Abstract
Glucagon and insulin maintain blood glucose homeostasis and are used to treat hypoglycemia and hyperglycemia, respectively, in patients with diabetes. Whereas insulin is stable for weeks in its solution formulation, glucagon fibrillizes rapidly at the acidic pH required for solubility and is therefore formulated as a lyophilized powder that is reconstituted in an acidic solution immediately before use. Here we use solid-state NMR to determine the atomic-resolution structure of fibrils of synthetic human glucagon grown at pharmaceutically relevant low pH. Unexpectedly, two sets of chemical shifts are observed, indicating the coexistence of two β-strand conformations. The two conformations have distinct water accessibilities and intermolecular contacts, indicating that they alternate and hydrogen bond in an antiparallel fashion along the fibril axis. Two antiparallel β-sheets assemble with symmetric homodimer cross sections. This amyloid structure is stabilized by numerous aromatic, cation-π, polar and hydrophobic interactions, suggesting mutagenesis approaches to inhibit fibrillization could improve this important drug.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
NMR chemical shifts and distance restraints have been deposited in the Biological Magnetic Resonance Bank (BMRB) under entry 30572. Structural coordinates have been deposited online in the Protein Data Bank under accession code PDB 6NZN. The 2D spectra are available in Supplementary Data Set 1. All distance restraints derived from 13C–13C and 15N–13C correlation spectra are available in Supplementary Data Set 2. All other data are available from the authors upon reasonable request.
References
Chiti, F. & Dobson, C. M. Protein misfolding, amyloid formation, and human disease: a summary of progress over the last decade. Annu. Rev. Biochem. 86, 27–68 (2017).
Frokjaer, S. & Otzen, D. E. Protein drug stability: a formulation challenge. Nat. Rev. Drug Discov. 4, 298 (2005).
Brange, J., Andersen, L., Laursen, E. D., Meyn, G. & Rasmussen, E. Toward understanding insulin fibrillation. J. Pharm. Sci. 86, 517–525 (1997).
Habegger, K. M. et al. The metabolic actions of glucagon revisited. Nat. Rev. Endocrinol. 6, 689 (2010).
Caputo, N. et al. Mechanisms of glucagon degradation at alkaline pH. Peptides 45, 40–47 (2013).
Beaven, G. H., Gratzer, W. B. & Davies, H. G. Formation and structure of gels and fibrils from glucagon. Eur. J. Biochem. 11, 37–42 (1969).
Andersen, C. B. et al. Glucagon fibril polymorphism reflects differences in protofilament backbone structure. J. Mol. Biol. 397, 932–946 (2010).
Oliveira, C. L. et al. A SAXS study of glucagon fibrillation. J. Mol. Biol. 387, 147–161 (2009).
Dong, M. et al. AFM-based force spectroscopy measurements of mature amyloid fibrils of the peptide glucagon. Nanotechnology 19, 384013 (2008).
Pedersen, J. S. et al. The changing face of glucagon fibrillation: structural polymorphism and conformational imprinting. J. Mol. Biol. 355, 501–523 (2006).
De Jong, K. L., Incledon, B., Yip, C. M. & DeFelippis, M. R. Amyloid fibrils of glucagon characterized by high-resolution atomic force microscopy. Biophys. J. 91, 1905–1914 (2006).
Gratzer, W. B., Bailey, E. & Beaven, G. H. Conformational states of glucagon. Biochem. Biophys. Res. Commun. 28, 914–919 (1967).
Colvin, M. T. et al. Atomic resolution structure of monomorphic Aβ42 amyloid fibrils. J. Am. Chem. Soc. 138, 9663–9674 (2016).
Wälti, M. A. et al. Atomic-resolution structure of a disease-relevant Aβ(1–42) amyloid fibril. Proc. Natl Acad. Sci. USA 113, E4976–E4984 (2016).
