Abstract
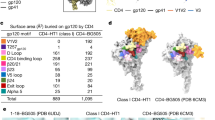
The human immunodeficiency virus (HIV-1) envelope (Env) glycoprotein, a (gp120–gp41)3 trimer, mediates fusion of viral and host cell membranes after gp120 binding to host receptor CD4. Receptor binding triggers conformational changes allowing coreceptor (CCR5) recognition through CCR5’s tyrosine-sulfated amino (N) terminus, release of the gp41 fusion peptide and fusion. We present 3.3 Å and 3.5 Å cryo-EM structures of E51, a tyrosine-sulfated coreceptor-mimicking antibody, complexed with a CD4-bound open HIV-1 native-like Env trimer. Two classes of asymmetric Env interact with E51, revealing tyrosine-sulfated interactions with gp120 mimicking CCR5 interactions, and two conformations of gp120–gp41 protomers (A and B protomers in AAB and ABB trimers) that differ in their degree of CD4-induced trimer opening and induction of changes to the fusion peptide. By integrating the new structural information with previous closed and open envelope trimer structures, we modeled the order of conformational changes on the path to coreceptor binding site exposure and subsequent viral–host cell membrane fusion.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The structural coordinates were deposited into the Worldwide Protein Data Bank (wwPDB) with accession code 6U0L (class I E51–BG505 SOSIP.664–sCD4 complex) and 6U0N (class II E51–BG505 SOSIP.664–sCD4 complex). EM density maps were deposited into EMDB with accession numbers EMD-20605 (class I) and EMD-20608 (class II). Other data are available upon reasonable request.
Change history
21 January 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
16 December 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Harrison, S. C. Viral membrane fusion. Virology 479–480, 498–507 (2015).
Choe, H. et al. The beta-chemokine receptors CCR3 and CCR5 facilitate infection by primary HIV-1 isolates. Cell 85, 1135–1148 (1996).
Feng, Y., Broder, C. C., Kennedy, P. E. & Berger, E. A. HIV-1 entry cofactor: functional cDNA cloning of a seven-transmembrane, G protein-coupled receptor. Science 272, 872–877 (1996).
Liu, J., Bartesaghi, A., Borgnia, M. J., Sapiro, G. & Subramaniam, S. Molecular architecture of native HIV-1 gp120 trimers. Nature 455, 109–113 (2008).
Ozorowski, G. et al. Open and closed structures reveal allostery and pliability in the HIV-1 envelope spike. Nature 547, 360–363 (2017).
Wang, H. et al. Cryo-EM structure of a CD4-bound open HIV-1 envelope trimer reveals structural rearrangements of the gp120 V1V2 loop. Proc. Natl Acad. Sci. USA 113, E7151–E7158 (2016).
Wang, H., Barnes, C. O., Yang, Z., Nussenzweig, M. C. & Bjorkman, P. J. Partially open HIV-1 envelope structures exhibit conformational changes relevant for coreceptor binding and fusion. Cell Host Microbe 24, 579–592 e4 (2018).
Sanders, R. W. et al. A next-generation cleaved, soluble HIV-1 Env trimer, BG505 SOSIP.664 gp140, expresses multiple epitopes for broadly neutralizing but not non-neutralizing antibodies. PLoS Pathog. 9, e1003618 (2013).
Alsahafi, N., Debbeche, O., Sodroski, J. & Finzi, A. Effects of the I559P gp41 change on the conformation and function of the human immunodeficiency virus (HIV-1) membrane envelope glycoprotein trimer. PLoS ONE 10, e0122111 (2015).
Ward, A. B. & Wilson, I. A. The HIV-1 envelope glycoprotein structure: nailing down a moving target. Immunol. Rev. 275, 21–32 (2017).
Burton, D. R. & Hangartner, L. Broadly neutralizing antibodies to HIV and their role in vaccine design. Annu. Rev. Immunol. 34, 635–659 (2016).
DeVico, A. L. CD4-induced epitopes in the HIV envelope glycoprotein, gp120. Curr. HIV Res. 5, 561–571 (2007).
Burton, D. R. et al. HIV vaccine design and the neutralizing antibody problem. Nat. Immunol. 5, 233–236 (2004).
Thali, M. et al. Characterization of conserved human immunodeficiency virus type 1 gp120 neutralization epitopes exposed upon gp120-CD4 binding. J. Virol. 67, 3978–3988 (1993).
Xiang, S. H., Doka, N., Choudhary, R. K., Sodroski, J. & Robinson, J. E. Characterization of CD4-induced epitopes on the HIV type 1 gp120 envelope glycoprotein recognized by neutralizing human monoclonal antibodies. AIDS Res. Hum. Retroviruses 18, 1207–1217 (2002).
Decker, J. M. et al. Antigenic conservation and immunogenicity of the HIV coreceptor binding site. J. Exp. Med. 201, 1407–1419 (2005).
Labrijn, A. F. et al. Access of antibody molecules to the conserved coreceptor binding site on glycoprotein gp120 is sterically restricted on primary human immunodeficiency virus type 1. J. Virol. 77, 10557–10565 (2003).
