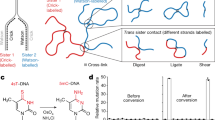
Abstract
In meiotic prophase, chromosomes are organized into compacted loop arrays to promote homolog pairing and recombination. Here, we probe the architecture of the mouse spermatocyte genome in early and late meiotic prophase using chromosome conformation capture (Hi-C). Our data support the established loop array model of meiotic chromosomes, and infer loops averaging 0.8–1.0 megabase pairs (Mb) in early prophase and extending to 1.5–2.0 Mb in late prophase as chromosomes compact and homologs undergo synapsis. Topologically associating domains (TADs) are lost in meiotic prophase, suggesting that assembly of the meiotic chromosome axis alters the activity of chromosome-associated cohesin complexes. While TADs are lost, physically separated A and B compartments are maintained in meiotic prophase. Moreover, meiotic DNA breaks and interhomolog crossovers preferentially form in the gene-dense A compartment, revealing a role for chromatin organization in meiotic recombination. Finally, direct detection of interhomolog contacts genome-wide reveals the structural basis for homolog alignment and juxtaposition by the synaptonemal complex.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All custom scripts and code are available at Github (https://github.com/lucaspatel/nsmb_mousehic) or from the authors (K.D.C.) upon request. All sequencing data have been deposited in the NCBI Gene Expression Omnibus database under accession number GSE122622: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE122622. All previously published data used in our analysis are available at the links below: GSM1908921: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSM1908921; GSM1954839: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSM1954839; ERS076381: ftp://ftp-mouse.sanger.ac.uk/current_snps/strain_specific_vcfs/CAST_EiJ.mgp.v5.snps.dbSNP142.vcf.gz; GSE101406: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE101406; GSM1083638: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSM1083638; SRR772029/SRR7720230 (processed): http://www.smallrnagroup.uni-mainz.de/piRNAclusterDB.html; SRR772029/GSM1096583 (raw data): https://www.ncbi.nlm.nih.gov/sra/SRX248863; SRR7720230/GSM1096584 (raw data): https://www.ncbi.nlm.nih.gov/sra/SRX248864
References
Keeney, S., Lange, J. & Mohibullah, N. Self-organization of meiotic recombination initiation: general principles and molecular pathways. Annu. Rev. Genet. 48, 187–214 (2014).
Zickler, D. & Kleckner, N. Meiotic chromosomes: integrating structure and function. Annu. Rev. Genet. 33, 603–754 (1999).
Hassold, T., Hall, H. & Hunt, P. The origin of human aneuploidy: where we have been, where we are going. Hum. Mol. Genet. 2, R203–R208 (2007).
McNicoll, F., Stevense, M. & Jessberger, R. Cohesin in gametogenesis. Curr. Top. Dev. Biol. 102, 1–34 (2013).
Lee, J. Roles of cohesin and condensin in chromosome dynamics during mammalian meiosis. J. Reprod. Dev. 59, 431–436 (2013).
Rankin, S. Complex elaboration: making sense of meiotic cohesin dynamics. FEBS. J. 282, 2426–2443 (2015).
West, A. M. V. et al. A conserved mechanism for meiotic chromosome organization through self-assembly of a filamentous chromosome axis core. Preprint at bioRxiv https://doi.org/10.1101/375220 (2018).
Blat, Y., Protacio, R. U., Hunter, N. & Kleckner, N. Physical and functional interactions among basic chromosome organizational features govern early steps of meiotic chiasma formation. Cell 111, 791–802 (2002).
Panizza, S. et al. Spo11-accessory proteins link double-strand break sites to the chromosome axis in early meiotic recombination. Cell 146, 372–383 (2011).
Page, S. L. & Hawley, R. S. The genetics and molecular biology of the synaptonemal complex. Annu. Rev. Cell. Dev. Biol. 20, 525–558 (2004).
Margolin, G., Khil, P. P., Kim, J., Bellani, M. A. & Camerini-Otero, R. D. Integrated transcriptome analysis of mouse spermatogenesis. BMC Genomics 15, 39 (2014).