Tuttle, M. D. et al. Solid-state NMR structure of a pathogenic fibril of full-length human α-synuclein. Nat. Struct. Mol. Biol. 23, 409 (2016).
Fitzpatrick, A. W. P. et al. Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 547, 185 (2017).
Sawaya, M. R. et al. Atomic structures of amyloid cross-beta spines reveal varied steric zippers. Nature 447, 453–457 (2007).
Antzutkin, O. N. et al. Multiple quantum solid-state NMR indicates a parallel, not antiparallel, organization of β-sheets in Alzheimer’s β-amyloid fibrils. Proc. Natl Acad. Sci. USA 97, 13045–13050 (2000).
Paravastu, A. K., Leapman, R. D., Yau, W.-M. & Tycko, R. Molecular structural basis for polymorphism in Alzheimer’s β-amyloid fibrils. Proc. Natl Acad. Sci. USA 105, 18349–18354 (2008).
Petkova, A. T., Yau, W.-M. & Tycko, R. Experimental constraints on quaternary structure in alzheimer’s β-amyloid fibrils. Biochemistry 45, 498–512 (2006).
Tycko, R. Amyloid polymorphism: structural basis and neurobiological relevance. Neuron 86, 632–645 (2015).
Hiller, S. et al. Solution structure of the integral human membrane protein VDAC-1 in detergent micelles. Science 321, 1206–1210 (2008).
Paëpe, G. D., Lewandowski, J. R., Loquet, A., Böckmann, A. & Griffin, R. G. Proton assisted recoupling and protein structure determination. J. Chem. Phys. 129, 245101 (2008).
Gelenter, M. D. & Hong, M. Efficient 15N–13C polarization transfer by third-spin-assisted pulsed cross-polarization magic-angle-spinning NMR for protein structure determination. J. Phys. Chem. B 122, 8367–8379 (2018).
Wasmer, C. et al. Amyloid fibrils of the HET-s(218-289) prion form a beta solenoid with a triangular hydrophobic core. Science 319, 1523–1526 (2008).
Qiang, W., Yau, W.-M., Luo, Y., Mattson, M. P. & Tycko, R. Antiparallel β-sheet architecture in Iowa-mutant β-amyloid fibrils. Proc. Natl Acad. Sci. USA 109, 4443–4448 (2012).
Lange, A., Luca, S. & Baldus, M. Structural constraints from proton-mediated rare-spin correlation spectroscopy in rotating solids. J. Am. Chem. Soc. 124, 9704–9705 (2002).
Wang, T., Jo, H., DeGrado, W. F. & Hong, M. Water distribution, dynamics, and interactions with Alzheimer’s β-amyloid fibrils investigated by solid-state NMR. J. Am. Chem. Soc. 139, 6242–6252 (2017).
Mandala, V. S., Gelenter, M. D. & Hong, M. Transport-relevant protein conformational dynamics and water dynamics on multiple time scales in an archetypal proton channel: Insights from solid-state NMR. J. Am. Chem. Soc. 140, 1514–1524 (2018).
Lesage, A. & Bockmann, A. Water-protein interactions in microcrystalline Crh measured by H-1-C-13 solid-state NMR spectroscopy. J. Am. Chem. Soc. 125, 13336–13337 (2003).
Liepinsh, E. & Otting, G. Proton exchange rates from amino acid side chains— implications for image contrast. Magn. Reson. Med. 35, 30–42 (1996).
Williams, J. K. & Hong, M. Probing membrane protein structure using water polarization transfer solid-state NMR. J. Magn. Reson. 247, 118–127 (2014).
Gallivan, J. P. & Dougherty, D. A. Cation-pi interactions in structural biology. Proc. Natl Acad. Sci. USA 96, 9459–9464 (1999).
Serrano, L., Bycroft, M. & Fersht, A. R. Aromatic-aromatic interactions and protein stability. Investigation by double-mutant cycles. J. Mol. Biol. 218, 465–475 (1991).
