Abstract
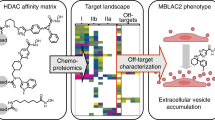
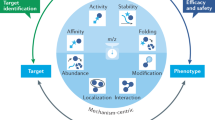
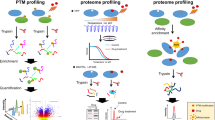
The identification of the full target spectrum of active molecules, known as target deconvolution, has become an indispensable step during the drug discovery process. It is now achievable thanks to mass spectrometry–based technologies. Here we discuss these approaches in the context of epigenetic drug discovery.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Portela, A. & Esteller, M. Epigenetic modifications and human disease. Nat. Biotechnol. 28, 1057–1068 (2010).
Arrowsmith, C. H., Bountra, C., Fish, P. V., Lee, K. & Schapira, M. Epigenetic protein families: a new frontier for drug discovery. Nat. Rev. Drug Discov. 11, 384–400 (2012).
Prachayasittikul, V. et al. Exploring the epigenetic drug discovery landscape. Expert Opin. Drug Discov. 12, 345–362 (2017).
Atadja, P. & Perez, L. in Successful Drug Discovery, Vol. 2, 59–88 (Wiley Online Library, 2016).
Dittmann, A. et al. The commonly used PI3-kinase probe LY294002 is an inhibitor of BET bromodomains. ACS Chem. Biol. 9, 495–502 (2014).
Bantscheff, M., Scholten, A. & Heck, A. J. Revealing promiscuous drug-target interactions by chemical proteomics. Drug Discov. Today 14, 1021–1029 (2009).
Rix, U. & Superti-Furga, G. Target profiling of small molecules by chemical proteomics. Nat. Chem. Biol. 5, 616–624 (2009).
Sharma, K. et al. Proteomics strategy for quantitative protein interaction profiling in cell extracts. Nat. Methods 6, 741–744 (2009).
Bantscheff, M. et al. Quantitative chemical proteomics reveals mechanisms of action of clinical ABL kinase inhibitors. Nat. Biotechnol. 25, 1035–1044 (2007).
Taunton, J., Hassig, C. A. & Schreiber, S. L. A mammalian histone deacetylase related to the yeast transcriptional regulator Rpd3p. Science 272, 408–411 (1996).
Weigt, D., Hopf, C. & Médard, G. Studying epigenetic complexes and their inhibitors with the proteomics toolbox. Clin. Epigenetics 8, 76 (2016).
Bantscheff, M. et al. Chemoproteomics profiling of HDAC inhibitors reveals selective targeting of HDAC complexes. Nat. Biotechnol. 29, 255–265 (2011).
Niphakis, M. J. & Cravatt, B. F. Enzyme inhibitor discovery by activity-based protein profiling. Annu. Rev. Biochem. 83, 341–377 (2014).
Salisbury, C. M. & Cravatt, B. F. Activity-based probes for proteomic profiling of histone deacetylase complexes. Proc. Natl Acad. Sci. USA 104, 1171–1176 (2007).
Chung, C. W. et al. Discovery and characterization of small molecule inhibitors of the BET family bromodomains. J. Med. Chem. 54, 3827–3838 (2011).
Dawson, M. A. et al. Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia. Nature 478, 529–533 (2011).
Savitski, M. M. et al. Tracking cancer drugs in living cells by thermal profiling of the proteome. Science 346, 1255784 (2014).
Franken, H. et al. Thermal proteome profiling for unbiased identification of direct and indirect drug targets using multiplexed quantitative mass spectrometry. Nat. Protoc. 10, 1567–1593 (2015).
Becher, I. et al. Thermal profiling reveals phenylalanine hydroxylase as an off-target of panobinostat. Nat. Chem. Biol. 12, 908–910 (2016).
Olsen, J. V. & Mann, M. Status of large-scale analysis of post-translational modifications by mass spectrometry. Mol. Cell. Proteomics 12, 3444–3452 (2013).
Greiner, D., Bonaldi, T., Eskeland, R., Roemer, E. & Imhof, A. Identification of a specific inhibitor of the histone methyltransferase SU(VAR)3-9. Nat. Chem. Biol. 1, 143–145 (2005).
Garcia, B. A., Busby, S. A., Shabanowitz, J., Hunt, D. F. & Mishra, N. Resetting the epigenetic histone code in the MRL-lpr/lpr mouse model of lupus by histone deacetylase inhibition. J. Proteome Res. 4, 2032–2042 (2005).
Lee, A. Y. et al. Quantitative analysis of histone deacetylase-1 selective histone modifications by differential mass spectrometry. J. Proteome Res. 7, 5177–5186 (2008).
Xu, K. et al. EZH2 oncogenic activity in castration-resistant prostate cancer cells is Polycomb-independent. Science 338, 1465–1469 (2012).
Schölz, C. et al. Acetylation site specificities of lysine deacetylase inhibitors in human cells. Nat. Biotechnol. 33, 415–423 (2015).
Musiani, D. et al. Proteomics profiling of arginine methylation defines PRMT5 substrate specificity. Sci. Signal. 12, eaat8388 (2019).
Noberini, R., Sigismondo, G. & Bonaldi, T. The contribution of mass spectrometry-based proteomics to understanding epigenetics. Epigenomics 8, 429–445 (2016).
Schirle, M., Bantscheff, M. & Kuster, B. Mass spectrometry-based proteomics in preclinical drug discovery. Chem. Biol. 19, 72–84 (2012).
Krönke, J. et al. Lenalidomide causes selective degradation of IKZF1 and IKZF3 in multiple myeloma cells. Science 343, 301–305 (2014).
Schopper, S. et al. Measuring protein structural changes on a proteome-wide scale using limited proteolysis-coupled mass spectrometry. Nat. Protoc. 12, 2391–2410 (2017).
Anders, L. et al. Genome-wide localization of small molecules. Nat. Biotechnol. 32, 92–96 (2014).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Noberini, R., Bonaldi, T. Epigenetic drug target deconvolution by mass spectrometry–based technologies. Nat Struct Mol Biol 26, 854–857 (2019). https://doi.org/10.1038/s41594-019-0279-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0279-x