Parsons, G. G. & Spencer, C. A. Mitotic repression of RNA polymerase II transcription is accompanied by release of transcription elongation complexes. Mol. Cell. Biol. 17, 5791–5802 (1997).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Gibcus, J. H. et al. A pathway for mitotic chromosome formation. Science 359, eaao6135 (2018).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Hogarth, C. A. et al. Turning a spermatogenic wave into a tsunami: synchronizing murine spermatogenesis using WIN 18,446. Biol. Reprod. 88, 40 (2013).
Cole, F. et al. Mouse tetrad analysis provides insights into recombination mechanisms and hotspot evolutionary dynamics. Nat. Genet. 46, 1072–1080 (2014).
Yan, J. et al. Histone H3 lysine 4 monomethylation modulates long-range chromatin interactions at enhancers. Cell Res. 28, 204–220 (2018).
Wutz, G. et al. Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J. 36, 3573–3599 (2017).
Jost, D., Carrivain, P., Cavalli, G. & Vaillant, C. Modeling epigenome folding: formation and dynamics of topologically associated chromatin domains. Nucleic Acids Res. 42, 9553–9561 (2014).
Di Pierro, M., Zhang, B., Aiden, E. L., Wolynes, P. G. & Onuchic, J. N. Transferable model for chromosome architecture. Proc. Natl Acad. Sci. USA 113, 12168–12173 (2016).
Falk, M. et al. Heterochromatin drives organization of conventional and inverted nuclei. Preprint at bioRxiv https://doi.org/10.1101/244038 (2018).
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Biswas, U., Hempel, K., Llano, E., Pendas, A. & Jessberger, R. Distinct roles of meiosis-specific cohesin complexes in mammalian spermatogenesis. PLoS Genet. 12, e1006389 (2016).
Muller, H. et al. Characterizing meiotic chromosomes’ structure and pairing using a designer sequence optimized for Hi-C. Mol. Syst. Biol. 14, e8293 (2018).
Schalbetter, S. A., Fudenberg, G., Baxter, J., Pollard, K. S. & Neale, M. J. Principles of meiotic chromosome assembly. Preprint at bioRxiv https://doi.org/10.1101/442038 (2018).
Sun, X. et al. Transcription dynamically patterns the meiotic chromosome-axis interface. eLife 4, 8522 (2015).
Shopland, L. S. et al. Folding and organization of a contiguous chromosome region according to the gene distribution pattern in primary genomic sequence. J. Cell. Biol. 174, 27–38 (2006).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320.e24 (2017).
Vagin, V. V. et al. A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313, 320–324 (2006).
Siomi, M. C., Sato, K., Pezic, D. & Aravin, A. A. PIWI-interacting small RNAs: the vanguard of genome defence. Nat. Rev. Mol. Cell Biol. 12, 246–258 (2011).
Buckley, M. S. & Lis, J. T. Imaging RNA Polymerase II transcription sites in living cells. Curr. Opin. Genet. Dev. 25, 126–130 (2014).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Chong, S. et al. Imaging dynamic and selective low-complexity domain interactions that control gene transcription. Science 361, eaar2555 (2018).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Cho, W.-K. et al. Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 361, 412–415 (2018).
Modzelewski, A. J. et al. Dgcr8 and Dicer are essential for sex chromosome integrity during meiosis in males. J. Cell. Sci. 128, 2314–2327 (2015).
Page, J. et al. Inactivation or non-reactivation: what accounts better for the silence of sex chromosomes during mammalian male meiosis? Chromosoma. 121, 307–326 (2012).
de Vries, M. et al. Human male meiotic sex chromosome inactivation. PLoS ONE 7, e31485 (2012).
Turner, J. M. A. et al. BRCA1, histone H2AX phosphorylation, and male meiotic sex chromosome inactivation. Curr. Biol. 14, 2135–2142 (2004).
Flyamer, I. M. et al. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 544, 110–114 (2017).
Marsden, M. P. & Laemmli, U. K. Metaphase chromosome structure: evidence for a radial loop model. Cell 17, 849–858 (1979).
Liang, Z. et al. Chromosomes progress to metaphase in multiple discrete steps via global compaction/expansion cycles. Cell 161, 1124–1137 (2015).
Novak, I. et al. Cohesin Smc1beta determines meiotic chromatin axis loop organization. J. Cell. Biol. 180, 83–90 (2008).