Meyer, E. A., Castellano, R. K. & Diederich, F. Interactions with aromatic rings in chemical and biological recognition. Angew. Chem. Int. Ed. Engl. 42, 1210–1250 (2003).
Choma, C., Gratkowski, H., Lear, J. D. & DeGrado, W. F. Asparagine-mediated self-association of a model transmembrane helix. Nat. Struct. Biol. 7, 161–166 (2000).
Lear, J. D., Gratkowski, H., Adamian, L., Liang, J. & DeGrado, W. F. Position-dependence of stabilizing polar interactions of asparagine in transmembrane helical bundles. Biochemistry 42, 6400–6407 (2003).
Deechongkit, S. et al. Context-dependent contributions of backbone hydrogen bonding to beta-sheet folding energetics. Nature 430, 101–105 (2004).
Elkins, M. R. et al. Structural polymorphism of alzheimer’s β-Amyloid fibrils as controlled by an E22 switch: a solid-state NMR study. J. Am. Chem. Soc. 138, 9840–9852 (2016).
Helmus, J. J., Surewicz, K., Nadaud, P. S., Surewicz, W. K. & Jaroniec, C. P. Molecular conformation and dynamics of the Y145Stop variant of human prion protein in amyloid fibrils. Proc. Natl Acad. Sci. USA 105, 6284–6289 (2008).
Mompeán, M. et al. The structure of the necrosome RIPK1-RIPK3 Core, a human hetero-amyloid signaling complex. Cell 173, 1244–1253 (2018).
Pedersen, J. S., Dikov, D. & Otzen, D. E. N- and C-terminal hydrophobic patches are involved in fibrillation of glucagon. Biochemistry 45, 14503–14512 (2006).
Boesch, C., Bundi, A., Oppliger, M. & Wüthrich, K. 1H nuclear-magnetic-resonance studies of the molecular conformation of monomeric glucagon in aqueous solution. Eur. J. Biochem. 91, 209–214 (1978).
Braun, W., Wider, G., Lee, K. H. & Wüthrich, K. Conformation of glucagon in a lipid-water interphase by 1H nuclear magnetic resonance. J. Mol. Biol. 169, 921–948 (1983).
Zhang, H. et al. Structure of the glucagon receptor in complex with a glucagon analogue. Nature 553, 106–110 (2018).
Sasaki, K., Dockerill, S., Adamiak, D. A., Tickle, I. J. & Blundell, T. X-ray analysis of glucagon and its relationship to receptor binding. Nature 257, 751–757 (1975).
Svane, A. S. et al. Early stages of amyloid fibril formation studied by liquid-state NMR: the peptide hormone glucagon. Biophys. J. 95, 366–377 (2008).
Moorthy, B. S., Ghomi, H. T., Lill, M. A. & Topp, E. M. Structural transitions and interactions in the early stages of human glucagon amyloid fibrillation. Biophys. J. 108, 937–948 (2015).
Andersen, C. B., Otzen, D., Christiansen, G. & Rischel, C. Glucagon amyloid-like fibril morphology is selected via morphology-dependent growth inhibition. Biochemistry 46, 7314–7324 (2007).
Chabenne, J. et al. A glucagon analog chemically stabilized for immediate treatment of life-threatening hypoglycemia. Mol. Metab. 3, 293–300 (2014).
Tian, Y. et al. Nanotubes, plates, and needles: pathway-dependent self-assembly of computationally designed peptides. Biomacromolecules 19, 4286–4298 (2018).
Amos, L. A. & Klug, A. Arrangement of subunits in flagellar microtubules. J. Cell Sci. 14, 523–549 (1974).
Bernard, G. M. et al. Methylammonium lead chloride: A sensitive sample for an accurate NMR thermometer. J. Magn. Reson. 283, 14–21 (2017).
Lee, W., Tonelli, M. & Markley, J. L. NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31, 1325–1327 (2015).
