Abstract
Amyotrophic lateral sclerosis (ALS) was initially thought to be associated with oxidative stress when it was first linked to mutant superoxide dismutase 1 (SOD1). The subsequent discovery of ALS-linked genes functioning in RNA processing and proteostasis raised the question of how different biological pathways converge to cause the disease. Both familial and sporadic ALS are characterized by the aggregation of the essential DNA- and RNA-binding protein TDP-43, suggesting a central role in ALS etiology. Here we report that TDP-43 aggregation in neuronal cells of mouse and human origin causes sensitivity to oxidative stress. Aggregated TDP-43 sequesters specific microRNAs (miRNAs) and proteins, leading to increased levels of some proteins while functionally depleting others. Many of those functionally perturbed gene products are nuclear-genome-encoded mitochondrial proteins, and their dysregulation causes a global mitochondrial imbalance that augments oxidative stress. We propose that this stress−aggregation cycle may underlie ALS onset and progression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All deep sequencing data from this study have been deposited in the Gene Expression Omnibus (GEO) under series accession number GSE126801. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE80 partner repository with the dataset identifier PXD021880. Source data are provided with this paper.
References
Morrice, J. R., Gregory-Evans, C. Y. & Shaw, C. A. Necroptosis in amyotrophic lateral sclerosis and other neurological disorders. Biochim. Biophys. Acta Mol. Basis Dis. 1863, 347–353 (2017).
Rosen, D. R. et al. Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 362, 59–62 (1993).
Bruijn, L. I. et al. Aggregation and motor neuron toxicity of an ALS-linked SOD1 mutant independent from wild-type SOD1. Science 281, 1851–1854 (1998).
Barber, S. C. & Shaw, P. J. Oxidative stress in ALS: key role in motor neuron injury and therapeutic target. Free Radic. Biol. Med 48, 629–641 (2010).
Ilieva, H., Polymenidou, M. & Cleveland, D. W. Non-cell autonomous toxicity in neurodegenerative disorders: ALS and beyond. J. Cell Biol. 187, 761–772 (2009).
Da Cruz, S. et al. Elevated PGC-1α activity sustains mitochondrial biogenesis and muscle function without extending survival in a mouse model of inherited ALS. Cell Metab. 15, 778–786 (2012).
Parone, P. A. et al. Enhancing mitochondrial calcium buffering capacity reduces aggregation of misfolded SOD1 and motor neuron cell death without extending survival in mouse models of inherited amyotrophic lateral sclerosis. J. Neurosci. 33, 4657–4671 (2013).
Neumann, M. et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006).
Al-Chalabi, A., van den Berg, L. H. & Veldink, J. Gene discovery in amyotrophic lateral sclerosis: implications for clinical management. Nat. Rev. Neurol. 13, 96–104 (2017).
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Saberi, S. et al. Sense-encoded poly-GR dipeptide repeat proteins correlate to neurodegeneration and uniquely co-localize with TDP-43 in dendrites of repeat-expanded C9orf72 amyotrophic lateral sclerosis. Acta Neuropathol. 135, 459–474 (2018).
Polymenidou, M. & Cleveland, D. W. The seeds of neurodegeneration: prion-like spreading in ALS. Cell 147, 498–508 (2011).
Ling, S. C., Polymenidou, M. & Cleveland, D. W. Converging mechanisms in ALS and FTD: disrupted RNA and protein homeostasis. Neuron 79, 416–438 (2013).
Conlon, E. G. & Manley, J. L. RNA-binding proteins in neurodegeneration: mechanisms in aggregate. Genes Dev. 31, 1509–1528 (2017).
Ito, D., Seki, M., Tsunoda, Y., Uchiyama, H. & Suzuki, N. Nuclear transport impairment of amyotrophic lateral sclerosis-linked mutations in FUS/TLS. Ann. Neurol. 69, 152–162 (2011).
Guo, W. et al. An ALS-associated mutation affecting TDP-43 enhances protein aggregation, fibril formation and neurotoxicity. Nat. Struct. Mol. Biol. 18, 822–830 (2011).
Lagier-Tourenne, C. & Cleveland, D. W. Rethinking ALS: the FUS about TDP-43. Cell 136, 1001–1004 (2009).
Zhang, Y. J. et al. Aberrant cleavage of TDP-43 enhances aggregation and cellular toxicity. Proc. Natl Acad. Sci. USA 106, 7607–7612 (2009).
