Abstract
Clostridium difficile is an opportunistic pathogen that establishes in the colon when the gut microbiota are disrupted by antibiotics or disease. C. difficile infection (CDI) is largely caused by two virulence factors, TcdA and TcdB. Here, we report a 3.87-Å-resolution crystal structure of TcdB holotoxin that captures a unique conformation of TcdB at endosomal pH. Complementary biophysical studies suggest that the C-terminal combined repetitive oligopeptides (CROPs) domain of TcdB is dynamic and can sample open and closed conformations that may facilitate modulation of TcdB activity in response to environmental and cellular cues during intoxication. Furthermore, we report three crystal structures of TcdB–antibody complexes that reveal how antibodies could specifically inhibit the activities of individual TcdB domains. Our studies provide novel insight into the structure and function of TcdB holotoxin and identify intrinsic vulnerabilities that could be exploited to develop new therapeutics and vaccines for the treatment of CDI.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Atomic coordinates and structure factors of the TcdB–5D–E3–7F, TcdB1072–1433–5D, GTD–E3, and GTDVPI10463–7F complexes have been deposited in the Protein Data Bank under accession codes PDB 6OQ5, PDB 6OQ6, PDB 6OQ7, and PDB 6OQ8, respectively. Source data for Fig. 4e,f are available with the paper online. Other data are available upon request.
References
Rupnik, M., Wilcox, M. H. & Gerding, D. N. Š infection: new developments in epidemiology and pathogenesis. Nat. Rev. Microbiol. 7, 526–536 (2009).
Lessa, F. C. et al. Burden of Clostridium difficile infection in the United States. N. Engl. J. Med. 372, 825–834 (2015).
Jank, T. & Aktories, K. Structure and mode of action of clostridial glucosylating toxins: the ABCD model. Trends Microbiol. 16, 222–229 (2008).
Seal, D. et al. Treatment of relapsing Clostridium difficile diarrhoea by administration of a non-toxigenic strain. Eur. J. Clin. Microbiol. 6, 51–53 (1987).
Kuehne, S. A. et al. The role of toxin A and toxin B in Clostridium difficile infection. Nature 467, 711–713 (2010).
Lyras, D. et al. Toxin B is essential for virulence of Clostridium difficile. Nature 458, 1176–1179 (2009).
Carter, G. P. et al. Defining the roles of TcdA and TcdB in localized gastrointestinal disease, systemic organ damage, and the host response during Clostridium difficile infections. MBio 6, e00551 (2015).
Carter, G. P., Rood, J. I. & Lyras, D. The role of toxin A and toxin B in the virulence of Clostridium difficile. Trends Microbiol. 20, 21–29 (2012).
Wilcox, M. H. et al. Bezlotoxumab for prevention of recurrent Clostridium difficile infection. N. Engl. J. Med. 376, 305–317 (2017).
Ho, J. G., Greco, A., Rupnik, M. & Ng, K. K. Crystal structure of receptor-binding C-terminal repeats from Clostridium difficile toxin A. Proc. Natl Acad. Sci. USA 102, 18373–18378 (2005).
Aktories, K., Schwan, C. & Jank, T. Clostridium difficile toxin biology. Annu Rev. Microbiol. 71, 281–307 (2017).
Greco, A. et al. Carbohydrate recognition by Clostridium difficile toxin A. Nat. Struct. Mol. Biol. 13, 460–461 (2006).
Chen, P. et al. Structural basis for recognition of frizzled proteins by Clostridium difficile toxin B. Science 360, 664–669 (2018).
Yuan, P. et al. Chondroitin sulfate proteoglycan 4 functions as the cellular receptor for Clostridium difficile toxin B. Cell Res. 25, 157–168 (2015).
Genisyuerek, S. et al. Structural determinants for membrane insertion, pore formation and translocation of Clostridium difficile toxin B. Mol. Microbiol. 79, 1643–1654 (2011).
Zhang, Z. et al. Translocation domain mutations affecting cellular toxicity identify the Clostridium difficile toxin B pore. Proc. Natl Acad. Sci. USA 111, 3721–3726 (2014).