Revenkova, E. et al. Cohesin SMC1 beta is required for meiotic chromosome dynamics, sister chromatid cohesion and DNA recombination. Nat. Cell Biol. 6, 555–562 (2004).
Yuan, L. et al. Female germ cell aneuploidy and embryo death in mice lacking the meiosis-specific protein SCP3. Science 296, 1115–1118 (2002).
Gassler, J. et al. A mechanism of cohesin-dependent loop extrusion organizes zygotic genome architecture. EMBO J. 36, 3600–3618 (2017).
Pan, J. et al. A hierarchical combination of factors shapes the genome-wide topography of yeast meiotic recombination initiation. Cell 144, 719–731 (2011).
Gerton, J. L. et al. Global mapping of meiotic recombination hotspots and coldspots in the yeast Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 97, 11383–11390 (2000).
Baudat, F. & Nicolas, A. Clustering of meiotic double-strand breaks on yeast chromosome III. Proc. Natl Acad. Sci. USA 94, 5213–5218 (1997).
Paigen, K. & Petkov, P. M. PRDM9 and its role in genetic recombination. Trends Genet. 34, 291–300 (2018).
Brick, K., Smagulova, F., Khil, P., Camerini-Otero, R. D. & Petukhova, G. V. Genetic recombination is directed away from functional genomic elements in mice. Nature 485, 642–645 (2012).
Plug, A. W., Xu, J., Reddy, G., Golub, E. I. & Ashley, T. Presynaptic association of Rad51 protein with selected sites in meiotic chromatin. Proc. Natl Acad. Sci. USA 93, 5920–5924 (1996).
Holmquist, G. P. Chromosome bands, their chromatin flavors, and their functional features. Am. J. Hum. Genet. 51, 17–37 (1992).
Walker, M. et al. Affinity-seq detects genome-wide PRDM9 binding sites and reveals the impact of prior chromatin modifications on mammalian recombination hotspot usage. Epigenetics Chromatin 8, 31 (2015).
Lange, J. et al. The landscape of mouse meiotic double-strand break formation, processing, and repair. Cell 167, 695–708.e16 (2016).
Smagulova, F., Brick, K., Pu, Y., Camerini-Otero, R. D. & Petukhova, G. V. The evolutionary turnover of recombination hot spots contributes to speciation in mice. Genes Dev. 30, 266–280 (2016).
Baker, C. L. et al. PRDM9 drives evolutionary erosion of hotspots in Mus musculus through haplotype-specific initiation of meiotic recombination. PLoS Genet. 11, e1004916 (2015).
Liu, E. Y. et al. High-resolution sex-specific linkage maps of the mouse reveal polarized distribution of crossovers in male germline. Genetics 197, 91–106 (2014).
Turner, J. M. A. Meiotic silencing in mammals. Annu. Rev. Genet. 49, 395–412 (2015).
Turner, J. M. A. et al. Silencing of unsynapsed meiotic chromosomes in the mouse. Nat. Genet. 37, 41–47 (2005).
Baarends, W. M. et al. Silencing of unpaired chromatin and histone H2A ubiquitination in mammalian meiosis. Mol. Cell. Biol. 25, 1041–1053 (2005).
Homolka, D., Ivanek, R., Capkova, J., Jansa, P. & Forejt, J. Chromosomal rearrangement interferes with meiotic X chromosome inactivation. Genome Res. 17, 1431–1437 (2007).
McKee, B. D. & Handel, M. A. Sex chromosomes, recombination, and chromatin conformation. Chromosoma 102, 71–80 (1993).
Blat, Y. & Kleckner, N. Cohesins bind to preferential sites along yeast chromosome III, with differential regulation along arms versus the centric region. Cell 98, 249–259 (1999).
Zhang, L. et al. Topoisomerase II mediates meiotic crossover interference. Nature 511, 551–556 (2014).
Li, X. Z. et al. An ancient transcription factor initiates the burst of piRNA production during early meiosis in mouse testes. Mol. Cell 50, 67–81 (2013).
