Abstract
The interplay between E2 and E3 enzymes regulates the polyubiquitination of substrates in eukaryotes. Among the several RING-domain E3 ligases in humans, many utilize two distinct E2s for polyubiquitination. For example, the cell cycle regulatory E3, human anaphase-promoting complex/cyclosome (APC/C), relies on UBE2C to prime substrates with ubiquitin (Ub) and on UBE2S to extend polyubiquitin chains. However, the potential coordination between these steps in ubiquitin chain formation remains undefined. While numerous studies have unveiled how RING E3s stimulate individual E2s for Ub transfer, here we change perspective to describe a case where the chain-elongating E2 UBE2S feeds back and directly stimulates the E3 APC/C to promote substrate priming and subsequent multiubiquitination by UBE2C. Our work reveals an unexpected model for the mechanisms of RING E3–dependent ubiquitination and for the diverse and complex interrelationship between components of the ubiquitination cascade.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data generated and analyzed in this study are available as source data.
Code availability
All codes utilized in this study are available from the authors upon request.
References
Metzger, M. B., Pruneda, J. N., Klevit, R. E. & Weissman, A. M. RING-type E3 ligases: master manipulators of E2 ubiquitin-conjugating enzymes and ubiquitination. Biochim. Biophys. Acta 1843, 47–60 (2014).
Streich, F. C. Jr. & Lima, C. D. Structural and functional insights to ubiquitin-like protein conjugation. Annu. Rev. Biophys. 43, 357–379 (2014).
Rape, M. Ubiquitylation at the crossroads of development and disease. Nat. Rev. Mol. Cell Biol. 19, 59–70 (2018).
Buetow, L. & Huang, D. T. Structural insights into the catalysis and regulation of E3 ubiquitin ligases. Nat. Rev. Mol. Cell Biol. 17, 626–642 (2016).
King, R. W., Deshaies, R. J., Peters, J. M. & Kirschner, M. W. How proteolysis drives the cell cycle. Science 274, 1652–1659 (1996).
Peters, J. M. SCF and APC: the Yin and Yang of cell cycle regulated proteolysis. Curr. Opin. Cell Biol. 10, 759–768 (1998).
Rodrigo-Brenni, M. C. & Morgan, D. O. Sequential E2s drive polyubiquitin chain assembly on APC targets. Cell 130, 127–139 (2007).
Wu, K., Kovacev, J. & Pan, Z. Q. Priming and extending: a UbcH5/Cdc34 E2 handoff mechanism for polyubiquitination on a SCF substrate. Mol. Cell 37, 784–796 (2010).
Haakonsen, D. L. & Rape, M. Branching out: improved signaling by heterotypic ubiquitin chains. Trends Cell Biol. 29, 704–716 (2019).
Wieser, S. & Pines, J. The biochemistry of mitosis. Cold Spring Harb. Perspect. Biol. 7, a015776 (2015).
Kernan, J., Bonacci, T. & Emanuele, M. J. Who guards the guardian? Mechanisms that restrain APC/C during the cell cycle. Biochim. Biophys. Acta Mol. Cell Res. 1865, 1924–1933 (2018).
Huang, J. & Bonni, A. A decade of the anaphase-promoting complex in the nervous system. Genes Dev. 30, 622–638 (2016).
Alfieri, C., Zhang, S. & Barford, D. Visualizing the complex functions and mechanisms of the anaphase promoting complex/cyclosome (APC/C). Open Biol. 7, 170204 (2017).
Watson, E. R., Brown, N. G., Peters, J. M., Stark, H. & Schulman, B. A. Posing the APC/C E3 ubiquitin ligase to orchestrate cell division. Trends Cell Biol. 29, 117–134 (2019).
Visintin, R., Prinz, S. & Amon, A. CDC20 and CDH1: a family of substrate-specific activators of APC-dependent proteolysis. Science 278, 460–463 (1997).
Kim, S. & Yu, H. Mutual regulation between the spindle checkpoint and APC/C. Semin. Cell Dev. Biol. 22, 551–558 (2011).
Musacchio, A. The molecular biology of spindle assembly checkpoint signaling dynamics. Curr. Biol. 25, R1002–R1018 (2015).
Reimann, J. D., Gardner, B. E., Margottin-Goguet, F. & Jackson, P. K. Emi1 regulates the anaphase-promoting complex by a different mechanism than Mad2 proteins. Genes Dev. 15, 3278–3285 (2001).