Kwong, P. D. et al. Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody. Nature 393, 648–659 (1998).
Shaik, M. M. et al. Structural basis of coreceptor recognition by HIV-1 envelope spike. Nature 565, 318–323 (2019).
Farzan, M. et al. Tyrosine sulfation of the amino terminus of CCR5 facilitates HIV-1 entry. Cell 96, 667–676 (1999).
Xiang, S. H. et al. Epitope mapping and characterization of a novel CD4-induced human monoclonal antibody capable of neutralizing primary HIV-1 strains. Virology 315, 124–134 (2003).
Huang, C. C. et al. Structural basis of tyrosine sulfation and VH-gene usage in antibodies that recognize the HIV type 1 coreceptor-binding site on gp120. Proc. Natl Acad. Sci. USA 101, 2706–2711 (2004).
Choe, H. et al. Tyrosine sulfation of human antibodies contributes to recognition of the CCR5 binding region of HIV-1 gp120. Cell 114, 161–170 (2003).
Diskin, R., Marcovecchio, P. M. & Bjorkman, P. J. Structure of a clade C HIV-1 gp120 bound to CD4 and CD4-induced antibody reveals anti-CD4 polyreactivity. Nat. Struct. Mol. Biol. 17, 608–613 (2010).
Huang, C. C. et al. Structures of the CCR5 N terminus and of a tyrosine-sulfated antibody with HIV-1 gp120 and CD4. Science 317, 1930–1934 (2007).
Scharf, L. et al. Broadly neutralizing antibody 8ANC195 recognizes closed and open states of HIV-1 Env. Cell 162, 1379–1390 (2015).
Gardner, M. R. et al. AAV-expressed eCD4-Ig provides durable protection from multiple SHIV challenges. Nature 519, 87–91 (2015).
Cormier, E. G. et al. Specific interaction of CCR5 amino-terminal domain peptides containing sulfotyrosines with HIV-1 envelope glycoprotein gp120. Proc. Natl Acad. Sci. USA 97, 5762–5767 (2000).
Lee, J. H., Ozorowski, G. & Ward, A. B. Cryo-EM structure of a native, fully glycosylated, cleaved HIV-1 envelope trimer. Science 351, 1043–1048 (2016).
Kong, R. et al. Fusion peptide of HIV-1 as a site of vulnerability to neutralizing antibody. Science 352, 828–833 (2016).
Kumar, S. et al. Capturing the inherent structural dynamics of the HIV-1 envelope glycoprotein fusion peptide. Nat. Commun. 10, 763 (2019).
Dingens, A. S. et al. Complete functional mapping of infection- and vaccine-elicited antibodies against the fusion peptide of HIV. PLoS Pathog. 14, e1007159 (2018).
Chan, D. C., Fass, D., Berger, J. M. & Kim, P. S. Core structure of gp41 from the HIV envelope glycoprotein. Cell 89, 263–273 (1997).
Weissenhorn, W., Dessen, A., Harrison, S. C., Skehel, J. J. & Wiley, D. C. Atomic structure of the ectodomain from HIV-1 gp41. Nature 387, 426–430 (1997).
Dorfman, T., Moore, M. J., Guth, A. C., Choe, H. & Farzan, M. A tyrosine-sulfated peptide derived from the heavy-chain CDR3 region of an HIV-1-neutralizing antibody binds gp120 and inhibits HIV-1 infection. J. Biol. Chem. 281, 28529–28535 (2006).
Fellinger, C. H. et al. eCD4-Ig Limits HIV-1 Escape More Effectively than CD4-Ig or a Broadly Neutralizing Antibody. J. Virol. 93, e00443-19 (2019).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Scheres, S. H. RELION: implementation of a bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive python-based system for macromolecular structure solution. Acta Crystallogr. D66, 213–221 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Dolinsky, T. J. et al. PDB2PQR: expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res. 35, W522–W525 (2007).
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001).
Acknowledgements
We thank M. Farzan, A.P. West and C.O. Barnes for helpful discussions. Structural studies were performed in the Biological and Cryogenic Transmission Electron Microscopy Center at Caltech with assistance from directors A. Malyutin and S. Chen. We thank the Gordon and Betty Moore and Beckman Foundations for gifts to Caltech to support electron microscopy, Z. Yu (Janeila Farm) for advice on cryo-EM, J. Vielmetter and the Caltech Protein Expression Center for protein production, and M. Shahgholi and the Caltech Protein Exploration Laboratory for mass spectrometry analyses. This work was supported by the National Institute of Allergy and Infectious Diseases of the National Institutes of Health Grant No. HIVRAD P01 P01AI10014 (P.J.B.) and the National Institutes of Health Grant No. P50 8 P50 AI150464-13 (P.J.B.).
