Abstract
The noncoding RNA Xist recruits silencing factors to the inactive X chromosome (Xi) and facilitates re-organization of Xi structure. Here, we examine the mouse epigenomic landscape of Xi and assess how Xist alters chromatin accessibility. Xist deletion triggers a gain of accessibility of select chromatin regions that is regulated by BRG1, an ATPase subunit of the SWI/SNF chromatin-remodeling complex. In vitro, RNA binding inhibits nucleosome-remodeling and ATPase activities of BRG1, while in cell culture Xist directly interacts with BRG1 and expels BRG1 from the Xi. Xist ablation leads to a selective return of BRG1 in cis, starting from pre-existing BRG1 sites that are free of Xist. BRG1 re-association correlates with cohesin binding and restoration of topologically associated domains (TADs) and results in the formation of de novo Xi ‘superloops’. Thus, Xist binding inhibits BRG1’s nucleosome-remodeling activity and results in expulsion of the SWI/SNF complex from the Xi.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
ATAC-seq, nChIP-seq and RNA-seq data have been deposited in GEO under accession code GSE109395. All other data are available from the corresponding author upon reasonable request.
References
Smeets, D. et al. Three-dimensional super-resolution microscopy of the inactive X chromosome territory reveals a collapse of its active nuclear compartment harboring distinct Xist RNA foci. Epigenetics Chromatin 7, 8 (2014).
Cremer, T. et al. The 4D nucleome: evidence for a dynamic nuclear landscape based on co-aligned active and inactive nuclear compartments. FEBS Lett. 589, 2931–2943 (2015).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Lee, J. T. Gracefully ageing at 50, X-chromosome inactivation becomes a paradigm for RNA and chromatin control. Nat. Rev. Mol. Cell. Biol. 12, 815–826 (2011).
Disteche, C. M. Dosage compensation of the sex chromosomes and autosomes. Semin. Cell Dev. Biol. 56, 9–18 (2016).
Monfort, A. & Wutz, A. Progress in understanding the molecular mechanism of Xist RNA function through genetics. Philos Trans. R Soc. Lond. B Biol. Sci. 372, 20160368 (2017).
Robert Finestra, T. & Gribnau, J. X chromosome inactivation: silencing, topology and reactivation. Curr. Opin. Cell Biol. 46, 54–61 (2017).
Clemson, C. M., McNeil, J. A., Willard, H. F. & Lawrence, J. B. XIST RNA paints the inactive X chromosome at interphase: evidence for a novel RNA involved in nuclear/chromosome structure. J. Cell Biol. 132, 259–275 (1996).
Simon, M. D. et al. High-resolution Xist binding maps reveal two-step spreading during X-chromosome inactivation. Nature 504, 465–469 (2013).
Minajigi, A. et al. Chromosomes. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 349, aab2276 (2015).
Giorgetti, L. et al. Structural organization of the inactive X chromosome in the mouse. Nature 535, 575–579 (2016).
Darrow, E. M. et al. Deletion of DXZ4 on the human inactive X chromosome alters higher-order genome architecture. Proc. Natl Acad. Sci. USA 113, E4504–E4512 (2016).
Costanzi, C. & Pehrson, J. R. Histone macroH2A1 is concentrated in the inactive X chromosome of female mammals. Nature 393, 599–601 (1998).
Heard, E. et al. Methylation of histone H3 at Lys-9 is an early mark on the X chromosome during X inactivation. Cell 107, 727–738 (2001).
Plath, K. et al. Role of histone H3 lysine 27 methylation in X inactivation. Science 300, 131–135 (2003).
Zhao, J., Sun, B. K., Erwin, J. A., Song, J. J. & Lee, J. T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322, 750–756 (2008).
Zhang, L. F., Huynh, K. D. & Lee, J. T. Perinucleolar targeting of the inactive X during S phase: evidence for a role in the maintenance of silencing. Cell 129, 693–706 (2007).
Splinter, E. et al. The inactive X chromosome adopts a unique three-dimensional conformation that is dependent on Xist RNA. Genes Dev. 25, 1371–1383 (2011).
Calabrese, J. M. et al. Site-specific silencing of regulatory elements as a mechanism of X inactivation. Cell 151, 951–963 (2012).