Chang, L. F., Zhang, Z., Yang, J., McLaughlin, S. H. & Barford, D. Molecular architecture and mechanism of the anaphase-promoting complex. Nature 513, 388–393 (2014).
da Fonseca, P. C. et al. Structures of APC/C(Cdh1) with substrates identify Cdh1 and Apc10 as the D-box co-receptor. Nature 470, 274–278 (2011).
Buschhorn, B. A. et al. Substrate binding on the APC/C occurs between the coactivator Cdh1 and the processivity factor Doc1. Nat. Struct. Mol. Biol. 18, 6–13 (2011).
Kraft, C., Vodermaier, H. C., Maurer-Stroh, S., Eisenhaber, F. & Peters, J. M. The WD40 propeller domain of Cdh1 functions as a destruction box receptor for APC/C substrates. Mol. Cell 18, 543–553 (2005).
Kimata, Y., Baxter, J. E., Fry, A. M. & Yamano, H. A role for the Fizzy/Cdc20 family of proteins in activation of the APC/C distinct from substrate recruitment. Mol. Cell 32, 576–583 (2008).
Chang, L., Zhang, Z., Yang, J., McLaughlin, S. H. & Barford, D. Atomic structure of the APC/C and its mechanism of protein ubiquitination. Nature 522, 450–454 (2015).
Brown, N. G. et al. RING E3 mechanism for ubiquitin ligation to a disordered substrate visualized for human anaphase-promoting complex. Proc. Natl Acad. Sci. USA 112, 5272–5279 (2015).
Li, Q. et al. WD40 domain of Apc1 is critical for the coactivator-induced allosteric transition that stimulates APC/C catalytic activity. Proc. Natl Acad. Sci. USA 113, 10547–10552 (2016).
Brown, N. G. et al. Dual RING E3 architectures regulate multiubiquitination and ubiquitin chain elongation by APC/C. Cell 165, 1440–1453 (2016).
Aristarkhov, A. et al. E2-C, a cyclin-selective ubiquitin carrier protein required for the destruction of mitotic cyclins. Proc. Natl Acad. Sci. USA 93, 4294–4299 (1996).
Yu, H., King, R. W., Peters, J. M. & Kirschner, M. W. Identification of a novel ubiquitin-conjugating enzyme involved in mitotic cyclin degradation. Curr. Biol. 6, 455–466 (1996).
Kirkpatrick, D. S. et al. Quantitative analysis of in vitro ubiquitinated cyclin B1 reveals complex chain topology. Nat. Cell Biol. 8, 700–710 (2006).
Garnett, M. J. et al. UBE2S elongates ubiquitin chains on APC/C substrates to promote mitotic exit. Nat. Cell Biol. 11, 1363–1369 (2009).
Wu, T. et al. UBE2S drives elongation of K11-linked ubiquitin chains by the anaphase-promoting complex. Proc. Natl Acad. Sci. USA 107, 1355–1360 (2010).
Williamson, A. et al. Identification of a physiological E2 module for the human anaphase-promoting complex. Proc. Natl Acad. Sci. USA 106, 18213–18218 (2009).
Lu, Y., Wang, W. & Kirschner, M. W. Specificity of the anaphase-promoting complex: a single-molecule study. Science 348, 1248737 (2015).
Kelly, A., Wickliffe, K. E., Song, L., Fedrigo, I. & Rape, M. Ubiquitin chain elongation requires E3-dependent tracking of the emerging conjugate. Mol. Cell 56, 232–245 (2014).
Wickliffe, K. E., Lorenz, S., Wemmer, D. E., Kuriyan, J. & Rape, M. The mechanism of linkage-specific ubiquitin chain elongation by a single-subunit E2. Cell 144, 769–781 (2011).
Brown, N. G. et al. Mechanism of polyubiquitination by human anaphase-promoting complex: RING repurposing for ubiquitin chain assembly. Mol. Cell 56, 246–260 (2014).
Frye, J. J. et al. Electron microscopy structure of human APC/C(CDH1)-EMI1 reveals multimodal mechanism of E3 ligase shutdown. Nat. Struct. Mol. Biol. 20, 827–835 (2013).