Li, Q., Yokoshi, M., Okada, H. & Kawahara, Y. The cleavage pattern of TDP-43 determines its rate of clearance and cytotoxicity. Nat. Commun. 6, 6183 (2015).
Dewey, C. M. et al. TDP-43 is directed to stress granules by sorbitol, a novel physiological osmotic and oxidative stressor. Mol. Cell. Biol. 31, 1098–1108 (2011).
Goh, C. W. et al. Chronic oxidative stress promotes GADD34-mediated phosphorylation of the TAR DNA-binding protein TDP-43, a modification linked to neurodegeneration. J. Biol. Chem. 293, 163–176 (2018).
Lei, Y. et al. DJ-1 suppresses cytoplasmic TDP-43 aggregation in oxidative stress-induced cell injury. J. Alzheimers Dis. 66, 1001–1014 (2018).
Wang, W. et al. The inhibition of TDP-43 mitochondrial localization blocks its neuronal toxicity. Nat. Med. 22, 869–878 (2016).
Rot, G. et al. High-resolution RNA maps suggest common principles of splicing and polyadenylation regulation by TDP-43. Cell Rep. 19, 1056–1067 (2017).
Melamed, Z. et al. Premature polyadenylation-mediated loss of stathmin-2 is a hallmark of TDP-43-dependent neurodegeneration. Nat. Neurosci. 22, 180–190 (2019).
Wils, H. et al. TDP-43 transgenic mice develop spastic paralysis and neuronal inclusions characteristic of ALS and frontotemporal lobar degeneration. Proc. Natl Acad. Sci. USA 107, 3858–3863 (2010).
Igaz, L. M. et al. Expression of TDP-43 C-terminal fragments in vitro recapitulates pathological features of TDP-43 proteinopathies. J. Biol. Chem. 284, 8516–8524 (2009).
Krach, F. et al. Transcriptome-pathology correlation identifies interplay between TDP-43 and the expression of its kinase CK1E in sporadic ALS. Acta Neuropathol. 136, 405–423 (2018).
Kapeli, K. et al. Distinct and shared functions of ALS-associated proteins TDP-43, FUS and TAF15 revealed by multisystem analyses. Nat. Commun. 7, 12143 (2016).
Butovsky, O. et al. Targeting miR-155 restores abnormal microglia and attenuates disease in SOD1 mice. Ann. Neurol. 77, 75–99 (2015).
Prudencio, M. et al. Distinct brain transcriptome profiles in C9orf72-associated and sporadic ALS. Nat. Neurosci. 18, 1175–1182 (2015).
King, I. N. et al. The RNA-binding protein TDP-43 selectively disrupts microRNA-1/206 incorporation into the RNA-induced silencing complex. J. Biol. Chem. 289, 14263–14271 (2014).
Fan, Z., Chen, X. & Chen, R. Transcriptome-wide analysis of TDP-43 binding small RNAs identifies miR-NID1 (miR-8485), a novel miRNA that represses NRXN1 expression. Genomics 103, 76–82 (2014).
Tollervey, J. R. et al. Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat. Neurosci. 14, 452–458 (2011).
Xiao, S. et al. RNA targets of TDP-43 identified by UV-CLIP are deregulated in ALS. Mol. Cell Neurosci. 47, 167–180 (2011).
Shu, P. et al. Opposing gradients of microRNA expression temporally pattern layer formation in the developing neocortex. Dev. Cell 49, 764–785.e4 (2019).
Nishimoto, Y. et al. Characterization of alternative isoforms and inclusion body of the TAR DNA-binding protein-43. J. Biol. Chem. 285, 608–619 (2010).
Agarwal, V., Bell, G. W., Nam, J. W. & Bartel, D. P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 4, e05005 (2015).
John, B. et al. Human MicroRNA targets. PLoS Biol. 2, e363 (2004).
Polymenidou, M. et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat. Neurosci. 14, 459–468 (2011).
Iguchi, Y. et al. Oxidative stress induced by glutathione depletion reproduces pathological modifications of TDP-43 linked to TDP-43 proteinopathies. Neurobiol. Dis. 45, 862–870 (2012).
Hu, J. Y. et al. Pathological concentration of zinc dramatically accelerates abnormal aggregation of full-length human Tau and thereby significantly increases Tau toxicity in neuronal cells. Biochim. Biophys. Acta Mol. Basis Dis. 1863, 414–427 (2017).
Chou, C. C. et al. TDP-43 pathology disrupts nuclear pore complexes and nucleocytoplasmic transport in ALS/FTD. Nat. Neurosci. 21, 228–239 (2018).
von Mering, C. et al. STRING: known and predicted protein–protein associations, integrated and transferred across organisms. Nucleic Acids Res. 33, D433–D437 (2005).