Qa’Dan, M., Spyres, L. M. & Ballard, J. D. pH-induced conformational changes in Clostridium difficile toxin B. Infect. Immun. 68, 2470–2474 (2000).
Li, S. et al. Critical roles of Clostridium difficile toxin B enzymatic activities in pathogenesis. Infect. Immun. 83, 502–513 (2015).
Sehr, P. et al. Glucosylation and ADP ribosylation of rho proteins: effects on nucleotide binding, GTPase activity, and effector coupling. Biochemistry 37, 5296–5304 (1998).
Egerer, M., Giesemann, T., Jank, T., Satchell, K. J. & Aktories, K. Auto-catalytic cleavage of Clostridium difficile toxins A and B depends on cysteine protease activity. J. Biol. Chem. 282, 25314–25321 (2007).
Reineke, J. et al. Autocatalytic cleavage of Clostridium difficile toxin B. Nature 446, 415–419 (2007).
Just, I. et al. Glucosylation of Rho proteins by Clostridium difficile toxin B. Nature 375, 500–503 (1995).
Hofmann, F., Busch, C., Prepens, U., Just, I. & Aktories, K. Localization of the glucosyltransferase activity of Clostridium difficile toxin B to the N-terminal part of the holotoxin. J. Biol. Chem. 272, 11074–11078 (1997).
Chumbler, N. M. et al. Crystal structure of Clostridium difficile toxin A. Nat. Microbiol. 1, 15002 (2016).
Murase, T. et al. Structural basis for antibody recognition in the receptor-binding domains of toxins A and B from Clostridium difficile. J. Biol. Chem. 289, 2331–2343 (2014).
Reinert, D. J., Jank, T., Aktories, K. & Schulz, G. E. Structural basis for the function of Clostridium difficile toxin B. J. Mol. Biol. 351, 973–981 (2005).
Shen, A. et al. Defining an allosteric circuit in the cysteine protease domain of Clostridium difficile toxins. Nat. Struct. Mol. Biol. 18, 364–371 (2011).
Orth, P. et al. Mechanism of action and epitopes of Clostridium difficile toxin B-neutralizing antibody bezlotoxumab revealed by X-ray crystallography. J. Biol. Chem. 289, 18008–18021 (2014).
Pruitt, R. N., Chambers, M. G., Ng, K. K., Ohi, M. D. & Lacy, D. B. Structural organization of the functional domains of Clostridium difficile toxins A and B. Proc. Natl Acad. Sci. USA 107, 13467–13472 (2010).
Yang, G. L. et al. Expression of recombinant Clostridium difficile toxin A and B in Bacillus megaterium. BMC Microbiol. 8, 192 (2008).
Yang, Z. et al. A novel multivalent, single-domain antibody targeting TcdA and TcdB prevents fulminant Clostridium difficile infection in mice. J. Infect. Dis. 210, 964–972 (2014).
Kobe, B. & Kajava, A. V. When protein folding is simplified to protein coiling: the continuum of solenoid protein structures. Trends Biochem. Sci. 25, 509–515 (2000).
Fernandez-Tornero, C., Lopez, R., Garcia, E., Gimenez-Gallego, G. & Romero, A. A novel solenoid fold in the cell wall anchoring domain of the pneumococcal virulence factor LytA. Nat. Struct. Biol. 8, 1020–1024 (2001).
Zhang, Y. et al. A segment of 97 amino acids within the translocation domain of Clostridium difficile toxin B is essential for toxicity. PLoS One 8, e58634 (2013).
Lanis, J. M., Barua, S. & Ballard, J. D. Variations in TcdB activity and the hypervirulence of emerging strains of Clostridium difficile. PLoS Pathog. 6, e1001061 (2010).
Lanis, J. M., Hightower, L. D., Shen, A. & Ballard, J. D. TcdB from hypervirulent Clostridium difficile exhibits increased efficiency of autoprocessing. Mol. Microbiol. 84, 66–76 (2012).