Meyer, H. J. & Rape, M. Enhanced protein degradation by branched ubiquitin chains. Cell 157, 910–921 (2014).
Qiao, R. et al. Mechanism of APC/CCDC20 activation by mitotic phosphorylation. Proc. Natl Acad. Sci. USA 113, E2570–E2578 (2016).
Van Voorhis, V. A. & Morgan, D. O. Activation of the APC/C ubiquitin ligase by enhanced E2 efficiency. Curr. Biol. 24, 1556–1562 (2014).
Sako, K. et al. Emi2 mediates meiotic MII arrest by competitively inhibiting the binding of Ube2S to the APC/C. Nat. Commun. 5, 3667 (2014).
Matyskiela, M. E. & Morgan, D. O. Analysis of activator-binding sites on the APC/C supports a cooperative substrate-binding mechanism. Mol. Cell 34, 68–80 (2009).
Craney, A. et al. Control of APC/C-dependent ubiquitin chain elongation by reversible phosphorylation. Proc. Natl Acad. Sci. USA 113, 1540–1545 (2016).
Yamaguchi, M. et al. Cryo-EM of mitotic checkpoint complex-bound APC/C reveals reciprocal and conformational regulation of ubiquitin ligation. Mol. Cell 63, 593–607 (2016).
Alfieri, C. et al. Molecular basis of APC/C regulation by the spindle assembly checkpoint. Nature 536, 431–436 (2016).
Rodriguez, C. et al. A novel human Cdh1 mutation impairs anaphase promoting complex/cyclosome activity resulting in microcephaly, psychomotor retardation, and epilepsy. J. Neurochem. 151, 103–115 (2019).
Paiva, S. L. & Crews, C. M. Targeted protein degradation: elements of PROTAC design. Curr. Opin. Chem. Biol. 50, 111–119 (2019).
Fujimitsu, K., Grimaldi, M. & Yamano, H. Cyclin-dependent kinase 1-dependent activation of APC/C ubiquitin ligase. Science 352, 1121–1124 (2016).
Zhang, S. et al. Molecular mechanism of APC/C activation by mitotic phosphorylation. Nature 533, 260–264 (2016).
Baek, K. et al. NEDD8 nucleates a multivalent cullin-RING-UBE2D ubiquitin ligation assembly. Nature 578, 461–466 (2020).
Kelsall, I. R. et al. TRIAD1 and HHARI bind to and are activated by distinct neddylated Cullin-RING ligase complexes. EMBO J. 32, 2848–2860 (2013).
Scott, D. C. et al. Two distinct types of E3 ligases work in unison to regulate substrate ubiquitylation. Cell 166, 1198–1214.e24 (2016).
Dueber, E. C. et al. Antagonists induce a conformational change in cIAP1 that promotes autoubiquitination. Science 334, 376–380 (2011).
Feltham, R. et al. Smac mimetics activate the E3 ligase activity of cIAP1 protein by promoting RING domain dimerization. J. Biol. Chem. 286, 17015–17028 (2011).
Stewart, M. D., Ritterhoff, T., Klevit, R. E. & Brzovic, P. S. E2 enzymes: more than just middle men. Cell Res. 26, 423–440 (2016).
Pierce, N. W., Kleiger, G., Shan, S. O. & Deshaies, R. J. Detection of sequential polyubiquitylation on a millisecond timescale. Nature 462, 615–619 (2009).
Kleiger, G., Saha, A., Lewis, S., Kuhlman, B. & Deshaies, R. J. Rapid E2-E3 assembly and disassembly enable processive ubiquitylation of cullin-RING ubiquitin ligase substrates. Cell 139, 957–968 (2009).
Koegl, M. et al. A novel ubiquitination factor, E4, is involved in multiubiquitin chain assembly. Cell 96, 635–644 (1999).
Jarvis, M. A. et al. Measuring APC/C-dependent ubiquitylation in vitro. Methods Mol. Biol. 1342, 287–303 (2016).
Yamaguchi, M. et al. Structure of an APC3–APC16 complex: insights into assembly of the anaphase-promoting complex/cyclosome. J. Mol. Biol. 427, 1748–1764 (2015).
Pickart, C. M. & Raasi, S. Controlled synthesis of polyubiquitin chains. Methods Enzymol. 399, 21–36 (2005).
White, H. D. & Rayment, I. Kinetic characterization of reductively methylated myosin subfragment 1. Biochemistry 32, 9859–9865 (1993).