Hogarth, C. A. & Griswold, M. D. The key role of vitamin A in spermatogenesis. J. Clin. Invest. 120, 956–962 (2010).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
van de Geijn, B., McVicker, G., Gilad, Y. & Pritchard, J. K. WASP: allele-specific software for robust molecular quantitative trait locus discovery. Nat. Methods 12, 1061–1063 (2015).
Keane, T. M. et al. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature 477, 289–294 (2011).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst 3, 95–98 (2016).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst 3, 99–101 (2016).
Knight, P. A. & Ruiz, D. A fast algorithm for matrix balancing. IMA J. Numer. Anal. 33, 1029–1047 (2013).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinformatics 14, 178–192 (2013).
Gel, B. et al. regioneR: an R/Bioconductor package for the association analysis of genomic regions based on permutation tests. Bioinformatics 32, 289–291 (2016).
Rosenkranz, D. piRNA cluster database: a web resource for piRNA producing loci. Nucleic Acids Res. 44, D223–D230 (2016).
Rosenkranz, D. & Zischler, H. proTRAC—a software for probabilistic piRNA cluster detection, visualization and analysis. BMC Bioinformatics 13, 5 (2012).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Acknowledgements
We thank M. Handel for the kind gift of H1T antibodies, members of the Corbett, Cole and Ren labs, and A. Desai for helpful discussions. We thank S. Aigner, M. Neale, G. Fudenberg and L. Mirny for helpful suggestions on Hi-C data interpretation, and M. Griswold and C. Hogarth for assistance with synchronization of spermatogenesis. K.D.C. acknowledges support from the National Institutes of Health (grant No. R01GM104141). F.C. acknowledges support from the National Institutes of Health (grant No. DP2HD087943). K.D.C. and B.R. acknowledge support from the Ludwig Institute for Cancer Research. We acknowledge the National Institutes of Health (grant No. CA16672) for support of the Research Animal Support Facility Smithville, and the CPRIT (grant No. RP170628) for support of the Flow Cytometry and Cellular Imaging Core. R.K. is supported by a CPRIT Research Training Award (No. RP170067). R.R. was supported by a Ruth L. Kirschstein National Research Service Award (no. NIH/NCI T32 CA009523).
Author information
Authors and Affiliations
Contributions
K.D.C., F.C. and B.R. conceived and planned the study. R.K. performed spermatocyte isolation and characterization. L.P. adapted and implemented the Hi-C data analysis pipeline and performed Hi-C data analysis. S.C. and R.H. prepared sequencing libraries and performed initial Hi-C data analysis. S.R., Y.Q. and R.R. provided valuable input for data processing and analysis. K.D.C. and F.C. wrote the manuscript with input from B.R. and all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Synchronization and isolation of meiotic prophase spermatocytes.
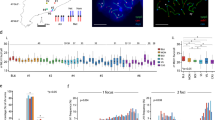
(a) Experimental workflow. C57BL/6 x CAST/EiJ F1 hybrid male mice were injected daily from 2–8 days post-partum with WIN18,446, then injected with retinoic acid at 9 days post-partum to synchronize spermatogenesis. Treated animals were allowed to recover for 21–47 days, then spermatocytes were isolated at time-points enriched at specific stages of prophase (J. Clin. Invest. 120, 956–962, 2010). The zygonema/pachynema stages for each wave of spermatogenesis are indicated by color: red (second wave), orange (third), green (fourth), and blue (fifth). (b) Spermatocytes were isolated, stained with Hoescht 33342 and sorted by FACS to further enrich for either zygonema or pachynema-stage cells (1C: spermatids; SP: spermatogonia) (Nat. Genet. 46, 1072–1080, 2014). (c) Samples of FACS-sorted cells were removed for chromosome spreads, and stained for SYCP3 and H1T for stage scoring (Supplementary Table 1). Scale bar = 10 μm. (d) Meiotic chromosome axis length in pachynema spermatocytes. Graph showing total measured chromosome axis length in pachynema spermatocytes of B6 x CAST F1 hybrid mice (orange; 48 cells from two animals) and B6 x DBA F1 hybrid mice (blue, 334 cells from two animals). Black line with error bar indicates mean ± standard deviation (also shown below graph). N = 48 cells for B6 x CAST, 34 for B6 x DBA. (e) Genome-wide Hi-C contact map for ES cells in interphase. Atypical contacts between chromosomes 8 and 14 indicate that a small percentage of analyzed cells possess a translocation between these two chromosomes. (f) Genome-wide Hi-C contact map for zygonema cells, showing X-shaped interchromosomal contact patterns between all chromosomes. (g) Genome-wide Hi-C contact map for pachynema cells. interchromosomal contacts are reduced in pachynema relative to zygonema, and the X chromosome in particular is isolated in pachynema as it is packaged into the XY body.
Supplementary Figure 2 Reproducibility of Hi-C contact maps in biological replicates.
(a) Hi-C contact map for combined zygonema data (top), and individual maps for samples #1 and #2 (bottom). The dotted box indicates the chromosome region shown in Fig. 3b. CPKM = contacts per kb per billion mapped contacts (see Methods). (b) Hi-C contact map for combined pachynema data (top), and individual maps for samples #1, #2, and #3 (bottom).
Supplementary Figure 3 Interchromosomal contacts reveal the meiotic bouquet.
(a) Schematic of the bouquet present in early meiotic prophase, and its expected signature in interchromosomal Hi-C contact maps. (b) X-shaped interchromosomal contacts between meiotic prophase chromosomes, a result of physical alignment in the bouquet. Interchromosomal contacts are particularly strong at the centromeric ends of chromosomes. Consistent with the loss of the bouquet following homolog synapsis in pachynema, the X-shaped interchromosomal contact patterns are strongly reduced in this stage compared to zygonema. Color scale for all panels is white (zero Hi-C contacts per bin) to red (indicated number or higher Hi-C contacts per bin). (c) Interchromosomal contacts between chromosome 3 and chromosome X reveal isolation of the X chromosome into the XY body in pachynema.
Supplementary Figure 4 Compartment identity is maintained in meiotic prophase and dictates large-scale recombination patterns.
Eigenvector analysis of chromosomes 3 (a), 6 (b), and 12 (c) in interphase, zygonema, and pachynema Hi-C contact maps. A and B compartments are shown in blue and green, respectively. For each chromosome, DSB hotspots (Genes Dev. 30, 266–280, 2016) and annotated genes are also shown. Correlations were calculated using a two-tailed, non-parametric Spearman correlation coefficient. (d) Eigenvector analysis of chromosome X in interphase, zygonema, and pachynema. While the calculated correlation coefficient between interphase and zygonema is low (0.18), the overall compartment structure is similar. Compartment structure is completely lost in pachynema.
Supplementary Figure 5 Transcription-mediated interaction hubs in meiotic chromosomes.
Regions of chromosome 1 (a), 2 (b), and 4 (c) in interphase, zygonema, and pachynema. Shown in blue are piRNA clusters transcribed at 12.5 dpp (Mol. Cell 50, 67–81, 2013), and shown in green are RNA polymerase II binding peaks at 10 dpp (zygonema) or 16 dpp (pachynema) (BMC Genomics 15, 39, 2014).
Supplementary Figure 6 Contact probability versus distance by chromosome and haplotype.
Contact probability (P(s)) plots for E14 ES cells (a), zygonema (b), zygonema interhomolog (c), pachynema (d), and pachynema interhomolog (e). For each graph, genome-wide data is shown in black, and individual chromosomes are shown in rainbow colors, offset in Y by 0.1 units for each chromosome to improve clarity. The X chromosome is not shown in interhomolog graphs, as these cells contain only one X chromosome. (f) Contact probability versus distance plot for zygonema, showing all data (black) and data for each haplotype (B6 orange, Y offset 0.2 units; CAST purple, Y offset 0.4 units). (g) Contact probability versus distance plot for pachynema (calculated from pachynema sample #1, see Supplementary Table 1), colored as in (a). (h) Contact probability vs. genomic distance (P(s)) curves for the X chromosome (green/purple) vs. autosomes (black) in zygonema. (i) Contact probability vs. genomic distance (P(s)) curves for the X chromosome (green/purple) vs. autosomes (black) in pachynema.
Supplementary Figure 7 Interhomolog Hi-C contact maps.