Larabee, J. L., Krumholz, A., Hunt, J. J., Lanis, J. M. & Ballard, J. D. Exposure of neutralizing epitopes in the carboxyl-terminal domain of TcdB is altered by a proximal hypervariable region. J. Biol. Chem. 290, 6975–6985 (2015).
Putnam, C. D., Hammel, M., Hura, G. L. & Tainer, J. A. X-ray solution scattering (SAXS) combined with crystallography and computation: defining accurate macromolecular structures, conformations and assemblies in solution. Q Rev. Biophys. 40, 191–285 (2007).
Kao, A. H. et al. Development of a novel cross-linking strategy for fast and accurate identification of cross-linked peptides of protein complexes. Mol. Cell. Proteomics 10, M110.002212 (2011).
Yu, C. & Huang, L. Cross-linking mass spectrometry: an emerging technology for interactomics and structural biology. Anal. Chem. 90, 144–165 (2018).
Hellenkamp, B. et al. Precision and accuracy of single-molecule FRET measurements-a multi-laboratory benchmark study. Nat. Methods 15, 669–676 (2018).
McCann, J. J., Choi, U. B., Zheng, L., Weninger, K. & Bowen, M. E. Optimizing methods to recover absolute FRET efficiency from immobilized single molecules. Biophys. J. 99, 961–970 (2010).
Gopich, I. V. & Szabo, A. FRET efficiency distributions of multistate single molecules. J. Phys. Chem. B 114, 15221–15226 (2010).
Chumbler, N. M. et al. Clostridium difficile Toxin B causes epithelial cell necrosis through an autoprocessing-independent mechanism. PLoS Pathog. 8, e1003072 (2012).
Lam, K. H. et al. A viral-fusion-peptide-like molecular switch drives membrane insertion of botulinum neurotoxin A1. Nat. Commun. 9, 5367 (2018).
Schmidt, D. J. et al. A tetraspecific VHH-based neutralizing antibody modifies disease outcome in three animal models of Clostridium difficile infection. Clin. Vaccin. Immunol. 23, 774–784 (2016).
Just, I. & Gerhard, R. Large clostridial cytotoxins. Rev. Physiol. Biochem. Pharm. 152, 23–47 (2004).
Egerer, M. & Satchell, K. J. Inositol hexakisphosphate-induced autoprocessing of large bacterial protein toxins. PLoS Pathog. 6, e1000942 (2010).
Lupardus, P. J., Shen, A., Bogyo, M. & Garcia, K. C. Small molecule-induced allosteric activation of the Vibrio cholerae RTX cysteine protease domain. Science 322, 265–268 (2008).
Puri, A. W. et al. Rational design of inhibitors and activity-based probes targeting Clostridium difficile virulence factor TcdB. Chem. Biol. 17, 1201–1211 (2010).
Olling, A. et al. The combined repetitive oligopeptides of Clostridium difficile toxin A counteract premature cleavage of the glucosyl-transferase domain by stabilizing protein conformation. Toxins (Basel) 6, 2162–2176 (2014).
Zhang, Y., Hamza, T., Gao, S. & Feng, H. Masking autoprocessing of Clostridium difficile toxin A by the C-terminus combined repetitive oligo peptides. Biochem. Biophys. Res Commun. 459, 259–263 (2015).
Geissler, B., Tungekar, R. & Satchell, K. J. Identification of a conserved membrane localization domain within numerous large bacterial protein toxins. Proc. Natl Acad. Sci. USA 107, 5581–5586 (2010).
Varela Chavez, C. et al. The tip of the four N-terminal alpha-helices of Clostridium sordellii lethal toxin contains the interaction site with membrane phosphatidylserine facilitating small GTPases glucosylation. Toxins (Basel) 8, 90 (2016).
Mesmin, B. et al. A phosphatidylserine-binding site in the cytosolic fragment of Clostridium sordellii lethal toxin facilitates glucosylation of membrane-bound Rac and is required for cytotoxicity. J. Biol. Chem. 279, 49876–49882 (2004).
