behavior of S-R/x2 in diagnostic ELISA and Luminex assays therefore correlated well with its structural behavior. S-R/x2 has a generally lower response than the proline-stabilized S-R/PP, consistent with the fact that it hides epitopes in the RBD. The S-R/x2 response is in most cases not altered by heat treatment, while the response of S-R/PP generally drops following heat treatment, concomitant with structural denaturation. We believe that thermostabilization of S by insertion of x2 may have direct applications in the production of more stable S derivatives for vaccine and diagnostic purposes.
Discussion
The pattern of immune response against SARS-CoV-2 is not yet fully understood and varies between individuals. In this context, an exclusively closed trimer with excellent stability like S-R/x2 has a number of potential applications. First, although we saw a correlation between response against S-R/PP and S-R/x2, there is variation in the relative responses (Extended Data Fig. 4) that probably reflects different antibody compositions in the sera. A recent preprint demonstrates that some monoclonals bind differently to an open trimer when compared to a closed trimer similar to our construct S-GSAS/PP/x120. Exclusively closed trimers may therefore prove useful as a serological reagent to assay differential immune responses.
Second, while SARS-CoV-2 neutralizing antibodies targeting alternate epitopes have been identified29, the majority of neutralizing monoclonals identified so far act by blocking the interaction between the RBD and the receptor ACE230,31,32. Antibodies that recognize the ACE2 binding site or cryptic epitopes in the open trimer can functionally mimic the receptor to trigger dissociation of S1 and S2 transition into the postfusion state8,19,33. It is not clear how the resulting S-protein products might modulate a further immune response. In addition, binding of IgG to the MERS S-receptor binding site and concomitant Fc-Fc receptor engagement promotes activation and internalization of the virus causing antibody enhancement of infection34. We find that S-R/PP or related, published PP-containing constructs, exhibit a higher fraction of open states than the closer-to-wild-type S-R. Exclusively closed trimers such as S-R/x2, if used as an antigen in vaccination or immunization, can be expected to induce an immune response that largely excludes receptor binding site targeting antibodies and will thus favor alternate antibodies that do not compete or functionally mimic ACE2 and which may have the potential to lock the S protein in a prefusion state to prevent infection. The stabilization of alternate or transient states to induce alternate immune responses has been a valuable approach in the elicitation of broadly neutralizing antibodies against HIV-1 Env, for example35.
We are optimistic that S-R/x2, or derivatives thereof, will be applicable as diagnostic reagents, as immunogens and as tools to generate or characterize novel monoclonal antibodies for immunotherapy.
Methods
Expression constructs
DNA constructs were designed to transiently express ectodomains of the SARS-CoV-2 S protein for secretion into cell culture media. The DNA sequence for the ectodomain (residues 14–1211) of S from 2019-nCoV BetaCoV/Wuhan/WIV04/2019 (GISAID accession no. EPI_ISL_402124)1, covering the region from the end of the signal peptide sequence (starting with QCVN) to the beginning of the transmembrane domain (ending with QYIK), was synthesized and codon-optimized for HEK293 human cell expression. We added a Kozak sequence as well as sequences encoding an exogenous N-terminal signal peptide MGILPSPGMPALLSLVSLLSVLLMGCVAETGT derived from µ-phosphatase and a C-terminal extension GSGRENLYFQGGGGSGYIPEAPRDGQAYVRKDGEWVLLSTFLGHHHHHH. The C-terminal extension contains a TEV site (italic), a T4 trimerization foldon (underlined) and a Hexa-histag (bold). The extended sequence was cloned into a pCDNA3.1 vector. Modification of the coding sequence of the multibasic S1/S2 cleavage site PRRAR to PGSAS or to a single R was carried out by PCR mutagenesis using Q5 polymerase (New England Biolabs). Introduction of cysteines and modification of residues 986 and 987 were carried out using Q5 polymerase PCR with primers containing desired substitutions, followed by In-Fusion HD (Takara Bio) assembly. Cysteine positions were designed based on visual inspection of published structures, and using Disulfide by Design 2.036.
