Abstract
Cohesin is a regulator of genome architecture with roles in sister chromatid cohesion and chromosome compaction. The recruitment and mobility of cohesin complexes on DNA is restricted by nucleosomes. Here, we show that the role of cohesin in chromosome organization requires the histone chaperone FACT (‘facilitates chromatin transcription’) in Saccharomyces cerevisiae. We find that FACT interacts directly with cohesin, and is dynamically required for its localization on chromatin. Depletion of FACT in metaphase cells prevents cohesin accumulation at pericentric regions and causes reduced binding on chromosome arms. Using the Hi-C technique, we show that cohesin-dependent TAD (topological associated domain)-like structures in G1 and metaphase chromosomes are reduced in the absence of FACT. Sister chromatid cohesion is intact in FACT-depleted cells, although chromosome segregation failure is observed. Our data show that FACT contributes to the formation of cohesin-dependent TADs, thus uncovering a new role for this complex in nuclear organization during interphase and mitotic chromosome folding.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
ChIP-seq and MNase-seq data supporting the findings of this study have been deposited in the GEO database and are accessible through accession no. GSE118534. Hi-C raw sequences are accessible via the SRA database through accession no. PRJNA486513. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD014896. Source data for Figs. 1b,c, 3d and 5 are available online. Any further data that support the findings of this study are available from the corresponding authors upon request.
References
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707 e14 (2017).
Lazar-Stefanita, L. et al. Cohesins and condensins orchestrate the 4D dynamics of yeast chromosomes during the cell cycle. EMBO J. 36, 2684–2697 (2017).
Schalbetter, S. A. et al. SMC complexes differentially compact mitotic chromosomes according to genomic context. Nat. Cell Biol. 19, 1071–1080 (2017).
Haering, C. H., Lowe, J., Hochwagen, A. & Nasmyth, K. Molecular architecture of SMC proteins and the yeast cohesin complex. Mol. Cell 9, 773–788 (2002).
Nasmyth, K. Disseminating the genome: joining, resolving and separating sister chromatids during mitosis and meiosis. Annu. Rev. Genet. 35, 673–745 (2001).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 e24 (2017).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Ganji, M. et al. Real-time imaging of DNA loop extrusion by condensin. Science 360, 102–105 (2018).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Stigler, J., Camdere, G. O., Koshland, D. E. & Greene, E. C. Single-molecule imaging reveals a collapsed conformational state for DNA-bound cohesin. Cell Rep. 15, 988–998 (2016).
Huang, J., Hsu, J. M. & Laurent, B. C. The RSC nucleosome-remodeling complex is required for cohesin’s association with chromosome arms. Mol. Cell 13, 739–750 (2004).
Litwin, I., Bakowski, T., Maciaszczyk-Dziubinska, E. & Wysocki, R. The LSH/HELLS homolog Irc5 contributes to cohesin association with chromatin in yeast. Nucleic Acids Res. 45, 6404–6416 (2017).
Hakimi, M. A. et al. A chromatin remodelling complex that loads cohesin onto human chromosomes. Nature 418, 994–998 (2002).
Megee, P. C., Mistrot, C., Guacci, V. & Koshland, D. The centromeric sister chromatid cohesion site directs Mcd1p binding to adjacent sequences. Mol. Cell 4, 445–450 (1999).
Blat, Y. & Kleckner, N. Cohesins bind to preferential sites along yeast chromosome III, with differential regulation along arms versus the centric region. Cell 98, 249–259 (1999).
Tanaka, T., Cosma, M. P., Wirth, K. & Nasmyth, K. Identification of cohesin association sites at centromeres and along chromosome arms. Cell 98, 847–858 (1999).
Petela, N. J. et al. Scc2 is a potent activator of cohesin’s ATPase that promotes loading by binding scc1 without Pds5. Mol. Cell 70, 1134–1148 e7 (2018).
Gonzalez, S. et al. Nucleosomal signatures impose nucleosome positioning in coding and noncoding sequences in the genome. Genome Res. 26, 1532–1543 (2016).
Yuan, G. C. et al. Genome-scale identification of nucleosome positions in S. cerevisiae. Science 309, 626–630 (2005).
Chen, K. et al. DANPOS: dynamic analysis of nucleosome position and occupancy by sequencing. Genome Res. 23, 341–351 (2013).
Oum, J. H. et al. RSC facilitates Rad59-dependent homologous recombination between sister chromatids by promoting cohesin loading at DNA double-strand breaks. Mol. Cell Biol. 31, 3924–3937 (2011).