Schorch, B. et al. LRP1 is a receptor for Clostridium perfringens TpeL toxin indicating a two-receptor model of clostridial glycosylating toxins. Proc. Natl Acad. Sci. USA 111, 6431–6436 (2014).
Kreimeyer, I. et al. Autoproteolytic cleavage mediates cytotoxicity of Clostridium difficile toxin A. Naunyn Schmiede. Arch. Pharm. 383, 253–262 (2011).
Gupta, P. et al. Functional defects in Clostridium difficile TcdB toxin uptake identify CSPG4 receptor-binding determinants. J. Biol. Chem. 292, 17290–17301 (2017).
Tao, L. et al. Frizzled proteins are colonic epithelial receptors for C. difficile toxin B. Nature 538, 350–355 (2016).
Laursen, N. S. et al. Universal protection against influenza infection by a multidomain antibody to influenza hemagglutinin. Science 362, 598–602 (2018).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Gu, S. et al. Botulinum neurotoxin is shielded by NTNHA in an interlocked complex. Science 335, 977–981 (2012).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
DiMaio, F. et al. Improved low-resolution crystallographic refinement with Phenix and Rosetta. Nat. Methods 10, 1102–1104 (2013).
Strong, M. et al. Toward the structural genomics of complexes: crystal structure of a PE/PPE protein complex from Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 103, 8060–8065 (2006).
Brunger, A. T. Free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–475 (1992).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Petoukhov, M. V., Konarev, P. V., Kikhney, A. G. & Svergun, D. I. ATSAS 2.1 - towards automated and web-supported small-angle scattering data analysis. J. Appl. Crystallogr. 40, S223–S228 (2007).
Svergun, D. I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 25, 495–503 (1992).
Svergun, D., Barberato, C. & Koch, M. H. J. CRYSOL - A program to evaluate x-ray solution scattering of biological macromolecules from atomic coordinates. J. Appl. Crystallogr. 28, 768–773 (1995).
Leitner, A., Walzthoeni, T. & Aebersold, R. Lysine-specific chemical cross-linking of protein complexes and identification of cross-linking sites using LC-MS/MS and the xQuest/xProphet software pipeline. Nat. Protoc. 9, 120–137 (2014).
Yu, C., Kandur, W., Kao, A., Rychnovsky, S. & Huang, L. Developing new isotope-coded mass spectrometry-cleavable cross-linkers for elucidating protein structures. Anal. Chem. 86, 2099–2106 (2014).
Yu, C. et al. Developing a multiplexed quantitative cross-linking mass spectrometry platform for comparative structural analysis of protein complexes. Anal. Chem. 88, 10301–10308 (2016).
Swoboda, M. et al. Enzymatic oxygen scavenging for photostability without pH drop in single-molecule experiments. ACS Nano. 6, 6364–6369 (2012).
Guttenberg, G. et al. Inositol hexakisphosphate-dependent processing of Clostridium sordellii lethal toxin and Clostridium novyi alpha-toxin. J. Biol. Chem. 286, 14779–14786 (2011).
Acknowledgements
This work was partly supported by National Institute of Health grants R01AI139087, R01AI125704, R21AI123920, R21AI139690, and R21CA235533 to R.J., R01GM074830 and R01GM130144 to L.H., R01DK084509, R01AI088748, R01AI132207, and U19AI109776 to H.F, and R01 MH081923 to M.E.B. NE-CAT at the Advanced Photon Source (APS) is supported by a grant from the National Institute of General Medical Sciences (P30GM124165). The Pilatus 6 M detector on 24-ID-C beam line is funded by an NIH-ORIP HEI grant (S10 RR029205). Use of the APS, an Office of Science User Facility operated for the U.S. Department of Energy (DOE) Office of Science by Argonne National Laboratory, was supported by the U.S. DOE under Contract No. DE-AC02-06CH11357. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, is supported by the U.S. DOE, Office of Science, Office of Basic Energy Sciences under Contract No. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research, and by the National Institutes of Health, National Institute of General Medical Sciences (including P41GM103393). The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of NIH.