Protein production and purification
Expi293 cells (Thermo Fisher Scientific) cultured in Expi293 medium (Thermo Fisher Scientific) were maintained at 37 °C. Cells were diluted to a density of 2 × 106 to 3 × 106 cells per ml before transfection. For protein production, 500 µg of DNA was mixed with 1,350 µg of polyethylenimine in 10 ml Expi293 medium, incubated for 10 min, then added to 500 ml of cells. Transfected cells were cultured at 33 °C. Except where otherwise indicated, for non-disulfide-stabilized constructs, transfected cell culture supernatant was collected twice at day 3 and day 6. For disulfide-stabilized mutants, supernatant was collected once at day 6.
To purify S-protein ectodomains, cells were removed from culture medium by centrifugation and 10× phosphate buffered saline (PBS) was added to 1× concentration. Cleared medium was supplemented with 2.5 mM imidazole, 300 mM NaCl and 0.5 mM phenylmethylsulfonyl fluoride and recirculated onto a 5-ml Talon Cobalt column 2.5 times. After sample application, the column was washed with 50 ml of buffer A (25 mM phosphate pH 8.0, 300 mM NaCl, 5 mM imidazole), and protein was eluted with a 100-ml linear gradient to 100% buffer B (25 mM phosphate pH 8.0, 300 mM NaCl, 500 mM imidazole). Fractions containing S protein were pooled, then proteins were concentrated and buffer exchanged into PBS in a 100-kDa MWCO spin concentrator. Purification from 500 ml of cells typically yielded 150–200 µg of disulfide-stabilized protein, ~700 µg of S-R/PP and ~300 µg of S-R. Except where otherwise indicated, proteins were flash-frozen in liquid nitrogen and stored at −80 °C.
Western blot
Cleared cell culture supernatant (30 µl) was heat-denatured ~80-h post-transfection by the addition of 4× NuPAGE LDS sample buffer (Thermo Fisher Scientific) in non-reducing and reducing (in the presence of 250 mM β-mercaptoethanol) conditions at 90 °C for 5 min. Proteins were separated by electrophoresis in NuPAGE 4% to 12%, Bis-Tris gels (Thermo Fisher Scientific). Protein bands were transferred onto Trans-Blot Turbo Midi 0.2-µm PVDF membranes (BioRad) in a Trans-Blot Turbo Transfer System (BioRad). Membranes were blocked with 5% skimmed milk in PBS supplemented with 0.1% Tween 20. The presence of His-tagged protein was detected by chemical luminescence using His-probe (H-3) horseradish peroxidase (HRP) (sc-8036 HRP, Santa Cruz Biotechnology) monoclonal antibody HRP conjugate and Pierce ECL western blotting substrate (Thermo Fisher Scientific).
Negative-staining electron microscopy
Proteins were diluted to ~0.05 mg ml−1 in water. CF200-Cu carbon film grids (EMS) were glow-discharged for 45 s at 25 mA in air, then 3 µl of diluted protein was applied and absorbed for 1 min. The grids were side-blotted and washed once in water and stained twice with Nano-W stain (Nanoprobes) with blotting in between. The grids were air-dried and imaged using a Tecnai T12 microscope operated at 120 kV.
Cryo-electron micoscopy
Grid preparation and image collection were performed similarly for all purified native, mutated and disulfide-stabilized SARS2 S proteins. C-Flat 2/2 3C grids (Protochips) were glow-discharged for 45 s at 25 mA, then 3 µl of freshly purified protein at ~0.2–0.6 mg ml−1 supplemented with 0.01% octyl-glucoside was applied to the grids, which were plunge-frozen in liquid ethane using a Vitrobot system (Thermo Fisher Scientific). Double loading with side blotting between loading was performed when the concentration of purified protein was below 0.3 mg ml−1 (S-R/x2). The grids were stored in liquid nitrogen and loaded into a Titan Krios electron microscope (Thermo Fisher Scientific) operated at 300 kV, equipped with a Gatan K3 BioQuantum direct electron detector with the slit retracted. Movies with 48 frames and an accumulated dose of 50 electrons per Å2 were acquired in counting mode using SerialEM-3.8.037 at the magnification of ×81,000, corresponding to a calibrated pixel size of 1.061 Å per pixel. Detailed data acquisition parameters are summarized in Table 1.