Wu, H. et al. Cellular resolution maps of X chromosome inactivation: implications for neural development, function, and disease. Neuron 81, 103–119 (2014).
Finn, E. H., Smith, C. L., Rodriguez, J., Sidow, A. & Baker, J. C. Maternal bias and escape from X chromosome imprinting in the midgestation mouse placenta. Dev. Biol. 390, 80–92 (2014).
Berletch, J. B. et al. Escape from X inactivation varies in mouse tissues. PLoS Genet. 11, e1005079 (2015).
Marks, H. et al. Dynamics of gene silencing during X inactivation using allele-specific RNA-seq. Genome Biol. 16, 149 (2015).
Nakayama, R. T. et al. SMARCB1 is required for widespread BAF complex-mediated activation of enhancers and bivalent promoters. Nat. Genet. 49, 1613–1623 (2017).
Blank, M. et al. A tumor suppressor function of Smurf2 associated with controlling chromatin landscape and genome stability through RNF20. Nat. Med. 18, 227–234 (2012).
Nozawa, R. S. et al. Human inactive X chromosome is compacted through a PRC2-independent SMCHD1-HBiX1 pathway. Nat. Struct. Mol. Biol. 20, 566–573 (2013).
Bao, X. et al. A novel ATAC-seq approach reveals lineage-specific reinforcement of the open chromatin landscape via cooperation between BAF and p63. Genome Biol. 16, 284 (2015).
Miller, E. L. et al. TOP2 synergizes with BAF chromatin remodeling for both resolution and formation of facultative heterochromatin. Nat. Struct. Mol. Biol. 24, 344–352 (2017).
Liu, Z., & Carter, A. C. & Chang, H. Y. Mechanistic insights in X-chromosome inactivation. Philos. Trans. R Soc. Lond. B Biol. Sci. 372, 20160356 (2017).
D, G. H., Kelley, D. R., Tenen, D., Bernstein, B. & Rinn, J. L. Widespread RNA binding by chromatin-associated proteins. Genome Biol. 17, 28 (2016).
Kawaguchi, T. et al. SWI/SNF chromatin-remodeling complexes function in noncoding RNA-dependent assembly of nuclear bodies. Proc. Natl Acad. Sci. USA 112, 4304–4309 (2015).
Tang, Y. et al. Linking long non-coding RNAs and SWI/SNF complexes to chromatin remodeling in cancer. Mol. Cancer 16, 42 (2017).
Lino Cardenas, C. L. et al. An HDAC9-MALAT1-BRG1 complex mediates smooth muscle dysfunction in thoracic aortic aneurysm. Nat. Commun. 9, 1009 (2018).
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Chu, C. et al. Systematic discovery of Xist RNA binding proteins. Cell 161, 404–416 (2015).
Cifuentes-Rojas, C., Hernandez, A., Sarma, K. & Lee, J. T. Regulatory interactions between RNA and Polycomb repressive complex 2. Mol. Cell 55, 171–185 (2014). ePub May 29, 2014.
Kaneko, S., Son, J., Bonasio, R., Shen, S. S. & Reinberg, D. Nascent RNA interaction keeps PRC2 activity poised and in check. Genes Dev. 28, 1983–1988 (2014).
Cajigas, I. et al. Evf2 lncRNA/BRG1/DLX1 interactions reveal RNA-dependent inhibition of chromatin remodeling. Development 142, 2641–2652 (2015).
Jeon, Y. & Lee, J. T. YY1 tethers Xist RNA to the inactive X nucleation center. Cell 146, 119–133 (2011).
de Dieuleveult, M. et al. Genome-wide nucleosome specificity and function of chromatin remodellers in ES cells. Nature 530, 113–116 (2016).
Lessing, D. et al. A high-throughput small molecule screen identifies synergism between DNA methylation and Aurora kinase pathways for X reactivation. Proc. Natl Acad. Sci. USA 113, 14366–14371 (2016).
Carrette, L. L. G. et al. A mixed modality approach towards Xi reactivation for Rett syndrome and other X-linked disorders. Proc. Natl Acad. Sci. USA 115, E668–E675 (2018).
Bhatnagar, S. et al. Genetic and pharmacological reactivation of the mammalian inactive X chromosome. Proc. Natl Acad. Sci. USA 111, 12591–12598 (2014).
Minkovsky, A. et al. A high-throughput screen of inactive X chromosome reactivation identifies the enhancement of DNA demethylation by 5-aza-2′-dC upon inhibition of ribonucleotide reductase. Epigenetics Chromatin 8, 42 (2015).
Sripathy, S. et al. Screen for reactivation of MeCP2 on the inactive X chromosome identifies the BMP/TGF-beta superfamily as a regulator of XIST expression. Proc. Natl Acad. Sci. USA 114, 1619–1624 (2017).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Bickmore, W. A. & van Steensel, B. Genome architecture: domain organization of interphase chromosomes. Cell 152, 1270–1284 (2013).
Bonev, B. et al. Multiscale 3D genome rewiring during mouse neural development. Cell 171, 557–572 (2017).
Phelan, M. L., Sif, S., Narlikar, G. J. & Kingston, R. E. Reconstitution of a core chromatin remodeling complex from SWI/SNF subunits. Mol. Cell 3, 247–253 (1999).
Wutz, A. & Jaenisch, R. A shift from reversible to irreversible X inactivation is triggered during ES cell differentiation. Mol. Cell 5, 695–705 (2000).
Csankovszki, G., Panning, B., Bates, B., Pehrson, J. R. & Jaenisch, R. Conditional deletion of Xist disrupts histone macroH2A localization but not maintenance of X inactivation. Nat. Genet. 22, 323–324 (1999).
Jegu, T., Aeby, E. & Lee, J. T. The X chromosome in space. Nat. Rev. Genet. 18, 377–389 (2017).
Hakimi, M. A. et al. A chromatin remodelling complex that loads cohesin onto human chromosomes. Nature 418, 994–998 (2002).
Wiechens, N. et al. The chromatin remodelling enzymes SNF2H and SNF2L position nucleosomes adjacent to CTCF and other transcription factors. PLoS Genet. 12, e1005940 (2016).
Chadwick, B. P. & Willard, H. F. Multiple spatially distinct types of facultative heterochromatin on the human inactive X chromosome. Proc. Natl Acad. Sci. USA 101, 17450–17455 (2004).
Pinter, S. F. et al. Spreading of X chromosome inactivation via a hierarchy of defined polycomb stations. Genome Res. 22, 1864–1876 (2012).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Luger, K., Rechsteiner, T. J. & Richmond, T. J. Preparation of nucleosome core particle from recombinant histones. Methods Enzymol. 304, 3–19 (1999).
Narlikar, G. J., Phelan, M. L. & Kingston, R. E. Generation and interconversion of multiple distinct nucleosomal states as a mechanism for catalyzing chromatin fluidity. Mol. Cell 8, 1219–1230 (2001).
Acknowledgements
T. Jégu is a European Molecular Biology Organization (EMBO) postdoctoral fellow (EMBO ALTF 1313–2015) and a young researcher Bettencourt Schueller Foundation awardee. R.E.K. is supported by NIH grant, R37-GM048405. This work was supported by funding to J.T.L. from the Rett Syndrome Research Trust, the LouLou Foundation, the National Institutes of Health (R01-DA36895) and the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
T.J., R.B. and J.T.L. designed the experiments and analyzed data. J.C.C. performed in vitro ATPase and remodeling assays. L.Y. performed the RNA-seq assays. D.C., A.S. and S.K.M. made all the plasmids. T.J. generated all data. R.B. performed all bioinformatics analyses, excepted the Hi-C analysis performed by C.Y.W. T.J. and M.E.G analyzed the immunoFISH. R.E.K. supervised J.C.C and S.K.M. T.J., R.B. and J.T.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.T.L. is a co-founder of and serves on the Scientific Advisory Boards of Translate Bio and Fulcrum Therapeutics.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Xist RNA regulates chromatin accessibility on Xi chromosome.
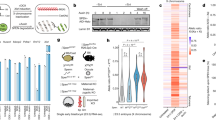
a, Scatterplots showing ATAC-seq signal correlation between two biological replicates of wild-type (left) or Xist-deletion (right) cells, n = 2 biological independent experiments. r2, Pearson coefficient. b, ATAC-seq peak-centered heat map of ATAC-seq signal (using composite reads) in Xist deletion and wild-type cells for the three major accessibility classes (left). Profiles presenting mean ATAC-seq signal (using composite reads) in the proximity of ATAC-seq peak center (right). c, Chromatin accessibility profiles measured by FAIRE–qPCR in wild-type and XaWT XiΔXist cells. Higher values express more accessible chromatin. Error bars represent mean ± s.d. P value indicates significant difference relative to wild-type cells (*P < 0.005; two-tailed Student’s t-test, n = 3 biologically independent experiments). Neg Ctl, negative control representing an inaccessible region. No signal was observed at Xist on the Xi allele in XaWT XiΔXist cells because the qPCR amplicon is localized in the deleted region
Supplementary Figure 2 Xist depletion in dermal fibroblasts induces increased chromatin accessibility, but not gene expression alterations on Xi chromosome.
a, Analysis performed for the second Xist-deletion line (XaWT XiΔXist_2) as indicated in Fig. 1b. P value was determined using a one-sided two-proportion z-test. b, Analysis performed for the second Xist-deletion line as described in Fig. 1c. c, Representative examples of peaks (highlighted) of the four accessibility classes in wild-type cells and a second Xist-deletion cell line. Similar results were obtained using another Xist-deletion line. d, Left, Venn diagrams showing relative overlap between two Xist-deletion lines per each accessibility class. Percentage in red indicates the fraction of classified regions of XaWT XiΔXist_2 that were classified identically (to the same accessibility class) in XaWT XiΔXist_1. Right, ATAC-seq peak-centered heat map of allele-specific ATAC-seq signal in the three cell lines. e, ATAC-seq peak-centered heat map of allele-specific ATAC-seq signal in two WT fibroblast lines (MEFs versus dermal (DF)). Xi-diff., ATAC peaks displaying difference on Xi chromosome between two WT cell lines. ATAC-seq peaks are clustered into three accessibility classes, and their relative percentage is designated (right). High concordance between two WT cell lines, as only four peaks (Xi-diff. peaks, corresponding to 1% of all peaks) display accessibility differences on Xi chromosome between two WT cell types. f, The four Xi-diff. peaks are located at the promoters of the four known escapee genes designated in the table. These four Xi-diff. peaks were classified as bi-allelic peaks when comparing the DF WT cell line to the two Xist-deletion cell lines (Supplementary Fig. 2d). These observations suggest that these four peaks display the same allelic characteristics in female dermal fibroblasts independently of Xist presence and that the differences observed between the two different WT fibroblast cell lines are rather tissue dependent (mouse embryonic versus dermal fibroblasts). g, Gene expression clustering based on SVM classification. SVM model trained to classify allelic signals into three accessibility classes (monoallelic, bi-allelic and Xi-restored) classified all expressed genes (the basis of their allelic signals) into two classes only (monoallelic and bi-allelic), whereas none of the genes was reproducibly classified as Xi-restored genes. h, Boxplots depicting gene expression as a function of ATAC-peak distance to the gene TSS. There is a correlation between gene expression and distance to the TSS of the closest ATAC peak; the closer the distance between the ATAC peak and its adjacent TSS, the higher the level that the gene is expressed. Box boundaries represent 25th and 75th percentiles; the center line represents the median; whiskers indicate ± 1.5 times the interquartile range (IQR), and points are the actual values of outliers
Supplementary Figure 3 Controls of MNase–qPCR and ChIP–qPCR experiments.
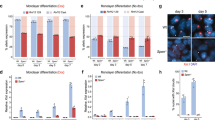
a, Time course of MNase sensitivity as measured by qPCR at representative Xi-restored genes and controls. Error bars represent mean ± s.d., n = 3 biologically independent experiments. b–d, Enrichment levels of the Polycomb repressive marks H3K27me3 (b), H2AK119Ub (c) and histone H3 (d) were measured by ChIP–qPCR in mutant and WT cells. Error bars represent mean ± s.d., n = 3 biologically independent experiments
Supplementary Figure 4 Xist RNA interaction inhibits BRG1 activities in vitro.
a,b. Quantification of SWI/SNF activities in the presence of different RNAs (top). The bar graph shows inhibition of in vitro SWI/SNF activities (IC50) by nine different RNAs (bottom). Error bars represent the s.d. of three biological replicates. c, The bar graph shows inhibition of in vitro SWI/SNF activities (IC50) by nine different RNAs normalized to RNA length
Supplementary Figure 5 Restoration of accessibility on the Xi depends on BRG1.
a, RT–qPCR analysis showing relative expression of the indicated genes in BRG1-knockdown (siBRG1) cells. Error bars represent mean ± s.d., n = 3 biologically independent experiments. b, Immunoblot analysis showing protein amounts of BRG1 and ZNF143 (control protein) in siBRG1 as compared to siCTL and siSNF2H cells. Each BRG1 western blot band was quantified and normalized to the ZNF143 band using ImageJ software (bottom). The experiment was repeated twice independently with similar results. c, Scatterplots showing ATAC-seq signal correlation between two biological replicates of WT cells treated with siCTL (top) or siBRG1 (bottom), n = 2 biologically independent experiments. d, Boxplots showing for each class distribution of allelic accessibility differences (siBRG1 versus siCTL; log2 FC) in WT cells. For each class, peak signals were obtained from allelic reads of Xa (top), monoallelic (n = 283), bi-allelic (n = 24), and Xi-restored (n = 30), or Xi (bottom), Bi-allelic (n = 22). P values were determined using a one-sided non-parametric Wilcoxon test. Monoallelic and Xi-restored classes are not shown for Xi read analysis because these classes do not display Xi reads (Fig. 1c). Box boundaries represent 25th and 75th percentiles; the center line represents the median; whiskers indicate ± 1.5 times the interquartile range (IQR), and points are the actual values of outliers. e, Scatterplots showing ATAC-seq signal correlation between two biological replicates of Xist-deletion cells treated with siCTL (top) or siBRG1 (bottom), n = 2 biologically independent experiments. f, Genome-wide scatterplots, using composite reads, showing chromatin accessibility changes between siBRG1- and siCTL-treated Xist-deletion cells. Peaks where chromatin accessibility was significantly increased (n = 1,233) or decreased (n = 16,328) as a result of BRG1 depletion are highlighted in red and green, respectively (FDR < 0.05). For each peak, an average value was calculated from two biological replicates. The Pearson correlation coefficient was calculated (r2). g, Boxplots showing the distribution of fold change accessibility values (log2 FC) between siBRG1- and siCTL-treated Xist-deletion cells. Fold change values between each condition were calculated based on all uniquely aligned reads (composite reads) of ATAC-seq peaks, monoallelic (n = 314), bi-allelic (n = 23), and Xi-restored (n = 35). P values were determined using a one-sided non-parametric Wilcoxon test. Box boundaries represent 25th and 75th percentiles; the center line represents the median; whiskers indicate ± 1.5 times the interquartile range (IQR), and points are the actual values of outliers. h, ATAC-seq read coverage of representative chromatin accessible regions in control and BRG1-knockdown Xist-deletion and WT cell lines. Two BRG1-dependnent ATAC-seq peaks with decreased accessibility post-BRG1 depletion are highlighted in green, and one BRG1-independent peak that remains unchanged upon BRG1 knockdown is highlighted in black (left). i, Chromatin accessibility profiles measured by FAIRE–qPCR in WT and XaWT XiΔXist cells treated with siCTL or siBRG1. Values were normalized to percentage input values obtained at the Gapdh and Actn1 promoters. Error bars represent mean ± s.d. P value indicates significant difference relative to siBRG1-treated cells (*P < 0.001; two-tailed Student’s t-test, n = 3 biologically independent experiments)
Supplementary Figure 6 Chromatin accessibility at Xi-restored regions is insensitive to CHD4 or SNF2H knockdown.
a,b, Same analyses as indicated before (see Supplementary Fig. 5a,b and Fig. 5c,d) for siCHD4 and siSNF2H, respectively. Error bars represent mean ± s.d., n = 3 biologically independent experiments. Peaks up in siCHD4 (n = 10,940) and peaks down in siCHD4 (n = 3,612). Boxplots show the distribution of fold change accessibility values (log2 FC) between siCHD4- (or siSNF2H-) and siCTL-treated WT cells. siCHD4: monoallelic (n = 310), bi-allelic (n = 24), and Xi-restored (n = 31). siSNF2H: monoallelic (n = 304), bi-allelic (n = 25), and Xi-restored (n = 33). P values were determined using a one-sided non-parametric Wilcoxon test. Box boundaries represent 25th and 75th percentiles; the center line represents the median; whiskers indicate ± 1.5 times the interquartile range (IQR), and points are actual values of outliers
Supplementary Figure 7 BRG1 regulates chromatin accessibility at Xi-restored regions.
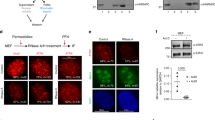
a, Immuno-RNA-FISH for H3K27me3 and Xist RNA. Percentages of cells displaying a colocalization between H3K27me3 and Xist cloud are shown. Scale bar, 5 μm. H3K27me3 intensities outside (green boxes) and inside (red) of the Xist cloud were calculated using ImageJ software. P value was determined using a two-tailed Student’s t-test. n = 135 cells. Box boundaries represent 25th and 75th percentiles; the center line represents the median; whiskers indicate ± 1.5 times the interquartile range (IQR), and points are the actual values of outliers. b, Immunoblot analysis showing the amount of BRG1 protein after BRG1 or IgG immunoprecipitation. Five times larger precipitate volume was loaded on the right well of IgG. The experiment was repeated twice independently with similar results. c, Representative examples of BRG1-bound regions displaying decreased chromatin accessibility upon BRG1 knockdown (highlighted). d, Scatterplots showing BRG1 ChIP-seq signal correlation between two biological replicates of WT (left) or Xist-deletion (right) cell lines, n = 2 biologically independent experiments. e, Boxplots showing the distribution of differences in allelic skewing for BRG1 peaks in WT cells on chromosomes X and 11. P value was determined using a one-sided non-parametric Wilcoxon test. Box boundaries represent 25th and 75th percentiles; the center line represents the median; whiskers indicate ± 1.5 times the interquartile range (IQR), and points are the actual values of outliers. Xa, cas reads. Xi, mus reads. f, Comparison of BRG1 ChIP-seq peak percentage on the Xi chromosome between XaWT XiΔXist and WT cells. P value was determined using a one-sided two-proportion z-test. g,h, Analyses conducted using composite reads as formerly indicated in Fig. 6c, d, respectively. i, BRG1 ChIP–qPCR performed in WT and Xist-deletion cells. Error bars represent mean ± s.d. P value indicates significant difference relative to WT cells (*P < 0.0001; two-tailed Student’s t-test, n = 3 biologically independent experiments). Neg Ctl, negative control representing a region not bound by BRG1. Pos Ctl, positive control representative of region bound by BRG1
Supplementary Figure 8 Annotation of BRG1 re-occupancy sites to pre-existing bi-allelic binding sites.
A superimposed heat map depicting all Xi-restored (red) and monoallelic (blue) BRG1 classified peaks within a window of ± 1 Mb centered on their adjacent bi-allelic peaks (designated by a green vertical line). All 60 bi-allelic BRG1 peaks are plotted as reference points. The majority of bi-allelic peaks (85%) manifest extended patterns of BRG1 re-occupancy emanating from pre-bound BRG1 sites (designated as ‘Nucleation-like’). Asterisks correspond to three representative regions presented in Fig. 7e
Supplementary Figure 9 Integrative ATAC-seq and SMC1a ChIP-seq analysis in Xist-deleted cells.
a, Left, ATAC-seq peak-centered heat map of allele-specific ATAC-seq and SMC1a ChIP-seq signals. Concordance between ATAC-seq and SMC1a profiles. Pearson correlation was calculated for each of the regions (rows) depicted in the heat map by comparing a 3-kb density matrix of allelic-specific ATAC-seq signal to a corresponding density matrix of allelic-specific SMC1A ChIP-seq. Heat map regions sorted (green vertical magnitude bar) based on their Pearson correlation coefficients (top is strongest) are presented. The distribution of correlation coefficients of each class is shown by boxplots (right), monoallelic (n = 356) and Xi-restored (n = 37). