Abstract
All known eukaryotic topoisomerases are only able to relieve torsional stress in DNA. Nevertheless, it has been proposed that the introduction of positive DNA supercoiling is required for efficient sister-chromatid disjunction by Topoisomerase 2a during mitosis. Here we identify a eukaryotic enzymatic activity that introduces torsional stress into DNA. We show that the human Plk1-interacting checkpoint helicase (PICH) and Topoisomerase 3a proteins combine to create an extraordinarily high density of positive DNA supercoiling. This activity, which is analogous to that of a reverse-gyrase, is apparently driven by the ability of PICH to progressively extrude hypernegatively supercoiled DNA loops that are relaxed by Topoisomerase 3a. We propose that this positive supercoiling provides an optimal substrate for the rapid disjunction of sister centromeres by Topoisomerase 2a at the onset of anaphase in eukaryotic cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Code availability
The custom-made program used to operate the tweezers and analyze the data can be obtained upon request to V. Croquette (ENS Paris).
References
Kschonsak, M. & Haering, C. H. Shaping mitotic chromosomes: from classical concepts to molecular mechanisms. BioEssays 37, 755–766 (2015).
Baumann, C., Körner, R., Hofmann, K. & Nigg, E. A. PICH, a centromere-associated SNF2 family ATPase, is regulated by Plk1 and required for the spindle checkpoint. Cell 128, 101–114 (2007).
Chan, K.-L., North, P. S. & Hickson, I. D. BLM is required for faithful chromosome segregation and its localization defines a class of ultrafine anaphase bridges. EMBO J. 26, 3397–3409 (2007).
Wang, L. H. C., Schwarzbraun, T., Speicher, M. R. & Nigg, E. A. Persistence of DNA threads in human anaphase cells suggests late completion of sister chromatid decatenation. Chromosoma 117, 123–135 (2008).
Bizard, A. H. & Hickson, I. D. Anaphase: a fortune-teller of genomic instability. Curr. Opin. Cell Biol. 52, 112–119 (2018).
Nielsen, C. F. et al. PICH promotes sister chromatid disjunction and co-operates with topoisomerase II in mitosis. Nat. Commun. 6, 8962 (2015).
Spence, J. M. et al. Depletion of topoisomerase II leads to shortening of the metaphase interkinetochore distance and abnormal persistence of PICH-coated anaphase threads. J. Cell Sci. 120, 3952–3964 (2007).
Clarke, D. J. & Azuma, Y. Non-catalytic roles of the topoisomerase IIα C-terminal domain. Int. J. Molec. Sci. 18, pii: E2438 (2017).
Guturi, K. K. N. et al. RNF168 and USP10 regulate topoisomerase IIα function via opposing effects on its ubiquitylation. Nat. Commun. 7, 12638 (2016).
Dykhuizen, E. C. et al. BAF complexes facilitate decatenation of DNA by topoisomerase IIα. Nature 497, 624–627 (2013).
Baxter, J. et al. Positive supercoiling of mitotic DNA drives decatenation by topoisomerase II in eukaryotes. Science 331, 1328–1332 (2011).
Vologodskii, A. Unlinking of supercoiled DNA catenanes by type IIA topoisomerases. Biophys. J. 101, 1403–1411 (2011).
Biebricher, A. et al. PICH: a DNA translocase specially adapted for processing anaphase bridge DNA. Mol. Cell 51, 691–701 (2013).
Liu, Y., Nielsen, C. F., Yao, Q. & Hickson, I. D. The origins and processing of ultra fine anaphase DNA bridges. Curr. Opin. Genet. Dev. 26, 1–5 (2014).
Sarlós, K. et al. Reconstitution of anaphase DNA bridge recognition and disjunction. Nat. Struct. Mol. Biol. 25, 868–876 (2018).
Champoux, J. J. DNA topoisomerases: structure, function, and mechanism. Annu. Rev. Biochem. 70, 369–413 (2001).
Pommier, Y., Sun, Y., Huang, S. Y. N. & Nitiss, J. L. Roles of eukaryotic topoisomerases in transcription, replication and genomic stability. Nature Rev. Molec. Cell Biol. 17, 703–721 (2016).
Hutchins, J. R. A. et al. Systematic analysis of human protein complexes identifies chromosome segregation proteins. Science 328, 593–599 (2010).
Xu, D. et al. RMI, a new OB-fold complex essential for Bloom syndrome protein to maintain genome stability. Genes Dev. 22, 2843–2855 (2008).