Cryo-electron microscopy image processing
RELION-3.1’s Scheduler (to be described elsewhere) was used for fully automated real-time processing up to initial three-dimensional (3D) classification while micrographs were being collected on the microscope. In a first schedule called preprocess, Movies were motion-corrected and dose-weighted using RELION’s implementation of the MotionCor2 algorithm38; the contrast transfer function (CTF) was estimated by CTFFIND-4.1.1339, and particles were automatically picked by template matching using a structure of the S protein filtered to 20-Å resolution as a 3D reference. In parallel, a second schedule called class3d used the particles generated by the preprocess schedule for 3D classification in batches of 500,000 particles without imposing symmetry to separate SARS-Cov-2 S protein particles from contaminating features.
The remainder of the processing was performed interactively. Particles belonging to 3D classes corresponding to S-protein particles were subjected to one round of 2D classification to remove any remaining contaminating features. Subsequently, a second round of 3D classification was used to assess the ratio of open and closed states and standard 3D auto-refinement was performed on pooled classes corresponding to the different states (Extended Data Fig. 2). Suitable closed and locked classes were subjected to Bayesian polishing to correct for per-particle beam-induced motions, followed by CTF refinement for per-particle defocus, per-micrograph astigmatism, beam-tilt and optical aberration correction40,41 and a final round of 3D auto-refinement.
Map resolutions were estimated at the 0.143 criterion of the phase-randomization-corrected FSC curve calculated between two independently refined half-maps multiplied by a soft-edged solvent mask. Final reconstructions were sharpened and locally filtered in RELION post-processing (Extended Data Fig. 3). The estimated B-factors of each map are listed in Table 1.
Model building and refinement
The SARS-CoV-2 S-protein ectodomain structure (PDB 6VXX17) was used as a starting model for all closed conformations. Models were built and adjusted manually in Coot42. Steric clash and side chain rotamer conformations were improved using the Namdinator web server43. After further manual adjustment, the structures were refined in PHENIX-1.18.26144 to good geometry and the statistics are provided in Table 1. The unmasked model-to-map FSC was calculated in PHENIX for the refined model against the full reconstruction to validate the confidence of the atomic model.
Enzyme-linked immunosorbent assays
The ELISA assay was performed in a two-step process whereby a positivity screen was then followed by an end-point titer determination following previously established protocols45. Briefly, 96-well EIA/RIA plates (Corning, Sigma) were coated with PBS or 0.1 μg per well of either S-R/PP or S-R/x2 at 4 °C overnight. On the following day, the coating solution was removed and wells were blocked with 3% skimmed milk prepared in PBS with 0.1% Tween 20 (PBST) at room temperature for 1 h. Serum samples previously inactivated by heating at 56 °C for 1 h were diluted to 1:60 or serially diluted threefold, six times in 1% skimmed milk in PBST. The blocking solution was removed and the diluted sera were added to the plates and incubated for 2 h at room temperature. Following incubation, the diluted sera were removed and the plates were washed three times with PBST. Goat anti-human IgG secondary antibody-peroxidase (Fc-specific, Sigma) prepared at 1:3,000 in PBST was then added and plates were incubated for 1 h at room temperature. The plates were then washed three times with PBST. The ELISA was developed using 3,5,3′,5′-tetramethylbenzidine (Thermo Fisher Scientific) solution and the reaction was stopped after 10 min incubation using 0.16 M sulfuric acid. The OD450 was measured using a Spectramax i3 plate reader.
The absorbance values for each sample were determined after subtracting the OD values from uncoated wells. All data analyses were performed using Prism 8 version 8.4.2 (GraphPad). To investigate the impact of conformation on reactivity responses, AUC values for all positive samples were determined and Pearson’s correlation analysis comparing the responses to S-R/PP and S-R/x2 was calculated. In addition, the dilution of seropositive samples required to give a 50% maximum signal (EC50) against different S-protein constructs was determined using nonlinear regression analysis in Prism 8 version 8.4.2.
Serum samples for the ELISA assays were obtained from patients attending Addenbrooke’s Hospital with a suspected or confirmed diagnosis of COVID-19 together with health care workers identified through the staff screening as PCR-positive for SARS-CoV-2. Ethical approval was obtained from the East of England Cambridge Central Research Ethics Committee (REC ref. 17/EE/0025). All participants provided informed consent.
Multiplex particle-based flow cytometry (Luminex)
S-GSAS/PP, S-R/PP, S-R, S-R/x2 and RBD were covalently coupled to distinctive carboxylated bead sets (Luminex) to form a five-plex assay. The RBD protein used in this assay includes residues 319–541 with addition of the first 14 residues of S to the N terminus and a 6×His tag to the C terminus46. It was expressed and purified from HEK293 cells.
Beads were first activated with 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide hydrochloride (Thermo Fisher Scientific) in the presence of N-hydroxysuccinimide (Thermo Fisher Scientific), according to the manufacturer’s instructions, to form amine-reactive intermediates. The activated bead sets were incubated with the corresponding proteins at a concentration of 50 μg ml−1 in the reaction mixture for 3 h at room temperature on a rotator. Beads were washed and stored in a blocking buffer (10 mM PBS, 1% BSA, 0.05% NaN3).
The S-variant and RBD coupled bead sets were incubated with SARS-CoV-2 patient sera at three dilutions (1/100; 1/1,000 and 1/10,000) for 1 h in 96-well filter plates (MultiScreenHTS; Millipore) at room temperature in the dark on a horizontal shaker. Fluids were aspirated with a vacuum manifold and beads were washed three times with 10 mM PBS/0.05% Tween 20. Beads were incubated for 30 min with a PE-labeled anti-human IgG-Fc antibody (Leinco/Biotrend), washed as described above, and resuspended in 100 μl PBS/Tween. They were then analyzed on a Luminex analyzer (Luminex/R&D Systems) using Exponent Software V31. Specific binding was reported as MFI. MFI values at each of the three dilutions were summed to generate the points shown in Fig. 4c,d as representative values for comparison purposes.
Patient sera for Luminex assays were collected following informed consent, then aliquoted and held at −70 °C before use. Samples were taken from patients with PCR-confirmed COVID-19 who were being looked after at Royal Papworth Hospital NHS Foundation Trust (RPH) and RPH healthy staff who presented with either no history or a mild history of COVID-19 symptoms. Samples screened positive for SARS-CoV-2 N- and S-binding antibodies by Luminex assay as described were selected for further investigation (ethics approval: IRAS project ID: 96194 REC: 12/WA/0148).
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this Article.
Data availability
EM density maps and atomic models have been deposited in the EMDB and wwPDB with accession codes EMD-11329 and PDB 6ZOX (S-R/x2); EMD-11330 and PDB 6ZOY (S-R/PP/x1, closed); EMD-11331 and PDB 6ZOZ (S-R/PP/x1, locked); EMD-11332 and PDB 6ZP0 (S-R, closed); EMD-11333 and PDB 6ZP1 (S-R/PP, closed); EMD-11334 and PDB 6ZP2 (S-R/PP, locked). Source data are provided with this paper.
References
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
Zhu, N. et al. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 382, 727–733 (2020).
Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 5, 536–544(2020)
Li, F. Structure, function and evolution of coronavirus spike proteins. Annu. Rev. Virol. 3, 237–261 (2016).
Graham, B. S., Gilman, M. S. A. & McLellan, J. S. Structure-based vaccine antigen design. Annu. Rev. Med. 70, 91–104 (2019).