Shintomi, K., Takahashi, T. S. & Hirano, T. Reconstitution of mitotic chromatids with a minimum set of purified factors. Nat. Cell Biol. 17, 1014–1023 (2015).
Formosa, T. The role of FACT in making and breaking nucleosomes. Biochim. Biophys. Acta 1819, 247–255 (2013).
Nishimura, K. & Kanemaki, M. T. Rapid depletion of budding yeast proteins via the fusion of an auxin-inducible degron (AID). Curr. Protoc. Cell Biol. 64, 20.9.1–20.9.16 (2014).
Hu, B. et al. Biological chromodynamics: a general method for measuring protein occupancy across the genome by calibrating ChIP-seq. Nucleic Acids Res. 43, e132 (2015).
Lopez-Serra, L., Kelly, G., Patel, H., Stewart, A. & Uhlmann, F. The Scc2–Scc4 complex acts in sister chromatid cohesion and transcriptional regulation by maintaining nucleosome-free regions. Nat. Genet. 46, 1147–1151 (2014).
Fernius, J. & Marston, A. L. Establishment of cohesion at the pericentromere by the Ctf19 kinetochore subcomplex and the replication fork-associated factor, Csm3. PLoS Genet. 5, e1000629 (2009).
Hu, B. et al. ATP hydrolysis is required for relocating cohesin from sites occupied by its Scc2/4 loading complex. Curr. Biol. 21, 12–24 (2011).
Lengronne, A. et al. Cohesin relocation from sites of chromosomal loading to places of convergent transcription. Nature 430, 573–578 (2004).
Glynn, E. F. et al. Genome-wide mapping of the cohesin complex in the yeast Saccharomyces cerevisiae. PLoS Biol. 2, E259 (2004).
Gullerova, M. & Proudfoot, N. J. Cohesin complex promotes transcriptional termination between convergent genes in S. pombe. Cell 132, 983–995 (2008).
Busslinger, G. A. et al. Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl. Nature 544, 503–507 (2017).
Formosa, T. The role of FACT in making and breaking nucleosomes. Biochim. Biophys. Acta 1819, 247–255 (2012).
Saunders, A. et al. Tracking FACT and the RNA polymerase II elongation complex through chromatin in vivo. Science 301, 1094–1096 (2003).
Guacci, V., Koshland, D. & Strunnikov, A. A direct link between sister chromatid cohesion and chromosome condensation revealed through the analysis of MCD1 in S. cerevisiae. Cell 91, 47–57 (1997).
Michaelis, C., Ciosk, R. & Nasmyth, K. Cohesins: chromosomal proteins that prevent premature separation of sister chromatids. Cell 91, 35–45 (1997).
Kemble, D. J., McCullough, L. L., Whitby, F. G., Formosa, T. & Hill, C. P. FACT disrupts nucleosome structure by binding H2A-H2B with conserved peptide motifs. Mol. Cell 60, 294–306 (2015).
Bondarenko, M. T. et al. [Structure and function of histone chaperone FACT]. Mol. Biol. ( Mosk. ) 49, 891–904 (2015).
Simic, R. et al. Chromatin remodeling protein Chd1 interacts with transcription elongation factors and localizes to transcribed genes. EMBO J. 22, 1846–1856 (2003).
Boginya, A., Detroja, R., Matityahu, A., Frenkel-Morgenstern, M. & Onn, I. The chromatin remodeler Chd1 regulates cohesin in budding yeast and humans. Sci. Rep. 9, 8929 (2019).
Tan, B. C., Chien, C. T., Hirose, S. & Lee, S. C. Functional cooperation between FACT and MCM helicase facilitates initiation of chromatin DNA replication. EMBO J. 25, 3975–3985 (2006).
McCullough, L. L. et al. Establishment and maintenance of chromatin architecture are promoted independent of transcription by the histone chaperone FACT and H3-K56 acetylation in Saccharomyces cerevisiae. Genetics 211, 877–892 (2019).
Heidinger-Pauli, J. M., Mert, O., Davenport, C., Guacci, V. & Koshland, D. Systematic reduction of cohesin differentially affects chromosome segregation, condensation and DNA repair. Curr. Biol. 20, 957–963 (2010).
Belton, J. M. et al. Hi-C: a comprehensive technique to capture the conformation of genomes. Methods 58, 268–276 (2012).
Cournac, A., Marie-Nelly, H., Marbouty, M., Koszul, R. & Mozziconacci, J. Normalization of a chromosomal contact map. BMC Genom. 13, 436 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Quintales, L., Vazquez, E. & Antequera, F. Comparative analysis of methods for genome-wide nucleosome cartography. Brief. Bioinform. 16, 576–587 (2015).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Acknowledgements
We thank the members of the L.A., R.K. and P.A. laboratories for fruitful discussions and advice. The work in the L.A. laboratory was supported by a Wellcome Trust Senior Investigator award to L.A. (100955, ‘Functional dissection of mitotic chromatin’) and the London Institute of Medical Research (LMS), which receives its core funding from the UK Medical Research Council. This research was further supported by funding from The European Research Council (R.K.), Agence Nationale pour la Recherche (R.K.) and the the Spanish Ministerio de Economía, Industria y Competitividad (BFU2017-89622-P; F.A.).
Author information
Authors and Affiliations
Contributions
J.G.-L. performed all yeast experiments, providing samples for nucleosome mapping, HiC and sequencing. J.G.-L. performed IP and cell biology experiments. L.L.-S. and A.T. performed Hi-C experiments and analysis. A.C. analyzed Hi-C data. A.G., S.G. and M.S. performed and analyzed nucleosome position experiments. M.D. and M.M.K. performed bioinformatic analysis. P.G.-E. performed experiments related to the pulldowns for mass spectrometry and analyzed results data. H.K. and A.M. performed technical mass spectrometry analysis. A.J. performed molecular biology experiments to generate constructs. J.G.-L. and L.A. conceived the project. L.A. wrote the manuscript. L.A., F.A. and R.K. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Anke Sparmann was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Auxin-mediated degradation of Spt16 in G2–M arrested cells.
Cells carrying either SMC1-6HA or SMC1-6HA SPT16-AID (which allows auxin-mediated degradation of Spt16) were arrested in metaphase (G2–M-nocodazole) and exposed to auxin to degrade Spt16. Nuclear and cellular morphology was used following auxin addition to confirm that the cell population remained arrested in metaphase (top). Western analysis for Spt16 confirms its degradation in SPT16-AID cells upon auxin addition (bottom). The bar graph data show the mean and error bars the s.d. of three independent experiments.
Supplementary Figure 2 Auxin-mediated degradation of Sth1 in G2–M arrested cells.
Cells carrying either SMC1-6HA or SMC1-6HA STH1-AID (which allows auxin-mediated degradation of Sth1) were arrested in metaphase (G2–M-nocodazole) and exposed to auxin to degrade Sth1. Western analysis for Sth1 confirms its degradation in STH1-AID cells upon auxin addition.
Supplementary Figure 3 FACT function in chromosome organization.
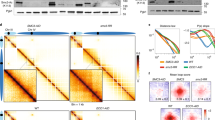
The 16 yeast chromosomes are displayed on Hi-C maps (10 kb bins) obtained from G2–M arrests in the presence of auxin of SMC1-6HA (wild-type, bottom left) and SMC1-6HA SPT16-AID (Spt16-aid) cells (top right). Black to white color scales reflect high to low contact frequencies, respectively (log2). Inset: Magnifications of chr7.
Supplementary Figure 4 FACT function in the establishment of cohesin-dependent structures in G1.
The 16 yeast chromosomes are displayed on Hi-C maps (10 kb bins) obtained from G1 arrests overexpressing MCD1 from the GAL1-10 promoter in the presence of auxin of wild-type (bottom left) and SPT16-AID (Spt16-aid) cells (top right). Black to white color scales reflect high to low contact frequencies, respectively (log2). Inset: magnifications of chr7.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4, Supplementary Table 1 and Supplementary Data Set 1
Rights and permissions
About this article
Cite this article
Garcia-Luis, J., Lazar-Stefanita, L., Gutierrez-Escribano, P. et al. FACT mediates cohesin function on chromatin. Nat Struct Mol Biol 26, 970–979 (2019). https://doi.org/10.1038/s41594-019-0307-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0307-x
This article is cited by
-
Cohesin regulates homology search during recombinational DNA repair
Nature Cell Biology (2021)
-
A compendium of chromatin contact maps reflecting regulation by chromatin remodelers in budding yeast
Nature Communications (2021)
-
Wie das Histon-Chaperon FACT aktives und stilles Chromatin bewahrt
BIOspektrum (2021)
-
Computer vision for pattern detection in chromosome contact maps
Nature Communications (2020)