Author information
Authors and Affiliations
Contributions
Z.Y. and P.J.B. conceived the study. Z.Y. solved and interpreted cryo-EM structures with help from H.W. Z.Y., H.W., H.B.G. and P.J.B. analyzed the data. A.Z.L. and H.B.G. prepared and isolated E51 Fabs. Z.Y. and P.J.B. wrote the paper with contributions from other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Beth Moorefield was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Mass spectrometry characterization of E51 Fab.
a, Peak fractions from anion exchange chromatography of E51 Fab. b, Mass spectra of the three E51 peaks from panel a. The molecular mass for Peak 1 corresponded within 27 Da to the predicted mass for unmodified E51 Fab (48,791 Da), suggesting this E51 Fab fraction contained no sulfated tyrosines. The molecular masses for Peaks 2 and 3 were each increased by 80 Da from the preceding peak, corresponding to the molecular weight of a SO3− group (80 Da). These results are consistent with the identification of Peaks 2 and 3 as E51 Fab with one and two sulfated tyrosines, respectively. Peak 3 was used for preparing complexes for cryo-EM.
Extended Data Fig. 2 Data processing for BG505 SOSIP-sCD4-E51 complex structures.
a, Example of a motion-corrected and dose-weighted micrograph of E51-sCD4-BG505 complexes (representative individual particles in boxes). Scale bar = 50 nm. Defocus was ~2.6 μm underfocus. b, Data processing scheme. Collected micrographs were motion-corrected using MotionCor2 (ref. 1) without binning. Contrast transfer functions (CTFs) were estimated using Gctf 1.06 (ref. 2). ~1,000 particles were manually picked and subjected to 2D classification in RELION3,4. Particles from good classes were used as references for automatic particle picking using RELION AutoPicking. 941,841 autopicked particles were subjected to three rounds of reference-free 2D classification. In each round, particles from good 2D classes were selected, which gave 656,059 particles. Subsequently, using the Env trimer portion of an 80 Å low-pass filtered partially-open trimeric BG505 SOSIP structure (PDB 6CM3) as the reference model, first round of 3D classification was performed assuming C1 symmetry. Particles from good classes were combined and used for a second round of 3D classification with C1 symmetry. The resulting maps could be grouped into two classes: class I (320,895 particles) and class II (182,970 particles). Maps from the two classes were refined separately assuming C1 symmetry with the sCD4 D2 domain and E51 Fab CHCL domains masked out. After CTF refinement, movie refinement, and particle polishing, the class I and class II post-processed maps were refined to 3.3 Å and 3.5 Å resolution (FSC 0.143), respectively (Supplementary Fig. 3).
Extended Data Fig. 3 Validation of the BG505 SOSIP-sCD4-E51 complex structures.
Class I (panel a) and class II (panel b) E51-sCD4-BG505 complex structures. Top left in both panels: Gold-standard Fourier Shell Correlations (FSCs) of two classes of maps. Top right: Orientation distributions for class I and class II structures. Bottom: Local resolution estimations for class I and class II density maps (calculated using the local resolution program in RELION3,4).
Extended Data Fig. 4 Representative densities for class I and class II E51-sCD4-BG505 complex structures.
a, Densities for residues involved in E51 CDRH3 sulfotyrosine interactions. b, Densities for fusion peptides in protomers A (top) and B (bottom). c, Densities for HR1C helices in protomers A (top) and B (bottom).
Extended Data Fig. 5 Comparison of fusion peptide conformations in Env structures.
The fusion peptide is orange in the conformation A protomer, cyan in the conformation B protomer (from the E51-sCD4-BG505 complex structures reported here), teal in a partially-open 17b-sCD4-BG505-8ANC195 complex (PDB 6CM3), and pale cyan in a fully-open 17b-sCD4-B41 complex (PDB 5VN3). Fusion peptides from Env trimers in a closed, prefusion conformation are color coded as shown for their PDB IDs. References for structures are listed in Supplementary Table 2.
Supplementary information
Supplementary information
Supplementary Note 1 and Tables 1 and 2.
Supplementary Video
Asymmetric opening of HIV-1 Env trimer induced by CD4 revealed intermediate states. The first section of the video shows soluble HIV-1 Env trimer opening when bound to sCD4, as highlighted by conformational changes in gp120 V1V2 region, V3 and coreceptor binding region, as well as gp41 HR1C. The second section of the video shows CD4-induced protomer intermediate open states, which ultimately contribute to the asymmetric open states of Env trimer.
Rights and permissions
About this article
Cite this article
Yang, Z., Wang, H., Liu, A.Z. et al. Asymmetric opening of HIV-1 Env bound to CD4 and a coreceptor-mimicking antibody. Nat Struct Mol Biol 26, 1167–1175 (2019). https://doi.org/10.1038/s41594-019-0344-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0344-5
This article is cited by
-
Intermediate conformations of CD4-bound HIV-1 Env heterotrimers
Nature (2023)
-
HIV-1 Env trimers asymmetrically engage CD4 receptors in membranes
Nature (2023)
-
Neutralizing antibodies induced in immunized macaques recognize the CD4-binding site on an occluded-open HIV-1 envelope trimer
Nature Communications (2022)
-
Comparing methods for immobilizing HIV-1 SOSIPs in ELISAs that evaluate antibody binding
Scientific Reports (2022)
-
HIV-1 CD4-binding site germline antibody–Env structures inform vaccine design
Nature Communications (2022)