Author information
Authors and Affiliations
Contributions
P.C. and R.J. conceived the project. P.C., K.L., Z.L., B.C., and R.J. carried out the protein expression, purification, characterization, crystallization, structure determination and analysis, and all related biochemical studies. K.P. collected the X-ray diffraction data. T.M. carried out the SAXS studies. P.C., C.B.G., and L.H. performed the XL-MS. F.A.M. and M.E.B. carried out the smFRET studies. Y.Z., T.H., and H.F. provided the plasmids of TcdB and VHHs (5D, E3, 7F) and TcdB-expressing Bacillus megaterium. P.C. and R.J. wrote the manuscript with input from other authors.
Corresponding author
Ethics declarations
Competing interests
A provisional patent application has been filed by The Regents of the University of California on the use of the structural information described in this manuscript to prevent and/or treat CDI. R.J. is a cofounder of DesignerT Inc., and H.F. is a co-founder of FZata Inc., which had no role in this study.
Additional information
Peer review information: Beth Moorefield was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
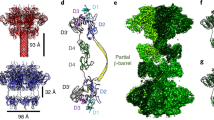
Supplementary Figure 1 Crystal structure of the TcdB–5D–E3–7F complex and the representative electron density maps.
(a) A 2Fo-Fc electron density map for the CROPs (residues G1835–E2367) contoured at 1.0 σ. (b–d) Representative 2Fo-Fc electron density maps for the newly identified SR (residues G1815–S1834) and two other regions in the CROPs (residues G1898–T1927 and G2345–E2367) contoured at 1.0 σ. The bulky residue Tyr was used as one of the markers to confirm the sequence register of the structural model. (e) An anomalous-difference electron density map generated from a crystal of the TcdB–5D–E3–7F complex soaked in tantalum bromide. The map was calculated using a 5.44 Å resolution data collected at λ = 1.2524 Å and phases derived from the final structure model. We found 10 tantalum bromide molecules in one asymmetric unit, which all bind to negatively charged glutamate or aspartate residues and thus confirmed the amino acid register of the complex. One representative tantalum bromide peak was shown in the inset. (f) Overall structure of the TcdB–5D–E3–7F complex. The color scheme for the TcdB domains are the same as that shown in Fig. 1. The three VHHs are colored purple (5D), brown (E3), and green (7F). (g) The structure of TcdB holotoxin is fitted into the negative stain EM map of TcdA that is displayed as a grey surface model (map was generously provided by Dr. Lacy D.B.) (Chumbler, N.M. et al., Nat Microbiol. 1, 15002, 2016). The CROPs of TcdA interacts with the DRBD and adopts a closed conformation.
Supplementary Figure 2 Sequence alignment in the CROPs and the hinge region.
(a) Sequence alignment of all the SRs and LRs in the CROPs of TcdB* (M68 strain), with the sequence conservation displayed on the top of the alignment. (b) Sequence alignment of the hinge region among TcdB*, TcdB (VPI 10463 strain), and TcdB2 (BI/NAP1/027 strain). (c)The hinge region (olive) directly interacts with the 3-HB (blue) and the β-flap (pink). Residues involved in the interactions are colored cyan for the hinge and green for the 3-HB and the β-flap.
Supplementary Figure 3 SEC-SAXS studies on TcdB holotoxin at pH 7.4 and 5.0.
(a, b) Rg (blue) and I(0) (orange) plots for SEC-SAXS at pH 7.4 and 5.0. (c) SAXS profile at pH 7.4 (blue) and pH 5.0 (orange) were superimposed with a curve fitting using the crystal structure of TcdB holotoxin (black). (d) Pairwise distribution function, P(r), of TcdB holotoxin at pH 7.4 (blue) and 5.0 (orange). (e) The maximum dimension of TcdB holotoxin at acidic pH and the core of TcdB composed of GTD, CPD, and DRBD.
Supplementary Figure 4 XL-MS and FRET studies of TcdB holotoxin.