Bonacci, T. et al. Cezanne/OTUD7B is a cell cycle-regulated deubiquitinase that antagonizes the degradation of APC/C substrates. EMBO J. 37, e98701 (2018).
Williamson, A., Jin, L. & Rape, M. Preparation of synchronized human cell extracts to study ubiquitination and degradation. Methods Mol. Biol. 545, 301–312 (2009).
Moggridge, S., Sorensen, P. H., Morin, G. B. & Hughes, C. S. Extending the compatibility of the SP3 paramagnetic bead processing approach for proteomics. J. Proteome Res. 17, 1730–1740 (2018).
Acknowledgements
We thank M. Brunner, R. VanderLinden and B. Schulman (St. Jude Children’s Research Hospital/HHMI/Max Planck Institute of Biochemistry) for providing reagents, E. Salmon (University of North Carolina) for providing cell lines and T. Kenakin for helpful discussions. Our work is supported by NIH T32GM008570 and NSF DGE-1650116 (T. Bodrug and M.E.G.); NIH T32CA009156 (G.D.G.); NIH P30CA016086 (UNC High-throughput Peptide Synthesis Facility and Array Facility); NIH R01GM083024, NIH R01GM102413 and the W.M. Keck Foundation (J.G.C.); Hertha Firnberg Program of the Austrian Science Fund (R.Q.); Boehringer Ingelheim, the Austrian Research Promotion Agency (Headquarter grant FFG-852936), the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (GA no. 693949) and Human Frontier Science Program grant RGP0057/2018 (J.-M.P.); NIH R01AG011085 (J.W.H.); UNC University Cancer Research Fund (UCRF), NIH R01GM120309 and the American Cancer Society RSG-18-220-01-TBG (M.J.E.); and NIH R35GM128855 and UCRF (N.G.B.).
Author information
Authors and Affiliations
Contributions
R.C.M.-C., T. Bodrug, K.M.K., T. Bonacci, A.O., M.E.G., J.-M.P., J.W.H., M.J.E. and N.G.B. designed the research and were supervised by J.G.C., J.-M.P., J.W.H., M.J.E. and N.G.B. R.C.M.-C., T. Bodrug, D.L.B., K.M.K., T. Bonacci, A.O., M.E.G., F.W., R.Q. and G.D.G. performed research and/or contributed new reagents. R.C.M.-C., T. Bodrug, K.M.K., T. Bonacci, A.O., M.E.G., J.G.C., J.-M.P., J.W.H., M.J.E. and N.G.B. analyzed data. R.C.M.-C., T. Bodrug, M.J.E. and N.G.B. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Katarzyna Marcinkiewicz was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Current model of structural architectures of APC/CCDH1–UBE2C and APC/CCDH1–UBE2S.
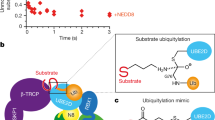
Schematic depicting the interplay of substrate ubiquitination mechanisms, including substrate priming, multiubiquitination, and Ub chain elongation, by APC/CCDH1, UBE2C, and UBE2S. Upon APC/C activation by CDH1 binding, catalytic core APC2WHB–APC11RING is mobile and exposed for UBE2C recruitment (1). APC/CCDH1–UBE2C structural architecture for substrate priming and multiubiquitination (2 and 3). APC/CCDH1–UBE2S structural architecture where APC11RING is repurposed and binds acceptor ubiquitin for K11-linked Ub chain elongation (4).
Extended Data Fig. 2 Substrate polyubiquitination assays by APC/CCDH1, UBE2C, and UBE2S.
UBE2S extends Ub chains on substrates, CycBNTD* and Securin*, and increases the rate of substrate modification in a dose-dependent manner. Fluorescence scan of full SDS-PAGE gels used in (1b). * Represents contaminant present in substrate stock. Uncropped images are available as source data.
Extended Data Fig. 3 UBE2SCTP increases substrate modification by APC/CCDH1–UBE2C.
a, Catalytically inactive UBE2SC95K enhances rapid turnover of CycBNTD* by the APC/CCDH1 and UBE2C. b, Quantification depicts relative fraction of remaining unmodified CycBNTD* shown in Extended Data Figure Fig. 3a. Average of n=3 independent experiments ± s.e.m. c, UBE2S UBC domain (UBE2S core) is defective for Ub chain elongation in an APC/C-dependent manner. d, UBE2SCTP enhances the rapid turnover of CycBNTD* by the APC/CCDH1 and UBE2C. Fluorescence scan of full SDS-PAGE gels used in (2b). Uncropped images for panels (a,c-d) and data for the graph in (b) are available as source data.