(a) Hi-C contact maps showing interhomolog contacts for chromosome 3 in zygonema and pachynema. (b) Hi-C contact maps showing interhomolog contacts for chromosome 6 in zygonema and pachynema. (c) Overall Hi-C contact map for a region of chromosome 7 showing strong clustering of piRNA loci in pachynema. Dotted box indicates the region shown in Fig. 3b, and blue circles indicate strong clustering interactions. (d) Hi-C contact maps showing interhomolog contact maps (at two different contrast levels) of the same region as panel (c). The maps show evidence of transcribed-loci clustering between homologs, despite their low resolution and signal-to-noise ratio.
Supplementary Figure 8 Modeling interhomolog contacts as a convolution of P(s) functions.
(a) Graphical illustration of the mathematical convolution of two power-law functions. (b) Applicability of the convolution to interhomolog interactions in meiotic chromosomes: Intra-homolog contact probability versus genomic distance follows a power-law scaling function (top). Interactions between two juxtaposed and aligned loop arrays with identical power-law scaling can be modeled by a convolution of the respective scaling functions, resulting in a wider and shallower scaling function for interhomolog contacts (bottom). (c) Plot of the function P(s) = s−0.5 (orange), and a convolution of this function with itself, P(s) * P(s) (gray, normalized to 1 at x = 1). Both functions are symmetrical with respect to the Y axis (as in panels a and b), but only positive values are shown. The convolution was calculated using integer values for s in the interval from −30 to 30. Since the value of 0–0.5 is infinity, we used the value of 0.1–0.5 = 3.16 for the purposes of this calculation. Fitting the convolution data series (gray) to a power-law trendline yields a function with scaling proportional to s−0.206, close to the observed scaling of interhomolog contacts in meiotic prophase chromosomes. (d) Log-log plot of the graph shown in (c); this plot is equivalent to the log-log P(s) plots in Fig. 4.
Supplementary Figure 9 Distribution of meiotic DSBs and other chromatin features by compartment.
(a) regioneR analysis (Bioinformatics 32, 289–291, 2016) of the genome-wide overlap between the gene-rich A compartment (as assigned from the control interphase dataset) and 14,951 DSB hotspots in spermatocytes from a B6xCAST F1 hybrid mouse (Genes Dev. 30, 266–280, 2016). (b) regioneR analysis of 6948 PRDM9 peaks measured by ChIP-Seq on spermatocytes from a B6xCAST F1 hybrid mouse (PLoS Genet. 11, e1004916, 2015). (c) regioneR analysis of 80,856 H3K4me3 ChIP-Seq peaks in spermatocytes from a B6xCAST F1 hybrid mouse (PLoS Genet. 11, e1004916, 2015). (d) regioneR analysis as in (a) of 784 mapped crossovers between B6 and CAST chromosomes in a multi-species cross (Genetics 197, 91–106, 2014). (e) For each chromosome, the cumulative intensity distribution of B6xCAST hotspots (Genes Dev. 30, 266–280, 2016) was calculated for hotspots located in A (blue) or B (green) compartments (compartment calls from the control interphase dataset). P values were calculated using a Kolmogorov-Smirnov test.
Supplementary information
Supplementary Figures and Tables
Supplementary Figures 1–9, Supplementary Tables 1–2
Rights and permissions
About this article
Cite this article
Patel, L., Kang, R., Rosenberg, S.C. et al. Dynamic reorganization of the genome shapes the recombination landscape in meiotic prophase. Nat Struct Mol Biol 26, 164–174 (2019). https://doi.org/10.1038/s41594-019-0187-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0187-0
This article is cited by
-
A-MYB and BRDT-dependent RNA Polymerase II pause release orchestrates transcriptional regulation in mammalian meiosis
Nature Communications (2023)
-
ATM signaling modulates cohesin behavior in meiotic prophase and proliferating cells
Nature Structural & Molecular Biology (2023)
-
Granulosa cell mevalonate pathway abnormalities contribute to oocyte meiotic defects and aneuploidy
Nature Aging (2023)
-
3D chromatin remodelling in the germ line modulates genome evolutionary plasticity
Nature Communications (2022)
-
Meiotic sex chromosome inactivation and the XY body: a phase separation hypothesis
Cellular and Molecular Life Sciences (2022)