Pallesen, J. et al. Immunogenicity and structures of a rationally designed prefusion MERS-CoV spike antigen. Proc. Natl Acad. Sci. USA 114, E7348–E7357 (2017).
Kirchdoerfer, R. N. et al. Stabilized coronavirus spikes are resistant to conformational changes induced by receptor recognition or proteolysis. Sci. Rep. 8, 15701 (2018).
Walls, A. C. et al. Unexpected receptor functional mimicry elucidates activation of coronavirus fusion. Cell 176, 1026–1039 (2019).
Walls, A. C. et al. Tectonic conformational changes of a coronavirus spike glycoprotein promote membrane fusion. Proc. Natl Acad. Sci. USA 114, 11157–11162 (2017).
Liu, C., Yang, Y., Gao, Y., Shen, C. & Ju, B. Viral architecture of SARS-CoV-2 with post-fusion spike revealed by cryo-EM. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2020.03.02.972927v1 (2020).
Cai, Y. et al. Distinct conformational states of SARS-CoV-2 spike protein. Science https://doi.org/10.1126/science.abd4251 (2020).
Sanders, R. W. et al. A next-generation cleaved, soluble HIV-1 Env trimer, BG505 SOSIP.664 gp140, expresses multiple epitopes for broadly neutralizing but not non-neutralizing antibodies. PLoS Pathog. 9, e1003618 (2013).
McLellan, J. S. et al. Structure-based design of a fusion glycoprotein vaccine for respiratory syncytial virus. Science 342, 592–598 (2013).
Yuan, Y. et al. Cryo-EM structures of MERS-CoV and SARS-CoV spike glycoproteins reveal the dynamic receptor binding domains. Nat. Commun. 8, 15892 (2017).
Gui, M. et al. Cryo-electron microscopy structures of the SARS-CoV spike glycoprotein reveal a prerequisite conformational state for receptor binding. Cell Res. 27, 119–129 (2017).
Wrapp, D. et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367, 1260–1263 (2020).
Walls, A. C. et al. Structure, function and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 181, 281–292 (2020).
Song, W., Gui, M., Wang, X. & Xiang, Y. Cryo-EM structure of the SARS coronavirus spike glycoprotein in complex with its host cell receptor ACE2. PLoS Pathog. 14, e1007236 (2018).
Huo, J. et al. Neutralization of SARS-CoV-2 by destruction of the prefusion spike. Cell Host Microbe https://doi.org/10.1016/j.chom.2020.06.010 (2020).
McCallum, M., Walls, A. C., Corti, D. & Veesler, D. Closing coronavirus spike glycoproteins by structure-guided design. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2020.06.03.129817v1 (2020).
Henderson, R. et al. Controlling the SARS-CoV-2 spike glycoprotein conformation. Nat. Struct. Mol. Biol. https://doi.org/10.1038/s41594-020-0479-4 (2020).
Toelzer, C. et al. Unexpected free fatty acid binding pocket in the cryo-EM structure of SARS-CoV-2 spike protein. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2020.06.18.158584v1 (2020).
Walls, A. C. et al. Cryo-electron microscopy structure of a coronavirus spike glycoprotein trimer. Nature 531, 114–117 (2016).
Walls, A. C. et al. Glycan shield and epitope masking of a coronavirus spike protein observed by cryo-electron microscopy. Nat. Struct. Mol. Biol. 23, 899–905 (2016).
Shang, J. et al. Cryo-EM structure of infectious bronchitis coronavirus spike protein reveals structural and functional evolution of coronavirus spike proteins. PLoS Pathog. 14, e1007009 (2018).
Xiong, X. et al. Glycan shield and fusion activation of a deltacoronavirus spike glycoprotein fine-tuned for enteric infections. J. Virol. 92, e01628 (2018).
Korber, B. et al. Tracking changes in SARS-CoV-2 spike: evidence that D614G increases infectivity of the COVID-19 virus. Cell https://doi.org/10.1016/j.cell.2020.06.043 (2020).
