Abstract
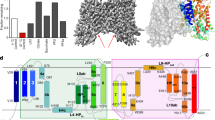
ATP-citrate lyase (ACLY) synthesizes cytosolic acetyl coenzyme A (acetyl-CoA), a fundamental cellular building block. Accordingly, aberrant ACLY activity is observed in many diseases. Here we report cryo-EM structures of human ACLY, alone or bound to substrates or products. ACLY forms a homotetramer with a rigid citrate synthase homology (CSH) module, flanked by four flexible acetyl-CoA synthetase homology (ASH) domains; CoA is bound at the CSH–ASH interface in mutually exclusive productive or unproductive conformations. The structure of a catalytic mutant of ACLY in the presence of ATP, citrate and CoA substrates reveals a phospho-citryl-CoA intermediate in the ASH domain. ACLY with acetyl-CoA and oxaloacetate products shows the products bound in the ASH domain, with an additional oxaloacetate in the CSH domain, which could function in ACLY autoinhibition. These structures, which are supported by biochemical and biophysical data, challenge previous proposals of the ACLY catalytic mechanism and suggest additional therapeutic possibilities for ACLY-associated metabolic disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Structures and EM maps of ACLY-apo (PDB-6POF, EMDB-20414), ACLY–citrate-CoA-D2 (PDB-6UUZ, EMDB-20903), ACLY–citrate-CoA-C1 assym open (PDB-6UIA, EMDB-20784), ACLY–citrate-CoA-C1 assym closed (PDB-6POE, EMDB-20413), ACLY–OAA–acetyl-CoA-C1 (PDB-6UV5, EMDB-20904), ACLY–OAA–acetyl-CoA-D2 (PDB-6UI9, EMDB-20783) and ACLY-E599Q–ATP-citrate-CoA (PDB-6UUW, EMDB-20902) have been deposited to the PDB and EMDB databases. Source data for Figs. 1f, 4d,e and 5a,b and Extended Data Fig. 1c are available with the online version of the paper.
Change history
14 February 2020
The callout to Extended Data Fig. 7c was linked to Fig. 4; the link has now been removed.
02 April 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41594-020-0421-9
References
Lee, J. V., Shah, S. A. & Wellen, K. E. Obesity, cancer and acetyl-CoA metabolism. Drug Discov. Today Dis. Mech. 10, e55–e61 (2013).
Chypre, M., Zaidi, N. & Smans, K. ATP-citrate lyase: a mini-review. Biochem. Biophys. Res. Commun. 422, 1–4 (2012).
Granchi, C. ATP citrate lyase (ACLY) inhibitors: an anti-cancer strategy at the crossroads of glucose and lipid metabolism. Eur. J. Med. Chem. 157, 1276–1291 (2018).
Bilen, O. & Ballantyne, C. M. Bempedoic acid (ETC-1002): an investigational inhibitor of ATP citrate lyase. Curr. Atheroscler. Rep. 18, 61 (2016).
Burke, A. C. & Huff, M. W. ATP-citrate lyase: genetics, molecular biology and therapeutic target for dyslipidemia. Curr. Opin. Lipidol. 28, 193–200 (2017).
Pinkosky, S. L. et al. Liver-specific ATP-citrate lyase inhibition by bempedoic acid decreases LDL-C and attenuates atherosclerosis. Nat. Commun. 7, 13457 (2016).
Ray, K. K. et al. Safety and efficacy of bempedoic acid to reduce LDL cholesterol. N. Engl. J. Med. 380, 1022–1032 (2019).
Sanchez, L. B., Galperin, M. Y. & Muller, M. Acetyl-CoA synthetase from the amitochondriate eukaryote Giardia lamblia belongs to the newly recognized superfamily of acyl-CoA synthetases (nucleoside diphosphate-forming). J. Biol. Chem. 275, 5794–5803 (2000).
Bond, D. R. et al. Characterization of citrate synthase from Geobacter sulfurreducens and evidence for a family of citrate synthases similar to those of eukaryotes throughout the Geobacteraceae. Appl. Environ. Microbiol. 71, 3858–3865 (2005).
Nguyen, N. T. et al. Comparative analysis of folding and substrate binding sites between regulated hexameric type II citrate synthases and unregulated dimeric type I enzymes. Biochemistry 40, 13177–13187 (2001).
Singh, M., Richards, E. G., Mukherjee, A. & Srere, P. A. Structure of ATP citrate lyase from rat liver. Physicochemical studies and proteolytic modification. J. Biol. Chem. 251, 5242–5250 (1976).
Walsh, C. T. Jr. & Spector, L. B. Citryl phosphate and the mode of action of the citrate cleavage enzyme. J. Biol. Chem. 244, 4366–4374 (1969).