Forterre, P. A hot story from comparative genomics: reverse gyrase is the only hyperthermophile-specific protein. Trends Genet. 18, 236–238 (2002).
Bettotti, P. et al. Structure and properties of DNA molecules over the full range of biologically relevant supercoiling states. Sci. Rep. 8, 6163 (2018).
Bazett-Jones, D. P., Kimura, K. & Hirano, T. Efficient supercoiling of DNA by a single condensin complex as revealed by electron spectroscopic imaging. Mol. Cell 9, 1183–1190 (2002).
Liu, L. F. & Wang, J. C. Supercoiling of the DNA template during transcription. Proc. Natl Acad. Sci. USA 84, 7024–7027 (1987).
Cejka, P., Plank, J. L., Dombrowski, C. C. & Kowalczykowski, S. C. Decatenation of DNA by the S. cerevisiae Sgs1-Top3-Rmi1 and RPA complex: a mechanism for disentangling chromosomes. Mol. Cell 47, 886–896 (2012).
Yang, J., Bachrati, C. Z., Ou, J., Hickson, I. D. & Brown, G. W. Human topoisomerase IIIα is a single-stranded DNA decatenase that is stimulated by BLM and RMI1. J. Biol. Chem. 285, 21426–21436 (2010).
Kirkegaard, K. & Wang, J. C. Bacterial DNA topoisomerase I can relax positively supercoiled DNA containing a single-stranded loop. J. Mol. Biol. 185, 625–637 (1985).
Charvin, G., Strick, T. R., Bensimon, D. & Croquette, V. Tracking topoisomerase activity at the single-molecule level. Annu. Rev. Biophys. Biomol. Struct. 34, 201–219 (2005).
Lulchev, P. & Klostermeier, D. Reverse gyrase: recent advances and current mechanistic understanding of positive DNA supercoiling. Nucleic Acids Res. 42, 8200–8213 (2014).
Prasad, T. K. et al. A DNA-translocating Snf2 molecular motor: Saccharomyces cerevisiae Rdh54 displays processive translocation and extrudes DNA loops. J. Mol. Biol. 369, 940–953 (2007).
Nimonkar, A. V., Amitani, I., Baskin, R. J. & Kowalczykowski, S. C. Single molecule imaging of Tid1/Rdh54, a Rad54 homolog that translocates on duplex DNA and can disrupt joint molecules. J. Biol. Chem. 282, 30776–30784 (2007).
Lia, G. et al. Direct observation of DNA distortion by the RSC complex. Mol. Cell 21, 417–425 (2006).
Ganji, M. et al. Real-time imaging of DNA loop extrusion by condensin. Science 360, 102–105 (2018).
Strick, T. R., Allemand, J. F., Bensimon, D., Bensimon, A. & Croquette, V. The elasticity of a single supercoiled DNA molecule. Science 271, 1835–1837 (1996).
Corless, S. & Gilbert, N. Investigating DNA supercoiling in eukaryotic genomes. Brief. Funct. Genomics 16, 379–389 (2017).
Davoli, T. & de Lange, T. Telomere-driven tetraploidization occurs in human cells undergoing crisis and promotes transformation of mouse cells. Cancer Cell 21, 765–776 (2012).
Fujiwara, T. et al. Cytokinesis failure generating tetraploids promotes tumorigenesis in p53-null cells. Nature 437, 1043–1047 (2005).
Logan, C. V. et al. Mutations in TOP3A cause a Bloom syndrome-like disorder. Am. J. Hum. Genet. 103, 221–231 (2018).
Chapin Rodríguez, A. & Stock, D. Crystal structure of reverse gyrase: Insights into the positive supercoiling of DNA. EMBO J. 21, 418–426 (2002).
Triebei, H., Bär, H., Walter, A., Burckhardt, G. & Zimmer, C. H. Modulation of DNA supercoiling by interaction with netropsin and other minor groove binders. J. Biomol. Struct. Dyn. 11, 1085–1105 (1994).
Manosas, M. et al. Magnetic tweezers for the study of DNA tracking motors. Methods Enzymol. 475, 297–320 (2010).
Moghadam-Kamrani, S. J. & Keyomarsi, K. Synchronization of the cell cycle using lovastatin. Cell Cycle 7, 2434–2440 (2008).