Extended Data Fig. 4 UBE2SCTP accelerates substrate modification in every UBE2C-dependent ubiquitination reaction condition using multiple substrates, APC/C variants, and different coactivators.
a, Fluorescence scan of full SDS-PAGE gels used in (2c) showing the effect of the UBE2SCTP on the polyubiquitination of multiple substrates, CycBNTD*, Ub-CycBNTD*, Securin*, Ub-Securin*, by APC/C and UBE2C. b, “No encounter” control assay done as in (2d) with the exception of swapping fluorescent substrate and cold substrate in the mixtures to prevent a reaction to occur. c, Ubiquitination reaction monitoring role of UBE2SCTP on UBE2C-dependent APC/C substrate priming using a single lysine substrate, CycBNTD*(1K), and methylated Ub (meUb). The unmodified substrate and monoubiquitinated product are followed over time in the absence or presence of UBE2SCTP. Quantitation shows effect of UBE2SCTP on substrate priming by APC/CCDH1-UBE2C, graphs depicts fraction of unmodified substrate and meUb~CycBNTD*(1K) over time. Average of n=3 independent experiments ± s.e.m. d, UBE2C-dependent substrate turnover is accelerated by UBE2SCTP when non-phosphorylated, APC/C-pA, or phopho-mimetic APC/C, APC/C-pE, is used. Fluorescence scan of full SDS-PAGE gels used in (2e). e, Substrate turnover by phosphomimetic APC/C, APC/C-pE, and CDC20 is enhanced by the addition of UBE2SCTP in UBE2C-dependent reactions. Fluorescence scan of full SDS-PAGE gels used in (2f). Uncropped images for panels (a-b, d-e) and data for the graph in (c) are available as source data.
Extended Data Fig. 5 UBE2SCTP stimulates the APC/C to activate UBE2C~Ub hydrolysis.
a, UBE2SCTP promotes a substrate-independent APC/C activation of UBE2C, monitored by the hydrolysis of UBE2S~Ub over time into UBE2C and Ub. Detected using Coomassie-stained gels, full gel from (3d). b, Western blot of (3e) showing successful knockdown of UBE2S and CDC20 in HeLa H2B-GFP. c, Time necessary for the metaphase to anaphase transition for different U2OS cells. Data were collected in 2 independent experiments, total number of analyzed cells n = 67 (control), n = 76 (UBE2S knockdown), n = 67 (CDC20 knockdown) and n = 62 (UBE2S and CDC20 knockdown). Median, 25% and 75% percentiles are shown. ** indicates p<0.001, **** indicates p<0.0001 as calculated with Kruskal Wallis test followed by Dunn’s multiple comparison test. Right panel, western blot showing successful knockdown of UBE2S and CDC20 in U2OS H2B-mCherry. d, Western blot in (3g) with the addition of levels of endogenous or recombinant variants of UBE2S. Uncropped images for panels (a,d) and data for the graph in (c) are available as source data.
Extended Data Fig. 6 UBE2SCTP activates an otherwise inactive APC/C variant (APC/CΔAPC1-WD40).
a, As previously reported, APC/CΔAPC1-WD40 is defective for UBE2C-dependent polyubiquitination, but rescued by the addition of the purified APC1-WD40 domain added in trans, monitored by SDS-PAGE and fluorescent scanning12. b, Assays as in (4b), showing the effects of the UBE2SCTP on the polyubiquitination of Securin* by APC/CCDH1 (wild-type and indicated variant), UBE2C, and UBE2S. c, Assays as in (4c), showing the effects of the UBE2SCTP on Ub chain elongation of Ub-Securin by APC/CCDH1 (wild-type and indicated variant) and UBE2S. d, Assays as in (4d-e), showing the effects of the UBE2SCTP on the polyubiquitination of multiple substrate by APC/CCDH1 (wild-type and indicated variant) and UBE2C. e, Representative fluorescent scans of full SDS-PAGE gels in (4f) for data used to determine kinetic parameters upon titrating UBE2C in assays measuring UBE2C-dependent ubiquitination for Ub-CycBNTD* with APC/CCDH1 (wild-type and indicated variant). f, Heat map of Ub-linkages from reactions in (4g) analyzed by label free mass spectrometry. Average of n=3 independent experiments. g, APC/C activation by UBE2SCTP bypasses the requirement for the APC1-WD40 domain. Representative full gel of hydrolysis of UBE2C~Ub monitored by Sypro-stained SDS-PAGE gels (4h). Uncropped image for panel (g) and data for the graph in (f) are available as source Data.