Zhang, L. et al. The D614G mutation in the SARS-CoV-2 spike protein reduces S1 shedding and increases infectivity. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2020.06.12.148726v1 (2020).
Chi, X. et al. A neutralizing human antibody binds to the N-terminal domain of the Spike protein of SARS-CoV-2. Science https://doi.org/10.1126/science.abc6952 (2020).
Ju, B. et al. Human neutralizing antibodies elicited by SARS-CoV-2 infection. Nature https://doi.org/10.1038/s41586-020-2380-z (2020).
Wu, Y. et al. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science 368, 1274–1278 (2020).
Shi, R. et al. A human neutralizing antibody targets the receptor binding site of SARS-CoV-2. Nature https://doi.org/10.1038/s41586-020-2381-y (2020).
Yuan, M. et al. A highly conserved cryptic epitope in the receptor binding domains of SARS-CoV-2 and SARS-CoV. Science 368, 630–633 (2020).
Wan, Y. et al. Molecular mechanism for antibody-dependent enhancement of coronavirus entry. J. Virol. 94, e02015 (2019).
Pancera, M., Changela, A. & Kwong, P. D. How HIV-1 entry mechanism and broadly neutralizing antibodies guide structure-based vaccine design. Curr. Opin. HIV AIDS 12, 229–240 (2017).
Craig, D. B. & Dombkowski, A. A. Disulfide by Design 2.0: a web-based tool for disulfide engineering in proteins. BMC Bioinformatics 14, 346 (2013).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Kidmose, R. T. et al. Namdinator—automatic molecular dynamics flexible fitting of structural models into cryo-EM and crystallography experimental maps. IUCrJ 6, 526–531 (2019).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Amanat, F. et al. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. https://doi.org/10.1038/s41591-020-0913-5 (2020).
Stadlbauer, D. et al. SARS-CoV-2 seroconversion in humans: a detailed protocol for a serological assay, antigen production, and test setup. Curr. Protoc. Microbiol. 57, e100 (2020).
Acknowledgements
We thank the staff of the MRC-LMB for generous support during the COVID-19 pandemic lockdown. We thank the staff of the MRC-LMB EM Facility, in particular G. Sharov and G. Cannone, for supporting EM; J. Grimmett and T. Darling for supporting scientific computing; and P. Edwards for supporting cell culture. We thank T. Nakane for advice on image processing, K. Dent for assistance with sample screening, A. Fountain for assistance with primer design, C. Lu for assistance with bioinformatics, F. Coscia for assistance with EM grid preparation and R. Aricescu for advice on protein expression. We thank D. Mallery and D. Paul for performing exploratory experiments. We thank the Royal Papworth Hospital Clinical Staff and R&D team in supporting patient and staff recruitment, J. Gronlund for support with sample collection and Royal Papworth Hospital healthy staff donors for their participation in this research. This study was supported by funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program (ERC-CoG-648432 MEMBRANEFUSION to J.A.G.B.), the Medical Research Council as part of United Kingdom Research and Innovation (MC_UP_A025_1011 to A.P.C., MC_UP_A025_1013 to S.H.W.S., MC_U105181010 to L.C.J. and MC_UP_1201/16 to J.A.G.B.), 100 Top Talents Program of Sun Yat-sen University (to Y.Z.) and a Wellcome Trust Senior Fellowship (207498/Z/17/Z to I.G.).