(a) Crosslinking workflow used to unambiguously identify DSSO crosslinks from 3 replicates (Rep1–3) of cross-linked TcdB holotoxin. (b) MSn identification of a DSSO inter-linked peptide from TcdB: K1117–K2249. The cross-linked peptide α–β4+ (m/z 510.9914+) was detected during MS1. Next, it was selected for MS2 where it was fragmented into two characteristic fragment ion pairs, that is αA/βT (m/z 308.672+/704.312+) and αT/βA (m/z 324.652+/688.322+). Finally, these peaks were selected for MS3 analysis. αA (m/z 308.672+) identified the sequence DKAATK, in which the lysine residue at position 2 was modified with an alkene moiety. βT (m/z 704.312+) identified the sequence as YYFDEKTGIMR, in which the lysine residue at position 6 was modified with unsaturated thiol moiety. (c) Dynamic light scattering showed a monodispersed peak for TcdB holotoxin at neutral pH. (d) The distribution plot of identified linkages versus their spatial distances measured in the structure of TcdB holotoxin. (e) 5D has no effect on the fluorescence emission of the dye-labeled VHHs and the TcdB–B39–7F complex based on an ensemble FRET study. TcdB holotoxin was incubated with equimolar ratio of the Alexa-555-labeled B39, the Alexa-647-labeled 7F, or both VHHs, with or without 5D at pH 7 or pH 5. Fluorescence emissions upon excitation by 540 nm were measured. Data are the average of duplicated experiments. (f) Fluorescence intensity over time for single molecules of TcdB in complex with dye-labeled VHHs. Alexa 555, donor-labeled B39 (magenta). Alexa-647, acceptor-labeled F7 (cyan). Data was recorded at a frame rate of 10 Hz. Only molecules showing single step photobleaching to baseline for both dyes were included in the analysis. From the magnitude of the anticorrelated bleaching event, we obtained the γ-correction factor for each molecule that allows us to report the absolute FRET efficiencies. FRET was calculated for each frame until acceptor photobleaching. (g) Histogram of average FRET efficiency for each molecule at either pH 5.0 (filled circles, n = 498) or pH 7.0 (open circles, n = 594). This reports a single value for each molecule, which averages out any conformational dynamics. The mean FRET efficiency was unchanged by changing the time binning of the data. (h) Mean smFRET efficiency for TcdB in complex with dye-labeled VHHs at intermediate pH. FRET was calculated from each 100 ms frame of the movie until acceptor photobleaching, compiled into histograms and fit to a Gaussian function to obtain the FRET efficiencies. Error values represent the standard deviation from replicate measurements.
Supplementary Figure 5 VHHs 5D, 7F, and E3 recognize distinct epitopes and exploit different mechanisms to neutralize TcdB.
(a, b) pH-dependent conformational change of TcdB1072–1433 probed by ANS. TcdB1072–1433, the TcdB1072–1433–5D complex, or 5D alone was incubated with ANS in buffers with different pH, and the peak fluorescence intensities of ANS at 474 nm are shown. Error bar represents SD of three replicate experiments. (c) The structure of the TcdB1072–1433–5D complex (light blue, residues 1090–1431 of TcdB were fully resolved in this crystal structure) was superimposed to the full length TcdB–5D complex (TcdB and 5D were colored yellow and pink, respectively). The overall Cα atom r.m.s.d. is ~0.608 Å. (d) Structural superposition to compare the conformations of the pore-forming region among the TcdB1072–1433–5D complex that was crystallized at pH 8.5 (light blue), the TcdB–5D complex that was crystallized at acidic pH (yellow), and TcdA at neutral pH (orange). (e, f) A close-up view into the interface between TcdB and 5D in the structure of the TcdB1072–1433–5D complex, where the CDR2 of 5D (e), as well as its CDR1 and CDR3 (f) bind TcdB with extensive interactions. (g) 7F fixes the conformation of the Helix 1 (residues 525–539) and the Helix 2 (residues 137–158) in the GTD, which are in close proximity to the active site of the CPD and the scissile bond (L544–G545). GTD, CPD, and 7F are colored red, light blue, and green, respectively. The peptide linker between the GTD and the CPD is colored blue. A close-up view of the boxed area is shown in (h). (i) Detailed interactions between GTDVPI10463 and 7F. The interacting residues are colored green and pink for 7F and GTD, respectively. The 7F-binding interfaces are largely identical between GTDVPI10463 and GTD. (j) Interactions between GTD and E3. The residues involved in interactions are colored wheat and pink for E3 and GTD, respectively.