Bizard, A. H., Nielsen, C. F. & Hickson, I. D. Detection of ultrafine anaphase bridges. Methods Mol. Biol. 1672, 495–508 (2018).
Acknowledgements
We thank R. Singh Thakur for purification of recombinant proteins, and members of the Hickson group for helpful discussions. We also thank E. Hoffmann, H. Mankouri and C. Nielsen for helpful comments on the manuscript, S. Kowalczykowski (University of California at Davis, USA) for purified E. coli Top3 and M. T. Kanemaki (National Institute of Genetics, Mishima, Shizuoka, Japan) for the degron cells. This work was supported by the Danish National Research Foundation (No. DNRF115, to I.D.H.), the European Union Horizon 2020 ‘Chromavision’ (No. 665233, to I.D.H.) and the Nordea Foundation (to I.D.H.). T. Hassenkam wishes to thank the Villum foundation “Experiment” for support.
Author information
Authors and Affiliations
Contributions
A.H.B. performed the experiments. A.H.B. and I.D.H. designed the experiments and analyzed the data. J.-F.A. helped carry out the experiments with magnetic tweezers and data analysis. T.H. assisted with A.F.M.’s experiments. M.P. assisted with cell biology experiments. K.S. and M.I.S. assisted with ensemble biochemistry experiments. I.D.H. supervised the work. A.H.B. and I.D.H. wrote the manuscript, and all authors edited it.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 PICH cooperates with TRR to induce DNA positive supercoiling.
(a–c) Representative 1D agarose gel electrophoresis showing the influence of increasing concentrations of PICH on the topoisomerase activity catalyzed by hTop1 (a), hTop2a (b), and TRR (c) using a negatively supercoiled plasmid as a substrate. Agarose gel electrophoresis was either performed in the absence (neutral; a, b) or in the presence of chloroquine (c) in order to reveal variations of topology within the negatively supercoiled topoisomers. For each experiment, relaxed (rel), negatively (-) moderately positively (rel+), and positively (+) supercoiled plasmids were used as markers. (d) Representative 2D agarose gel electrophoresis of the reaction products shown in (c). For each panel, representative images of at least 3 independent experiments are presented. Each independent experiment lead to similar results.
Supplementary Figure 2 PICH and TRR introduce a high density of positive supercoiling.
Representative AFM topographs of the relaxed substrate (a), a positively supercoiled (+8) marker (b), and the PICH-TRR reaction product (c). For each example, a cartoon depicting the conformation of two selected molecules is presented. For each panel, images representative of at least 20 images collected for 3 independent experiments are presented. All experiments lead to similar results. (d) Additional examples of representative AFM topographs of several PICH-TRR reaction products. For each topological form, a zoomed image is presented. White arrowheads in the zoomed panels indicate the presence of height peaks. White numbers indicate the amount of positive supercoils in the corresponding field of view as estimated by counting of the height peaks. Height scale bar is common for all images, and is shown on the right. XY scale bar is 100 nm.
Supplementary Figure 3 Evidence that supercoiling occurs via a translocation-dependent mechanism.
(a, b) Schematic representation of mechanistic models for PICH-mediated positive supercoiling. PICH is depicted as a red circle. Positive and negative torsional stress induced by PICH translocation are indicated by red “+” and “-” symbols (Translocation models; b). The (-) and (+) symbols indicate the topology of DNA segments resulting from PICH wrapping DNA (Wrapping model; a) or from PICH translocation (Translocation model; b). Net supercoiling level (ΔLk) of each intermediate is indicated in the gray bar below each panel. (a) In the wrapping model, the bound DNA segment might be wrapped around PICH in a positive supercoiling conformation, leading to the introduction of compensatory negative supercoils in the unbound DNA segment. Type IA (upper panel) and Type IB or II topoisomerases (lower panel) then relax the negative supercoils, resulting into the introduction of net positive supercoiling. (b) In the translocation model, and according to the twin-supercoiling domain model, PICH translocation might lead to the redistribution of DNA torsional stress leading to the accumulation of unconstrained positive and negative supercoiling, respectively, in front and behind PICH. Type IA (upper panel) would only relax the negative supercoils, resulting in the introduction of net positive supercoiling. Type IB or II topoisomerases (lower panel) would relax both positive and negative supercoils, leading to no net alteration in supercoiling. A topological barrier induced by PICH is depicted as a black line and red square. (c-f) Representative 1D agarose gel electrophoresis showing the influence of PICH on the activity catalyzed by increasing concentrations of wgTop1 (c), ecTop1 (d), ecTop3 (e), and TRR (f) using a negatively supercoiled plasmid as a substrate. Agarose gel electrophoresis was performed in the absence (neutral) (c, d) or in the presence of chloroquine (e, f) in order to reveal variations of topology within the negatively supercoiled topoisomers. For each experiment, open circular (oc), linear (lin), relaxed (rel), negatively (-) and positively (+) supercoiled plasmids were used as markers. For each panel, representative images out of at least 3 independent experiments are presented. Each independent experiment lead to similar results. (g) Additional examples of representative AFM topographs of open circular DNA plasmids incubated in the presence of PICH and ATPγS. Height scale bar is common for all images and is shown on the right. XY scale bar is 100 nm. Images representative of at least 10 images collected for 3 independent experiments are presented. All experiments lead to similar results.