Author information
Authors and Affiliations
Consortia
Contributions
X.X. conceived the project. X.X. and J.A.G.B. planned the study. X.X. and Y.Z. designed and synthesized the initial expression construct. X.X., A.P.C. and J.A.G.B. designed the disulfide-stabilized and mutated constructs, which were made by A.P.C. Protein purification was established by X.X. and performed by X.X. and K.A.C. X.X. performed negative staining and prepared EM grids. K.Q. and Z.K. performed cryo-EM, S.H.W.S. assisted with the RELION scheduler and K.Q. processed EM data, which X.X., K.Q. and J.A.G.B. interpreted. Recruitment of donors, management of sera collection and sera processing were performed by L.B., S.B., C.N.C.B.R.C. and H.E.B. ELISA assays were performed by M.H. and I.G. and data analysis by M.H., I.G. and J.A.G.B. Luminex assays were designed by R.D., performed by S.E. and data were analyzed by R.D. and J.A.G.B. G.L.G. and J.A.N. provided critical reagents. L.C.J. contributed to design and interpretation. A.J.C., S.H.W.S., Y.Z., L.J.C., I.G. and J.A.G.B. obtained funding. J.A.G.B. supervised and managed the project. X.X., K.Q., M.H. and J.A.G.B. created the figures. X.X., K.Q. and J.A.G.B. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Peer reviewer reports are available. Inês Chen was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Pipeline used for picking and classification of cryo-EM data for S-R/x2, S-R/PP/x1, S-R, and S-R/PP.
After automated picking, 3D and 2D classification steps were used to remove contaminating objects. 3D classification was then used to sort the data into open, closed and locked conformations.
Extended Data Fig. 2 Resolution assessment of cryo-EM structures.
a) Local resolution maps for all closed structures calculated using RELION. b) Global resolution assessment by Fourier shell correlation at the 0.143 criterion, and correlations of model vs map by Fourier shell correlation at the 0.5 criterion.
Extended Data Fig. 3 Structure of the 833-855 motif.
a) Structural comparison with other closed coronaviruses spikes at the 833-855 motif. SARS-CoV-2 S is compared to spike proteins of murine hepatitis virus (MHV, a betacoronavirus, PDBID: 3JCL), NL63 coronavirus (NL63-CoV, an alphacoronavirus, PDBID: 5SZS), infectious bronchitis virus (IBV, a gammacoronavirus, PDBID: 6CV0) and porcine deltacoronavirus (PDCoV, a deltacoronavirus, PDBID: 6BFU). S proteins are structurally aligned based on S2. S protein trimers from all 4 genera of coronaviruses show presence of folded motifs equivalent to the 833-855 motif identified in SARS-CoV-2 S, suggesting a universal mechanism of locking RBD in closed state in coronavirus spikes. Boxes indicate the positions of the expanded views. b) Interaction between residue D614 and the 833-855 motif by formation of a 2.9 Å salt bridge between D614 and K854. Surrounding structural elements, including fusion peptide and glycans at N616 are shown and indicated. Box indicates the position of the expanded view.
Extended Data Fig. 4 Comparison of immune responses to open and closed conformations of the SARS-CoV-2 spike protein.
a) The dilution of seropositive samples required to give a 50% maximum signal (EC50) against S-R/PP and S-R/x2 was determined using nonlinear regression analysis (n=24). The mean and the 95% confidence intervals are shown. The fold difference in reactivity against S-R/PP and S-R/x2 is shown in panel b.
Supplementary information
Source data
Source Data Fig. 1
Unprocessed western blots and gels.
Rights and permissions
About this article
Cite this article
Xiong, X., Qu, K., Ciazynska, K.A. et al. A thermostable, closed SARS-CoV-2 spike protein trimer. Nat Struct Mol Biol 27, 934–941 (2020). https://doi.org/10.1038/s41594-020-0478-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0478-5
This article is cited by
-
Exploring cell-free assays for COVID-19 serosurvey
Scientific Reports (2024)
-
Immunization with V987H-stabilized Spike glycoprotein protects K18-hACE2 mice and golden Syrian hamsters upon SARS-CoV-2 infection
Nature Communications (2024)
-
High-throughput screening of genetic and cellular drivers of syncytium formation induced by the spike protein of SARS-CoV-2
Nature Biomedical Engineering (2023)
-
Design, structure and plasma binding of ancestral β-CoV scaffold antigens
Nature Communications (2023)
-
Homologous Pairs of Low and High Temperature Originating Proteins Spanning the Known Prokaryotic Universe
Scientific Data (2023)