Abstract
We present structures of mouse TRPV3 in temperature-dependent open, closed and intermediate states that suggest two-step activation of TRPV3 by heat. During the strongly temperature-dependent first step, sensitization, the channel pore remains closed while S6 helices undergo α-to-π transitions. During the weakly temperature-dependent second step, channel opening, tight association of the S1–S4 and pore domains is stabilized by changes in the carboxy-terminal and linker domains.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Cryo-EM density maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession numbers EMD-20492 for TRPV3WT-Closed-42 °C, EMD-20493 for TRPV3WT-Sensitized-42 °C, EMD-20494 for TRPV3Y564A-Sensitized-4 °C, EMD-20495 for TRPV3Y564A-Sensitized-37 °C, EMD-20496 for TRPV3Y564A-Open-37 °C and EMD-20497 for TRPV3Y564A-Intermediate-37 °C. Model coordinates have been deposited in the Protein Data Bank (PDB) under accession numbers 6PVL for TRPV3WT-Closed-42 °C, 6PVM for TRPV3WT-Sensitized-42 °C, 6PVN for TRPV3Y564A-Sensitized-4 °C, 6PVO for TRPV3Y564A-Sensitized-37 °C, 6PVP for TRPV3Y564A-Open-37 °C and 6PVQ for TRPV3Y564A-Intermediate-37 °C. All other data are available from the corresponding author upon request.
Change history
13 January 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Caterina, M. J. et al. Nature 389, 816–824 (1997).
Patapoutian, A., Peier, A. M., Story, G. M. & Viswanath, V. Nat. Rev. Neurosci. 4, 529–539 (2003).
Jordt, S. E., McKemy, D. D. & Julius, D. Curr. Opin. Neurobiol. 13, 487–492 (2003).
Clapham, D. E. Nature 426, 517–524 (2003).
Peier, A. M. et al. Science 296, 2046–2049 (2002).
Guler, A. D. et al. J. Neurosci. 22, 6408–6414 (2002).
Caterina, M. J., Rosen, T. A., Tominaga, M., Brake, A. J. & Julius, D. Nature 398, 436–441 (1999).
Smith, G. D. et al. Nature 418, 186–190 (2002).
Liu, B. Y., Yao, J., Zhu, M. X. & Qin, F. J. Gen. Physiol. 138, 509–520 (2011).
Zubcevic, L. et al. Nat. Commun. 9, 4773 (2018).
Singh, A. K., McGoldrick, L. L. & Sobolevsky, A. I. Nat. Struct. Mol. Biol. 25, 805–813 (2018).
Diaz-Franulic, I., Poblete, H., Mino-Galaz, G., Gonzalez, C. & Latorre, R. Annu. Rev. Biophys. 45, 371–398 (2016).
Feng, Q. Curr. Top. Membr. 74, 19–50 (2014).
Voets, T. Rev. Physiol. Biochem. Pharmacol. 162, 91–119 (2012).
Xu, H. X. et al. Nature 418, 181–186 (2002).
Kasimova, M. A. et al. J. Gen. Physiol. 150, 1554–1566 (2018).
Liu, B. & Qin, F. Proc. Natl Acad. Sci. USA 114, 1589–1594 (2017).
Cao, E., Liao, M., Cheng, Y. & Julius, D. Nature 504, 113–118 (2013).
Vlachova, V. et al. J. Neurosci. 23, 1340–1350 (2003).
Brauchi, S., Orio, P. & Latorre, R. Proc. Natl Acad. Sci. USA 101, 15494–15499 (2004).
Brauchi, S., Orta, G., Salazar, M., Rosenmann, E. & Latorre, R. J. Neurosci. 26, 4835–4840 (2006).
Grandl, J. et al. Nat. Neurosci. 11, 1007–1013 (2008).
Grandl, J. et al. Nat. Neurosci. 13, 708–714 (2010).
Yao, J., Liu, B. Y. & Qin, F. Proc. Natl Acad. Sci. USA 108, 11109–11114 (2011).
Macikova, L., Vyklicka, L., Barvik, I., Sobolevsky, A. I. & Vlachova, V. Int. J. Mol. Sci. 20, 3990 (2019).
Brauchi, S. et al. Proc. Natl Acad. Sci. USA 104, 10246–10251 (2007).
Zubcevic, L., Borschel, W. F., Hsu, A. L., Borgnia, M. J. & Lee, S.-Y. Elife 8, e47746 (2019).
Shi, D. J., Ye, S., Cao, X., Zhang, R. G. & Wang, K. W. Protein Cell 4, 942–950 (2013).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. P. J. Mol. Graph. Model. 14, 354–360 (1996).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Nat. Methods 11, 63–65 (2014).
Goehring, A. et al. Nat. Protoc. 9, 2574–2585 (2014).
Russo, C. J. & Passmore, L. A. Science 346, 1377–1380 (2014).
Suloway, C. et al. J. Struct. Biol. 151, 41–60 (2005).
Zivanov, J. et al. eLife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. Nat. Methods 14, 290–296 (2017).
Zhang, K. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. & Chen, S. Nat. Methods 9, 853–854 (2012).
Chen, S. et al. Ultramicroscopy 135, 24–35 (2013).
Pettersen, E. F. et al. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Acta Crystallogr. D 68, 352–367 (2012).
Tang, G. et al. J. Struct. Biol. 157, 38–46 (2007).
Zakharian, E. Methods Mol. Biol. 998, 109–118 (2013).
Acknowledgements
We thank R. Grassucci for assistance with microscope operation, U. Baxa and T. Edwards for help with data collection and H. Kao for computational support. We thank J. Yoder for comments on the manuscript and M. Yelshanskaya for helpful discussions. L.L.M. is supported by the National Institutes of Health (NIH) (F31 CA232391-01). A.I.S. is supported by the NIH (R01 CA206573, R01 NS083660, R01 NS107253), National Science Foundation (1818213) and the Irma T. Hirschl Career Scientist Award. Data were collected at the Frederick National Laboratory for Cancer Research National Cryo-EM Facility (NIH) and at the Simons Electron Microscopy Center and National Resource for Automated Molecular Microscopy (New York Structural Biology Center) supported by grants from the Simons Foundation (349247), NYSTAR and the NIH (GM103310).
Author information
Authors and Affiliations
Contributions
A.K.S. and A.I.S. designed the project. A.K.S., L.L.M., L.D., E.Z. and A.I.S. analyzed data. A.K.S. and L.L.M. made constructs, prepared protein samples, carried out cryo-EM data collection and processing. A.K.S. and A.I.S. built molecular models. L.D. and M.L. carried out bilayer experiments. A.K.S., L.L.M., L.D., E.Z. and A.I.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Katarzyna Marcinkiewicz was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Overview of cryo-EM data collected for TRPV3WT and TRPV3Y564A.
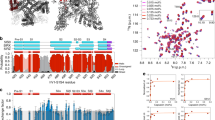
a, Example micrographs from different TRPV3-construct collections with example particles circled in red. b, Reference-free 2D class averages in different orientations. c, Euler angle distribution of particles contributing to the final reconstructions with larger red cylinders representing orientations comprising more particles.
Extended Data Fig. 2 Resolution of TRPV3WT and TRPV3Y564A cryo-EM reconstructions.
a, FSC curves calculated between half maps. b, FSC curves calculated between two unfiltered half-maps and the final map and a model whose coordinates were randomized and refined against only half map 1. c, Local resolution predicted by ResMap30.
Extended Data Fig. 3 Comparison of TRPV3WT closed-state structures at 4 °C and 42 °C.
a-b, Overall superposition of TRPV3WT-closed-4 °C (red) and TRPV3WT-closed-42 °C (blue) structures viewed from the side (a) or the bottom (b). c, Pore radii calculated using HOLE29. The vertical dashed line denotes the radius of a water molecule, 1.4 Å. d-e, Expanded view of the transmembrane domain of one TRPV3WT-Closed-4 °C (d) or TRPV3WT-Closed-42 °C (e) subunit with lipid-like densities shown as purple mesh.
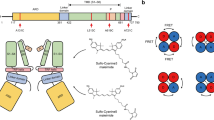
Extended Data Fig. 4 Architecture and domain organization of TRPV3.
a-b, Top (a) and side (b) views of the TRPV3 tetramer, with each subunit shown in a different color. c, Domain organization diagram of the TRPV3 subunit. d, Structure of TRPV3 subunit, with domains colored as in c. Alanine substituting tyrosine 564 in TRPV3Y564A is shown in red space-filling representation (a, b and d) or indicated by the red circle (c).
Extended Data Fig. 5
3D reconstruction workflow for TRPV3Y564A.
Extended Data Fig. 6 C-terminus unlatching during channel opening.
a-b, Expanded view of the cytosolic intersubunit interface in TRPV3Y564A-Sensitized-4 °C (a) and TRPV3Y564A-Open-37 °C (b). The C-termini and the putative N-terminus fragment from the TRPV3Y564A-Open-37 °C adjacent subunit are thickened for clarity. Conserved residues at the C-terminus-ARD interfaces are shown as sticks. Movement of the AR5 loop in the open state relative to the sensitized or closed state is indicated by a red arrow.
Extended Data Fig. 7 Comparison of heat- and ligand-activated open states.
a-c, Overall superposition of TRPV3Y564A-Open-37 °C (orange) and TRPV3Y564A-Open-2-APB (grey) viewed from the top (a) and the side (b) and an expanded view of the transmembrane domain of one subunit (c). 2-APB molecules bound to TRPV3Y564A-Open-2-APB structure are shown as space-filling models. d-f, Expanded views of binding sites 2 (d), 3 (e) and 4 (f). The 2-APB molecules bound to TRPV3Y564A-Open-2-APB are shown as sticks and the TRPV3Y564A-Open-37 °C density is shown as blue mesh. TRPV3Y564A-Open-37 °C residues that would clash with the 2-APB molecules are shown in stick representation.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Tables 1 and 2
Rights and permissions
About this article
Cite this article
Singh, A.K., McGoldrick, L.L., Demirkhanyan, L. et al. Structural basis of temperature sensation by the TRP channel TRPV3. Nat Struct Mol Biol 26, 994–998 (2019). https://doi.org/10.1038/s41594-019-0318-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0318-7
This article is cited by
-
Thermal gradient ring for analysis of temperature-dependent behaviors involving TRP channels in mice
The Journal of Physiological Sciences (2024)
-
Computational drug development for membrane protein targets
Nature Biotechnology (2024)
-
Involvement of skin TRPV3 in temperature detection regulated by TMEM79 in mice
Nature Communications (2023)
-
Structural titration reveals Ca2+-dependent conformational landscape of the IP3 receptor
Nature Communications (2023)
-
Structural basis of TRPV3 inhibition by an antagonist
Nature Chemical Biology (2023)