Supplementary Figure 4 Evidence that PICH exhibits a DNA loop extrusion activity and induces redistribution of DNA torsional stress.
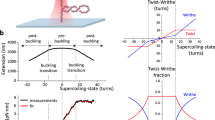
(a) Cartoon summarizing the features of the magnetic tweezer setup. (b) Additional representative traces showing the variation of DNA extension over time observed in the presence of PICH and ATP, at a force of 2 pN and on nicked DNA molecules. (c) Histogram to indicate the processivity of each event observed on nicked DNA molecules and at 2 pN (n = 273 events; bin size = 10 nm; errors bars are statistical errors; the red curve correspond to an exponential fit. (d) Representative traces showing the variation of DNA extension over time observed in the presence of PICH and ATP on nicked DNA molecules at various forces; 0.5 pN, 2 pN or 8 pN. (e) Scan showing the influence of rotation of the magnets on the extension of a nicked (gray curve) and a coilable (purple curve) DNA molecule at a force of 2 pN. (f) Additional representative traces showing the variation of DNA extension over time observed in the presence of PICH and ATP, at a force of 2 pN, on coiled (+ 100) DNA molecules.
Supplementary Figure 5 TRR facilitates decatenation of UFBs by Top2a.
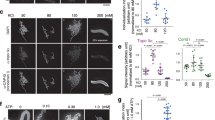
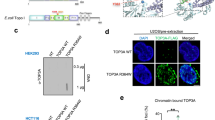
(a) Western blot analysis of Top3a, PICH and Top2a in HCT116-Top3aAID (Top3dg) and in the control cell line (430) in metaphase arrested cells incubated for 3 h in the absence (-) or in the presence (+) of Auxin. For each panel, representative images out of at least 3 independent experiments are presented. Uncropped blots images are shown in Supplementary Data Set 1. (b) Quantification of the average number of UFBs in the control cell line (430) exposed (red bars) or not (gray bars) to Auxin. Cells were treated with solvent (-) or either 50 nM or 100 nM ICRF193, as indicated below. Error bars denote the standard deviation of 3 independent replicates (n = 80cells/replicate) (c) Additional representative immunofluorescence images of HCT116-Top3aAID cells following Top3a depletion (+Auxin) in cells progressing through anaphase in the absence of ICRF193. UFBs are marked by PICH (blue) and centromeres are marked by CREST (red). The zoomed image shows CREST-positive staining at the tips of UFBs. Scale bar is 10 µm. Representative images of at least 20 images collected for 3 independent replicates are presented.
Supplementary information
Supplementary Figures and Supplementary Dataset
Supplementary Figures 1–5 and Supplementary Dataset 1
Rights and permissions
About this article
Cite this article
Bizard, A.H., Allemand, JF., Hassenkam, T. et al. PICH and TOP3A cooperate to induce positive DNA supercoiling. Nat Struct Mol Biol 26, 267–274 (2019). https://doi.org/10.1038/s41594-019-0201-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0201-6
This article is cited by
-
PICH deficiency limits the progression of MYC-induced B-cell lymphoma
Blood Cancer Journal (2024)
-
Replication-associated formation and repair of human topoisomerase IIIα cleavage complexes
Nature Communications (2023)
-
Human topoisomerases and their roles in genome stability and organization
Nature Reviews Molecular Cell Biology (2022)
-
PICH acts as a force-dependent nucleosome remodeler
Nature Communications (2022)
-
Visualizing RNA polymers produced by hot wet-dry cycling
Scientific Reports (2022)