Takegoshi, K., Nakamura, S. & Terao, T. 13C–1H dipolar-assisted rotational resonance in magic-angle spinning NMR. Chem. Phys. Lett. 344, 631–637 (2001).
Hou, G., Yan, S., Trébosc, J., Amoureux, J.-P. & Polenova, T. Broadband homonuclear correlation spectroscopy driven by combined R2nv sequences under fast magic angle spinning for NMR structural analysis of organic and biological solids. J. Magn. Reson. 232, 18–30 (2013).
Hing, A. W., Vega, S. & Schaefer, J. Transferred-echo double-resonance NMR. J. Magn. Reson. 96, 205–209 (1992).
Daviso, E., Eddy, M. T., Andreas, L. B., Griffin, R. G. & Herzfeld, J. Efficient resonance assignment of proteins in MAS NMR by simultaneous intra- and inter-residue 3D correlation spectroscopy. J. Biomol. NMR 55, 257–265 (2013).
Baldus, M., Petkova, A. T., Herzfeld, J. & Griffin, R. G. Cross polarization in the tilted frame: assignment and spectral simplification in heteronuclear spin systems. Mol. Phys. 95, 1197–1207 (1998).
Shen, Y. & Bax, A. Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J. Biomol. NMR 56, 227–241 (2013).
Güntert, P., Mumenthaler, C. & Wüthrich, K. Torsion angle dynamics for NMR structure calculation with the new program DYANA. J. Mol. Biol. 273, 283–298 (1997).
Jeppesen, M. D., Hein, K., Nissen, P., Westh, P. & Otzen, D. E. A thermodynamic analysis of fibrillar polymorphism. Biophys. Chem. 149, 40–46 (2010).
Acknowledgements
This work was funded by Merck Sharp & Dohme Corp., a subsidiary of Merck & Co., Inc, and by NIH grant no. AG059661 to M.H. M.D.G. was partially supported by an NIH Ruth L. Kirschstein Individual National Research Service Award (no. 1F31AI133989).
Author information
Authors and Affiliations
Contributions
M.H. and Y.S. designed and coordinated the project. M.D.G., S.Y.L., V.S.M. and M.H. conducted the solid-state NMR experiments and analyzed the spectra. M.D.G. and A.J.D. conducted structure calculations. K.J.S., W.X., T.J.T. and Y.S. prepared the fibril samples. K.J.S. conducted CD, DLS and negative-stain TEM experiments. M.S.L. measured AFM data. Y.T., M.S.L., Y.S. and D.J.P. measured and analyzed MPL data. M.D.G. and M.H. wrote the manuscript with input and contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
K.J.S, M.S.L, W.X., T.J.T. and Y.S. are Merck & Co., Inc. employees.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 13C and 15N CP-MAS NMR spectra of low-pH glucagon fibrils.
The spectra show narrow linewidths and peak doubling, indicating the coexistence of two well-ordered molecular conformations. a, 13C spectra. b, 15N spectra. Peak assignments obtained from 2D and 3D experiments are shown for samples 1–4. All spectra were collected on an 800 MHz spectrometer under 10–14 kHz MAS at a setpoint temperature of 273–293 K. Peaks assigned to fewer than three residues are labeled in samples 7c–9c.
Supplementary Figure 2 2D 13C-13C correlation spectra of glucagon fibrils.
a, Sample 1 spectrum. b, Sample 2 spectrum. c, Sample 3 spectrum. d, Sample 4 spectrum. e, Sample 7c spectrum. f, Sample 8c spectrum. g, Sample 9c spectrum. h, Spectrum of mixed samples 11 and 12. The spectra were measured using 50 ms or 30 ms 13C spin diffusion at 293 or 273 K on an 800 MHz spectrometer under 10–14 kHz MAS.