Supplementary Figure 6 Autoprocessing is self-inhibited in TcdB holotoxin.
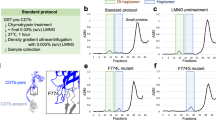
(a) A close-up view of a zinc atom bound in the active site of the CPD in TcdB holotoxin. (b, c) A simplified cartoon model showing the co-localization of the β-flap, the 3-HB, and the hinge. In TcdB holotoxin, the β-flap is kept in an inactive conformation (pink). Cytosolic InsP6 triggers a ~90° rotation of the β-flap (green) to activate CPD. (d) TcdB1–1805 showed a higher autoprocessing efficiency than the full length TcdB. The autoprocessing of TcdB was examined at 37°C for 1 hour incubation in the presence of various concentrations of InsP6. The results were assessed by SDS-PAGE. 7F potently inhibited GTD cleavage. The molecular weight of TcdB holotoxin, TcdB1–1805, GTD, and 7F is 270 kDa, 205 kDa, 64 kDa, and 15.2 kDa, respectively. The gel is representative of three independent experiments.
Supplementary Figure 7 Mapping the binding sites for CSPG4, FZDs, and bezlotoxumab in TcdB holotoxin.
(a) The reported CSPG4-binding site in TcdB, involving residues 1500–1900 (Gupta, P. et al., J Biol Chem. 292, 17290-17301, 2017 and Yuan, P. et al., Cell Res. 25, 157-68, 2015), was colored blue. (b) Binding of TcdB to the immobilized biotin-labeled CRD2 was examined using a pull-down assay at neutral and acidic pH. Samples were analyzed by SDS-PAGE and Coomassie Blue staining. (c) The structure of the cysteine-rich domain of FZD2 (CRD2, green cartoon model) in complex with TcdB1285–1804 (PDB code: 6C0B) (Chen, P. et al., Science, 360, 664-669, 2018) was superimposed to TcdB holotoxin. The pore-forming region of TcdB is shown in a purple ribbon model while the rest of the toxin is shown in a surface model. (d) The structure of the CROPs I–II of TcdB in complex with bezlotoxumab Fabs (PDB code: 4NP4) (Orth, P. et al., J Biol Chem, 289, 18008-21, 2014) was superimposed to TcdB holotoxin. Bezlotoxumab Fabs were shown as cartoon models, which bind to two sites on the CROPs (blue and pink). Bezlotoxumab clashes with the GTD and the DRBD when TcdB adopts the acidic conformation. A close-up view of the boxed area is shown in (e).
Supplementary information
Supplementary Information
Supplementary Figs. 1–7, Supplementary Tables 1–3, and Supplementary Note 1
Source data
Rights and permissions
About this article
Cite this article
Chen, P., Lam, Kh., Liu, Z. et al. Structure of the full-length Clostridium difficile toxin B. Nat Struct Mol Biol 26, 712–719 (2019). https://doi.org/10.1038/s41594-019-0268-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0268-0
This article is cited by
-
Molecular basis of TMPRSS2 recognition by Paeniclostridium sordellii hemorrhagic toxin
Nature Communications (2024)
-
Structural dynamics of the CROPs domain control stability and toxicity of Paeniclostridium sordellii lethal toxin
Nature Communications (2023)
-
Unconventional structure and mechanisms for membrane interaction and translocation of the NF-κB-targeting toxin AIP56
Nature Communications (2023)
-
Design of 8-mer peptides that block Clostridioides difficile toxin A in intestinal cells
Communications Biology (2023)
-
From signal transduction to protein toxins—a narrative review about milestones on the research route of C. difficile toxins
Naunyn-Schmiedeberg's Archives of Pharmacology (2023)