Higuchi-Sanabria, R., Frankino, P. A., Paul, J. W. III., Tronnes, S. U. & Dillin, A. A futile battle? Protein quality control and the stress of aging. Dev. Cell 44, 139–163 (2018).
Chiang, P. M. et al. Deletion of TDP-43 down-regulates Tbc1d1, a gene linked to obesity, and alters body fat metabolism. Proc. Natl Acad. Sci. USA 107, 16320–16324 (2010).
Kawahara, Y. & Mieda-Sato, A. TDP-43 promotes microRNA biogenesis as a component of the Drosha and Dicer complexes. Proc. Natl Acad. Sci. USA 109, 3347–3352 (2012).
Srivastava, T. et al. Regulation of neuronal mRNA translation by CaM-kinase I phosphorylation of eIF4GII. J. Neurosci. 32, 5620–5630 (2012).
Hazelett, D. J., Chang, J. C., Lakeland, D. L. & Morton, D. B. Comparison of parallel high-throughput RNA sequencing between knockout of TDP-43 and its overexpression reveals primarily nonreciprocal and nonoverlapping gene expression changes in the central nervous system of Drosophila. G3 (Bethesda) 2, 789–802 (2012).
Arai, T. et al. Phosphorylated TDP-43 in Alzheimer’s disease and dementia with Lewy bodies. Acta Neuropathol. 117, 125–136 (2009).
Hasegawa, M. et al. TDP-43 is deposited in the Guam parkinsonism–dementia complex brains. Brain 130, 1386–1394 (2007).
Schwab, C., Arai, T., Hasegawa, M., Yu, S. & McGeer, P. L. Colocalization of transactivation-responsive DNA-binding protein 43 and huntingtin in inclusions of Huntington disease. J. Neuropathol. Exp. Neurol. 67, 1159–1165 (2008).
Sena, L. A. & Chandel, N. S. Physiological roles of mitochondrial reactive oxygen species. Mol. Cell 48, 158–167 (2012).
Wong, A., Boutis, P. & Hekimi, S. Mutations in the clk-1 gene of Caenorhabditis elegans affect developmental and behavioral timing. Genetics 139, 1247–1259 (1995).
Gautam, M. et al. Mitochondria, ER, and nuclear membrane defects reveal early mechanisms for upper motor neuron vulnerability with respect to TDP-43 pathology. Acta Neuropathol. 137, 47–69 (2019).
Riedl, S. J. & Shi, Y. Molecular mechanisms of caspase regulation during apoptosis. Nat. Rev. Mol. Cell Biol. 5, 897–907 (2004).
Cui, H., Kong, Y. & Zhang, H. Oxidative stress, mitochondrial dysfunction, and aging. J. Signal Transduct. 2012, 646354 (2012).
Houtkooper, R. H. et al. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature 497, 451–457 (2013).
Masand, R. et al. Proteome imbalance of mitochondrial electron transport chain in brown adipocytes leads to metabolic benefits. Cell Metab. 27, 616–629.e4 (2018).
Rudnick, N. D. et al. Distinct roles for motor neuron autophagy early and late in the SOD1(G93A) mouse model of ALS. Proc. Natl Acad. Sci. USA 114, E8294–E8303 (2017).
Yang, Q. et al. Highly sensitive sequencing reveals dynamic modifications and activities of small RNAs in mouse oocytes and early embryos. Sci. Adv. 2, e1501482 (2016).
de Planell-Saguer, M., Rodicio, M. C. & Mourelatos, Z. Rapid in situ codetection of noncoding RNAs and proteins in cells and formalin-fixed paraffin-embedded tissue sections without protease treatment. Nat. Protoc. 5, 1061–1073 (2010).
Kim, K. H., Son, J. M., Benayoun, B. A. & Lee, C. The mitochondrial-encoded peptide MOTS-c translocates to the nucleus to regulate nuclear gene expression in response to metabolic stress. Cell Metab. 28, 516–524.e7 (2018).
Wang, G., Gong, Y., Burczynski, F. J. & Hasinoff, B. B. Cell lysis with dimethyl sulphoxide produces stable homogeneous solutions in the dichlorofluorescein oxidative stress assay. Free Radic. Res 42, 435–441 (2008).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 6, 11 (2015).