Supplementary Figure 3 2D 15N-13Cα correlation spectra of glucagon fibrils.
a, Sample 1. b, Sample 2. c, Sample 3. d, Sample 4. e, Sample 7c. f, Sample 8c. Note the presence of peak doubling for most residues. The spectra were measured using an out-and-back ZF- TEDOR sequence with a mixing time of 1.4–2.0 ms. All spectra were collected on an 800 MHz spectrometer under 10 kHz MAS at a setpoint temperature of 293 K.
Supplementary Figure 4 2D 15N-13C correlation spectra of glucagon fibrils for chemical shift measurement and extraction of long-range contacts.
a, Sample 7c. b, Sample 8c. c, Sample 9c. The spectra were measured using a 15N-13C TEDOR mixing time of 1.2 ms, followed by 79 ms of 13C-13C CORD mixing on an 800 MHz spectrometer under 10 kHz MAS at a setpoint temperature of 283 K. Sequential cross peaks (assigned in green) facilitate the assignment of the two β-strand conformations.
Supplementary Figure 5 Glucagon secondary chemical shifts and chemical shift differences between conformers I and II.
a, Cα (orange) and Cβ (red) secondary shifts of conformer I. b, CO secondary shifts of conformer I. c, Cα (light blue) and Cβ (blue) secondary shifts of conformer II. d, CO secondary shifts of conformer II. e, Difference in the δCα-δCβ chemical shift difference between conformers I and II. No systematic trend exists in the sign of the chemical shift differences, indicating that the two conformers are equivalently β-sheet.
Supplementary Figure 6 2D 13C-13C correlation spectra indicate intermolecular packing of the glucagon β-strands along the fibril axis.
a, PDSD spectrum of mixed sample 11 and 12 with 1.0 s 13C spin diffusion. Intermolecular T5–L26, T5–M27, S11–A19, and S11–Q20 cross peaks are observed (assigned in purple), indicating antiparallel packing of the β-strands. b, 500 ms 2D 13C SD spectrum of sample 9c. In addition to sequential correlations, R18-K12, A19-K12, S11-A19, S11-R18 and R17-L14 cross peaks between conformers I and II are observed, supporting antiparallel β-strand packing. c, 500 ms sample 2 spectrum shows multiple H1-T29 correlations, indicating that the antiparallel packing spans the entire peptide sequence. d, 500 ms SD spectrum of sample 7c shows multiple sequential and long-range cross peaks of W25, which constrain the indole conformation. e, 200 μs 1H SD CHHC spectrum of sample 7c. f, 200 μs 1H SD CHHC spectrum of sample 9c. Cα–Cα cross peaks are observed for residue numbers that add up to 31 (for example Q3–N28, M27–G4, T5–L26, K12–A19, and Y13–R18), indicating that the β-strands are hydrogen-bonded in antiparallel. The cross peaks occur between conformers I and II, indicating that the two conformers alternate along the fibril axis.
Supplementary Figure 7 2D 13C-13C correlation spectra of diluted samples indicate the intermolecular nature of all long-range correlations and the homodimer interfaces in the plane perpendicular to the fibril axis.
a, 500 ms PDSD spectra of sample 1 without (black) and with (red) dilution. b, 500 ms CORD spectra of sample 3 without (black) and with (red) dilution. The 1D cross sections of V23 Cα and T5 Cβ show reduced intensities for V23 and T5 cross peaks compared to the diagonal peaks due to dilution, indicating that in the undiluted sample, these cross peaks result from both intramolecular and intermolecular contacts. Sequential cross peaks such as T5–F6 have similar intensity reduction as the intra-residue peaks, indicating that these sequential contacts also have both intramolecular and intermolecular contributions. For comparison, long-range V23–S8 and M27–T5 correlations are suppressed by dilution, indicating that these correlations result from antiparallel intermolecular packing.