Extended Data Fig. 7 APC/C activation is mediated by UBE2SCTP binding to the APC2–APC4 groove.
a, Representative SDS-PAGE gels in (5b) for data used to determine kinetic parameters in titrating the UBE2SCTP in assays measuring Securin* polyubiquitination by APC/CΔAPC1-WD40, CDH1, and UBE2C. b, As in assays in (5e), UBE2SCTP restores UBE2C-dependent substrate polyubiquitination in APC/C variant defective in CDH1-dependent activation. c, Mutation of the APC2–APC4 groove prevents Ub chain elongation of Ub-CycBNTD* by APC/C (wild-type and indicated variants) and UBE2S. d, Coomassie-stained SDS-PAGE gel of purified wild-type and mutant recombinant APC/Cs. Uncropped image for panel (b) is available as source data.
Supplementary information
Source data
Source Data Fig. 1
Unprocessed gels Fig. 1b.
Source Data Fig. 1
Source data for Fig. 1c.
Source Data Fig. 2
Unprocessed gels Fig. 2b–f.
Source Data Fig. 2
Source data for Fig. 2b and 2d.
Source Data Fig. 3
Unprocessed gels Fig. 3b,d and unprocessed western blots Fig. 3f,g.
Source Data Fig. 3
Source data for Fig. 3c,e.
Source Data Fig. 4
Unprocessed gels Fig. 4b–e,g,h.
Source Data Fig. 4
Source data for Fig. 4f–h.
Source Data Fig. 5
Unprocessed gels Fig. 5d–f.
Source Data Fig. 5
Source data for Fig. 5b,c.
Source Data Extended Data Fig. 2
Unprocessed gels Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Unprocessed gels Extended Data Fig. 3a,c,d.
Source Data Extended Data Fig. 3
Source data for Extended Data Fig. 3b.
Source Data Extended Data Fig. 4
Unprocessed gels Extended Data Fig. 4a,b,d,e.
Source Data Extended Data Fig. 4
Source data for Extended Data Fig. 4c.
Source Data Extended Data Fig. 5
Unprocessed gel Extended Data Fig. 5a and unprocessed western blot Extended Data Fig. 5d.
Source Data Extended Data Fig. 5
Source data for Extended Data Fig. 5c
Source Data Extended Data Fig. 6
Unprocessed gel Extended Data Fig. 6g.
Source Data Extended Data Fig. 6
Source data for Extended Data Fig. 6f
Source Data Extended Data Fig. 7
Unprocessed gel Extended Data Fig. 7b.
Rights and permissions
About this article
Cite this article
Martinez-Chacin, R.C., Bodrug, T., Bolhuis, D.L. et al. Ubiquitin chain-elongating enzyme UBE2S activates the RING E3 ligase APC/C for substrate priming. Nat Struct Mol Biol 27, 550–560 (2020). https://doi.org/10.1038/s41594-020-0424-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0424-6
This article is cited by
-
Time-resolved cryo-EM (TR-EM) analysis of substrate polyubiquitination by the RING E3 anaphase-promoting complex/cyclosome (APC/C)
Nature Structural & Molecular Biology (2023)
-
Structural and functional asymmetry of RING trimerization controls priming and extension events in TRIM5α autoubiquitylation
Nature Communications (2022)
-
CDK4/6 inhibitors downregulate the ubiquitin-conjugating enzymes UBE2C/S/T involved in the ubiquitin–proteasome pathway in ER + breast cancer
Clinical and Translational Oncology (2022)
-
Screening and predicted value of potential biomarkers for breast cancer using bioinformatics analysis
Scientific Reports (2021)
-
UBE2S promotes cell chemoresistance through PTEN-AKT signaling in hepatocellular carcinoma
Cell Death Discovery (2021)