Lowe, T. M. & Chan, P. P. tRNAscan-SE On-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res. 44, W54–W57 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Lovci, M. T. et al. Rbfox proteins regulate alternative mRNA splicing through evolutionarily conserved RNA bridges. Nat. Struct. Mol. Biol. 20, 1434–1442 (2013).
Calvo, S. E., Clauser, K. R. & Mootha, V. K. MitoCarta2.0: an updated inventory of mammalian mitochondrial proteins. Nucleic Acids Res. 44, D1251–D1257 (2016).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Mi, H. et al. PANTHER version 11: expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. 45, D183–D189 (2017).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (91640115, 31670827, and 31871316 to Y.Z.; 31770833 and 31570779 to Y. Liang), and the Ministry of Science and Technology of China (2017YFA0504400 to Y.Z.; 2017YFA0504600 to X. Zhang). We thank W. Xu and Y. Liang (Wuhan University) for providing the Sarkosyl reagent and the related protocol. Part of the computation in this work was done on the supercomputing system in the Supercomputing Center of Wuhan University.
Author information
Authors and Affiliations
Contributions
X. Zuo, X.-D.F. and Y.Z. designed the experiments. X. Zuo and Y. Li carried out the majority of the experiments. J.Z., Z.C. and Y.Z. analyzed the sequencing data. K.W. performed mass spectrometric analysis. X. Zhang and Y. Liang contributed to critical experimental information. M.A.E. and Z.L. provided sodium arsenite and human iPSCs with related protocols. X. Zuo, Y.Z. and X.-D.F. wrote the paper. All authors discussed the results and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Structural & Molecular Biology thanks Defne Amado and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Anke Sparmann was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Transcriptomic comparison between TDP-43 knockdown and expression of the aggregation-prone CTF35.
a, Confirmation of the knockdown effect of multiple siRNAs against TDP-43 in N2a cells by Western blotting analysis. GAPDH served as loading control. The targeting regions in mouse TDP-43 are indicated with individual siRNAs. Note that siTDP-43-2290 corresponds to the siRNA against TDP-43-3′UTR. For RNA-seq analysis, total RNAs were extracted from N2a cells transfected with siTDP-43-1078 or control siRNA. b, Expression of wt TDP-43 or CTF35 in N2a cells depleted of endogenous TDP-43, analyzed by Western blotting. c, Reproducibility of duplicated RNA-seq libraries from siNC-treated (left) or siTDP-43 treated (right) N2a cells. d, Reproducibility of duplicated RNA-seq libraries from TDP-43 depleted N2a cells complemented with either wt full-length TDP-43 (left) or with CTF35 (right). e, Percentages of commonly affected genes in siTDP-43 treated N2a cells relative to three different sources of ALS patients. For a and b, a representative example of three independent experiments is shown. Uncropped images for panels a and b are available as source data online.
Extended Data Fig. 2 Characterization of TDP-43 and miRNA interactions.
a, Western blotting analysis of TDP-43 immunoprecipitation in N2a cells by probing TDP-43, Ago2 and GAPDH. b, Reproducibility of duplicated small RNA-seq libraries generated with total input (left) or TDP-43 IPed (right) miRNAs. c, Profile of total miRNAs versus those enriched with TDP-43 IP. Heat map show the expression miRNAs levels (left) and fold-enrichment compared to Input (right). d, Mass spectrometric analysis of proteins pulled down with biotin-labeled let-7c, biotin-labeled control RNA and naked beads without RNA bait. The data are presented in a triangular plot with yellow dots highlighting let-7c enriched proteins that include TDP-43 (top), which was further validated by Western blotting (bottom). e, RT-qPCR analysis of miRNAs enriched by myc-tag IP from N2a cells expressing myc-tagged full-length TDP-43 versus RRM1 and 2-deleted mutant. Red and Gray: target and non-target miRNAs analyzed. f, Biotin-let-7c capture of individually expressed wt and RRM1 or RRM2 deleted TDP-43 in N2a cells. Data in e (lower panel) are shown as mean ± s.d. (n = 3 independent experiments). For western blot in a and d-f, a representative example of three independent experiments is shown. Data for graph in e are available as source data online. Uncropped images for panels a and d-f are available as source data online.
Extended Data Fig. 3 TDP-43 modulation of miRNA target gene expression.
a, Western blotting analysis of IPed Ago2 under mock-treated (left) and TDP-43 overexpression (right) conditions. GAPDH served as loading control. b, RT-qPCR analysis of the expression of targets for TDP-43 bound miRNAs (red) and for non-bound miRNA (gray) in response to TDP-43 depletion. c, Dual luciferase reporter assay on individual target reporters. In each, the MRE sequence is shown with the seed region base-paired with the miRNA. Three copies of each MRE was inserted in each reporter. Corresponding to each wt reporter is the mutant one in which the seed region was mutated. Each pair of reporters was assayed in response to specific miRNA mimics or inhibitors. d, and e, Cumulative distribution plots of siTDP-43 induced expression changes among up-regulated genes with or without predicted target sites for TDP-43 sequestrated miRNAs (d), and of siTDP-43 induced expression changes among down-regulated genes with or without predicted target sites for TDP-43 non-bound miRNAs (e). Different colored lines indicate non-targets (gray), all predicted targets (blue), genes that contain more than one miRNA target (orange), and genes that contain more than two predicted miRNA target sites (red). The significance for each shift from non-target genes is indicated by P value determined by Wilcoxon signed-rank test. f, Boxplots of TDP-43 depletion-induced expression changes for genes with (n = 9,217) or without (n = 2,140) TDP-43 binding peaks. g, Boxplots of expression for genes with (n = 3,974) or without (n = 2,140) TDP-43 binding peaks in 3′UTRs. The lower and upper hinges of the boxplots correspond to the first and third quartiles (the 25th and 75th percentiles). The upper whisker extends from the hinge to the largest value no further than 1.5 * IQR from the hinge (where IQR is the interquartile range, or distance between the first and third quartiles). The lower whisker extends from the hinge to the smallest value at most 1.5 * IQR of the hinge. Data beyond the end of the whiskers are outlying points that are plotted individually. Data in b and c are presented as mean ± s.d. (n = 3 independent experiments). For b, c, f, and g, the P values are based on a two-tailed unpaired t test: *p < 0.05, **p < 0.01, ***p < 0.001; N.S., non-significant. For a, a representative example of three independent experiments is shown. Data for graphs in b and c are available as source data online. Uncropped images for panels a are available as source data online.
Extended Data Fig. 4 Characterization of isolated and induced neurons.
a, Western blotting analysis of isolated primary mouse neurons and N2a cells by probing both pan-neuron (Tuj1) and mature neuron (SYN1 and MAP2) markers. The non-neuronal MEF cells served as control. b, Immunostaining with Tuj1 (red) and SYN1 (green) on isolated primary mouse neurons. Scale bar: 100 μm. c,d,e, Characterization of iPSCs with the stem cell markers Oct-4A and SOX2 (c), induced NPCs with Nestin, Vimentin, SOX2, and Pax6 (d), and differentiated neurons with Tuj1, SYN1, and MAP2 (e). Scale bar: 100 μm. For a-e, a representative example of three independent experiments is shown. Uncropped images for panel a are available as source data online.
Extended Data Fig. 5 Partial purification of TDP-43 aggregates for global analysis.
a, The fractionation scheme for purifying TDP-43 aggregates. b, and c, Western blotting analysis of TDP-43 in N2a cells in response to sodium arsenite (b) and heat shock (c). d, and e, Western blotting analysis of TDP-43 in soluble and pellet fractions of two indicated neuronal (d) and two non-neuronal (e) cells. (f) Reproducibility of duplicated small RNA-seq on total (left) versus pellet fraction (right) miRNAs. g, RT-qPCR analysis of TDP-43 target and non-target miRNAs in the pellet. Data in g are shown as mean ± s.d. (n = 3 independent experiments). For b-e, a representative example of three independent experiments is shown. Data for graph in g are available as source data online. Uncropped images for panels b-e are available as source data online.
Extended Data Fig. 6 Analysis of the proteome in H2O2-induced TDP-43 pellet.
a, SDS-PAGE analysis of total versus pellet from mock-treated and H2O2-treated N2a cells. Proteins were stained with Coomassie brilliant blue. b, Western blotting validation of specific nDNA-encoded mitochondrial proteins in H2O2-induced pellet in N2a cells. c, and d, Immunofluorescence of N2a cells treated with DMSO or H2O2 and stained with antibodies against TDP-43 C-terminal, representative RBPs hnRNP A0 and hnRNP M (c), and against TDP-43 C-terminal, representative mitochondrial proteins LRPPRC and ATP5b (d). Arrows indicate colocalization of TDP-43 with indicated proteins in granules. Scale bar: 10 μm. e, The STRING DATABASE was used to connect the proteins identified in H2O2-induced pellet using a cutoff score > 0.15. The resulting protein interaction network contains 69 nodes and 456 non-redundant edges. Of these, 5 proteins do not interact with any other proteins, and 3 other proteins cannot be recognized by STRING. For panel a, a representative example of five independent experiments is shown. For b-d, a representative example of three independent experiments is shown. Uncropped images for panel b are available as source data online.
Extended Data Fig. 7 Mimicking mitochondrial imbalance by RNAi and overexpression.
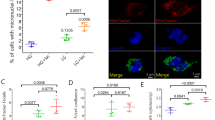
a, Overexpression of myc-tagged CTF35 in N2a cells, validated with the anti-myc antibody. GAPDH served as loading control. b, Western blotting validation of the siRNA effects on indicated nDNA-encoded mitochondrial proteins in N2a cells. c, Western blotting validation of the efficacy and specificity in double siRNA-treated N2a cells. d, and e, Western blotting of single and double expression of indicated nDNA-encoded mitochondrial proteins. f, Western blotting analysis of the effect of combinatory siRNA treatment (LRPPRC) and overexpression (Bcl2l2) of separate nDNA-encoded mitochondrial proteins in N2a cells. g, Quantification of ROS production upon knockdown of two representative “up-regulated” genes (Fam173b and Timm9; ATP5b as a positive control) or overexpression of two representative “down-regulated” genes (Ndufa9 and ATP5C1; Bcl2l2 as a positive control). Data in g were presented as mean ± s.d. (n = 3 independent experiments). The P values are based on a two-tailed unpaired t test: **p < 0.01, ***p < 0.001. For a-f and g (lower panel), a representative example of three independent experiments is shown. Data for graphs in g are available as source data online. Uncropped images for panels a-f and g are available as source data online.
Supplementary information
Supplementary Table 1
Differentially expressed genes in TDP-43-knockdown and CTF35-overexpression N2a cells.
Supplementary Table 2
TDP-43 RIP-enriched miRNAs and a subset of enriched miRNAs contributing to the shift of downregulated targets.
Supplementary Table 3
Enriched miRNAs in H2O2-induced TDP-43 pellet.
Supplementary Table 4
Enriched proteins in H2O2-induced TDP-43 pellet.
Supplementary Table 5
Mitochondrial proteins co-aggregated with TDP-43 and upregulated mitochondrial genes induced by CTF35 overexpression.
Supplementary Table 6
Primers for vector construction and small RNA oligos.
Supplementary Table 7
Primers for quantifying miRNA expression.
Supplementary Table 8
Oligos used for small RNA library preparation.
Supplementary Table 9
Modified RNA probes.
Source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 2
Unprocessed western blots
Source Data Fig. 3
Statistical source data
Source Data Fig. 3
Unprocessed western blots
Source Data Fig. 4
Unprocessed western blots
Source Data Fig. 5
Statistical source data
Source Data Fig. 5
Unprocessed western blots
Source Data Fig. 6
Unprocessed western blots
Source Data Fig. 7
Statistical source data
Source Data Fig. 7
Unprocessed western blots
Source Data Extended Data Fig. 1
Unprocessed western blots
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 2
Unprocessed western blots
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 3
Unprocessed western blots
Source Data Extended Data Fig. 4
Unprocessed western blots
Source Data Extended Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 5
Unprocessed western blots
Source Data Extended Data Fig. 6
Unprocessed western blots
Source Data Extended Data Fig. 7
Statistical source data
Source Data Extended Data Fig. 7
Unprocessed western blots
Rights and permissions
About this article
Cite this article
Zuo, X., Zhou, J., Li, Y. et al. TDP-43 aggregation induced by oxidative stress causes global mitochondrial imbalance in ALS. Nat Struct Mol Biol 28, 132–142 (2021). https://doi.org/10.1038/s41594-020-00537-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-00537-7
This article is cited by
-
Guggulsterone Selectively Modulates STAT-3, mTOR, and PPAR-Gamma Signaling in a Methylmercury-Exposed Experimental Neurotoxicity: Evidence from CSF, Blood Plasma, and Brain Samples
Molecular Neurobiology (2024)
-
Ergothioneine promotes longevity and healthy aging in male mice
GeroScience (2024)
-
Ferroptosis regulation through Nrf2 and implications for neurodegenerative diseases
Archives of Toxicology (2024)
-
Mitochondrial dysfunction: roles in skeletal muscle atrophy
Journal of Translational Medicine (2023)
-
Implications of TDP-43 in non-neuronal systems
Cell Communication and Signaling (2023)