Supplementary Figure 8 Water-to-peptide polarization transfer data.
a, Pulse sequence for transferring water 1H polarization to the protein and detected through 2D 13C-13C correlation spectra (Wang, T., et al., J. Am. Chem. Soc. 139, 6242-6252, 2017). Control spectra have a 1H T2 filter of 0 ms, whereas water-edited spectra have a T2 filter of 1.4–1.7 ms. All spectra were collected with a 2.25 ms 1H-1H tmix period. b-d, Aliphatic regions of the water-edited 2D 13C-13C correlation spectra of b, sample 7c, c, sample 8c, and d, sample 9c. Representative 1D cross sections of the sample 7c 2D spectrum illustrate the differential water-transferred intensities for the same residue in different conformers. For example, T5 in conformer I has lower water-transferred intensities than T5 in conformer II, indicating that T5 lies in a dry interface in conformer I but is water-exposed in conformer II.
Supplementary Figure 9 Intermolecular assembly of glucagon β-strands differs from two other canonical amyloid fibrils.
a, Schematic of the glucagon β-strand packing. The fibrils contain two distinct conformers that hydrogen-bond in antiparallel. The two strands in each cross section have C2 symmetry around the axis perpendicular to the fibril axis and parallel to the peptide backbone. b, Schematic of the two-fold symmetric Aβ1-40 fibrils, which consist of strand-turn-strand monomers that hydrogen-bond in a parallel-in-register fashion and that are C2 symmetric around the fibril axis (Petkova, A.T., et al., Biochemistry. 45, 498-512, 2006). c, Schematic of the D23N mutant of Aβ1-40 fibrils (Qiang. W. et al., Proc. Natl. Acad. Sci. USA. 109. 4443-4448, 2012). The β-strands hydrogen-bond in an antiparallel fashion along the fibril axis under certain conditions, but with time convert into a thermodynamically stable parallel-in-register structure. d, Observed intramolecular inter-residue contacts for each glucagon conformer, viewed from the homodimer cross sections. Sequential contacts are shown in gray and medium-range contacts in orange. e, Observed intermolecular contacts (green) between conformer I and conformer II of glucagon.
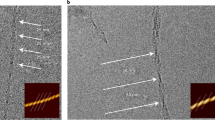
Supplementary Figure 10 STEM images used to determine the MPL of low-pH glucagon fibrils.
a, Stained and b, unstained images of microtubulin. c, Unstained image of glucagon fibrils diluted by 160-fold to isolate individual fibrils. d-f, Representative STEM images used to determine the glucagon MPL. g, Glucagon MPL histogram, showing a broad distribution with a maximum at 40.1 ± 0.5 kDa/nm. The glucagon molecular weight is 3.48 kDa, thus a single cross-β sheet with a 4.8 Å inter-chain spacing has an MPL of 7.25 kDa/nm. Therefore, the maximum MPL value corresponds to 5.5 ± 0.1 cross-β sheets as the predominant fibril species, indicating that three dimers stack in the plane normal to the fibril axis.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Supplementary Tables 1–3.
Supplementary Data Set 1
All 2D NMR spectra showing long-range correlations.
Supplementary Data Set 2
Excel table of measured inter-residue contacts.
Rights and permissions
About this article
Cite this article
Gelenter, M.D., Smith, K.J., Liao, SY. et al. The peptide hormone glucagon forms amyloid fibrils with two coexisting β-strand conformations. Nat Struct Mol Biol 26, 592–598 (2019). https://doi.org/10.1038/s41594-019-0238-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0238-6
This article is cited by
-
Polymorphic amyloid nanostructures of hormone peptides involved in glucose homeostasis display reversible amyloid formation
Nature Communications (2023)
-
Cell-free synthesis of amyloid fibrils with infectious properties and amenable to sub-milligram magic-angle spinning NMR analysis
Communications Biology (2022)
-
The Cryo-EM structures of two amphibian antimicrobial cross-β amyloid fibrils
Nature Communications (2022)
-
Solid-state NMR spectroscopy
Nature Reviews Methods Primers (2021)
-
Comparative analysis of 13C chemical shifts of β-sheet amyloid proteins and outer membrane proteins
Journal of Biomolecular NMR (2021)