Sun, T., Hayakawa, K., Bateman, K. S. & Fraser, M. E. Identification of the citrate-binding site of human ATP-citrate lyase using X-ray crystallography. J. Biol. Chem. 285, 27418–27428 (2010).
Inoue, H., Fujio, S., Keihachi, F., Kozaburo, A. & Takeda, Y. Studies on ATP citrate lyase of rat liver: I. Purification and some properties. J. Biol. Chem. 60, 543–553 (1966).
Inoue, H., Suzuki, F., Tanioka, H. & Takeda, Y. Role of ATP in the ATP citrate lyase reaction. Biochem. Biophys. Res. Commun. 26, 602–608 (1967).
Inoue, H., Suzuki, F., Tanioka, H. & Takeda, Y. Studies on ATP citrate lyase of rat liver. 3. The reaction mechanism. J. Biochem. 63, 89–100 (1968).
Inoue, H., Tsunemi, T., Suzuki, F. & Takeda, Y. Studies on ATP citrate lyase of rat liver. IV. The role of CoA. J. Biochem. 65, 889–900 (1969).
Srere, P. A. The citrate cleavage enzyme. I. Distribution and purification. J. Biol. Chem. 234, 2544–2547 (1959).
Srere, P. A. The citrate cleavage enzyme: II. Stoichiometry substrate specificity and its use for coenzyme A assay. J. Biol. Chem. 236, 50–53 (1961).
Srere, P. A. & Lipmann, F. An enzymatic reaction between citrate, adenosine triphosphate and coenzyme A1. J. Am. Chem. Soc. 75, 4874–4874 (1953).
Bazilevsky, G. A. et al. ATP-citrate lyase multimerization is required for coenzyme-A substrate binding and catalysis. J. Biol. Chem. 294, 7259–7268 (2019).
Verschueren, K. H. G. et al. Structure of ATP citrate lyase and the origin of citrate synthase in the Krebs cycle. Nature 568, 571–575 (2019).
Wei, J. et al. An allosteric mechanism for potent inhibition of human ATP-citrate lyase. Nature 568, 566–570 (2019).
Hu, J., Komakula, A. & Fraser, M. E. Binding of hydroxycitrate to human ATP-citrate lyase. Acta Crystallogr. D Struct. Biol. 73, 660–671 (2017).
Wiegand, G., Remington, S., Deisenhofer, J. & Huber, R. Crystal structure analysis and molecular model of a complex of citrate synthase with oxaloacetate and S-acetonyl-coenzyme A. J. Mol. Biol. 174, 205–219 (1984).
Pentyala, S. N. & Benjamin, W. B. Effect of oxaloacetate and phosphorylation on ATP-citrate lyase activity. Biochemistry 34, 10961–10969 (1995).
Rokita, S. E., Srere, P. A. & Walsh, C. T. 3-Fluoro-3-deoxycitrate: a probe for mechanistic study of citrate-utilizing enzymes. Biochemistry 21, 3765–3774 (1982).
Fan, F. et al. On the catalytic mechanism of human ATP citrate lyase. Biochemistry 51, 5198–5211 (2012).
Tanner, K. G. et al. Catalytic mechanism and function of invariant glutamic acid 173 from the histone acetyltransferase GCN5 transcriptional coactivator. J. Biol. Chem. 274, 18157–18160 (1999).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Kim, D. E., Chivian, D. & Baker, D. Protein structure prediction and analysis using the Robetta server. Nucleic Acids Res. 32, W526–W531 (2004).
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 40 (2008).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Delano, W. L. The PyMol Molecular Graphics System (DeLano Scientific, 2002).
Linn, T. C. & Srere, P. A. Identification of ATP citrate lyase as a phosphoprotein. J. Biol. Chem. 254, 1691–1698 (1979).
Michnik, A., Drzazga, Z., Kluczewska, A. & Michalik, K. Differential scanning microcalorimetry study of the thermal denaturation of haemoglobin. Biophys. Chem. 118, 93–101 (2005).
Acknowledgements
Negative staining and cryo-EM grids screening were carried out in EMRL at the Perelman School of Medicine, University of Pennsylvania. We thank B. Zuo and S. Molugu for their help with negative staining and cryo-EM grids screening. Cryo-EM data collection was carried out at the University of Massachusetts Cryo-EM Core Facility. Molecular graphics and structural analyses were performed with UCSF Chimera, developed by the Resource for Biocomputing, Visualization and Informatics at the University of California, San Francisco, with support from NIH P41-GM103311. We thank K. Song, K. Lee, C. Xu and other staff members at the University of Massachusetts Cryo-EM Core Facility for their support with cryo-EM data collection; K. Wellen, A. Olia, S. Deng and M. Ricketts for helpful discussions; T. Eeuwen and K. Murakami for advice with EM data collection; and S. Zeng for organizing cryo-EM data collection trips. This work was supported by NIH grants R35 GM118090 and P01 AG031862 to R.M. and NIH grant no. F31CA189559 to G.B.
Author information
Authors and Affiliations
Contributions
Conceptualization was provided by X.W., K.S., G.A.B., A.V. and R.M.; methodology by X.W., K.S., G.A.B., A.V. and R.M.; investigation by X.W., K.S., G.A.B. and A.V.; and formal analysis by X.W., K.S., G.A.B. and A.V. The original draft was written by X.W. Visualization was provided by X.W. and G.A.B. The manuscript was reviewed and edited by X.W., K.S., G.A.B., A.V. and R.M. Funding acquisition was carried out by R.M. Resources were provided by R.M. Supervision was performed by R.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Anke Sparmann was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Production of recombinant human ACLY.
a, SDS-PAGE (12% acrylamide) verification of intact, homogeneous human ACLY protein produced in Escherichia coli. b, Size-exclusion chromatography of ACLY on a superdex 200 increase column. c, Differential Scanning Fluorimetry (DSF) of ACLY in the absence and presence of metabolite ligands (Mean + /- Standard Deviation, n = 3 biologically independent samples). P-values were calculated by two-way ANOVA with Dunnett’s multiple comparisons test.
Extended Data Fig. 2 Electron micrographs of ACLY.
a, Representative negative stain images of human ACLY. b, 2D class average from 3,000 negative stain ACLY. c, Representative cryo-EM images of human ACLY in complex with citrate-CoA. d, Representative 2D class averages of cryo-EM ACLY particles.
Extended Data Fig. 3
Image processing workflow for single particle reconstruction of ACLY–citrate-CoA.
Extended Data Fig. 4 Analysis of single particle cryo-EM reconstructions.
a, Fourier Shell Correlation (FSC) curves for 3D reconstructions of reported structures, marked with resolutions corresponding to FSC = 0.143. b, Cryo-EM density of representative helical segments (residues 1055-1077) from ACLY–citrate-CoA (left) and ACLY–OAA–acetyl-CoA structures. c, Local resolution estimation of cryo-EM maps of ACLY–citrate-CoA-D2 (left), ACLY–citrate-CoA-C1 asymm closed (middle) and ACLY–OAA–acetyl-CoA (right) by Resmap.
Extended Data Fig. 5 Structural comparison of the CSH module of ACLY with citrate synthase.
a, Side-by-side views of Sulfolobus solfataricus citrate synthase (left) with the ACLY CSH module. b, Superposition of CSH/OAA2 with pig heart citrate synthase bound to OAA.
Extended Data Fig. 6 Model of alternate acetyl-CoA orientations against the citrate synthase homology (CSH) domain.
a, Both the extended and ‘flipped’ orientations of acetyl-CoA with the observed cryo-EM density (blue chicken wire) are shown. b, The flipped orientation of acetyl-CoA is modeled into ACLY-apo showing no significant steric clashes.
Extended Data Fig. 7 Cryo-EM density of ACLY ligands.
a, CoA in ACLY–CoA–citrate. b, Acetyl-CoA in ACLY–OAA–acetyl-CoA. c, Phosphor-citryl-CoA in ACLY-E599Q–ATP-citrate-CoA. d, Superposition of Acetyl-CoA and Phosphor-citryl-CoA showing movement of F537 as a function of bound intermediate/product.
Supplementary information
Supplementary Video 1
Side view of ACLY morphed through four conformational states reported here (ACLY-apo, ACLY–citrate-CoA, ACLY-E599Q–ATP-citrate-CoA, ACLY–OAA-acetyl-CoA and back to ACLY-apo). CPK-white carbon coloring is used for bound ligands. The binding of ATP and citrate prior to CoA binding is modeled based on the ACLY ASH domain crystal structures bound to citrate (3MWD) and ADP (5TES).
Source data
Rights and permissions
About this article
Cite this article
Wei, X., Schultz, K., Bazilevsky, G.A. et al. Molecular basis for acetyl-CoA production by ATP-citrate lyase. Nat Struct Mol Biol 27, 33–41 (2020). https://doi.org/10.1038/s41594-019-0351-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0351-6
This article is cited by
-
The role of fatty acid metabolism in acute lung injury: a special focus on immunometabolism
Cellular and Molecular Life Sciences (2024)
-
Allosteric role of the citrate synthase homology domain of ATP citrate lyase
Nature Communications (2023)
-
Metabolic regulation of proteome stability via N-terminal acetylation controls male germline stem cell differentiation and reproduction
Nature Communications (2023)
-
Acetyl-CoA metabolism in cancer
Nature Reviews Cancer (2023)
-
Physiological and pathological roles of lipogenesis
Nature Metabolism (2023)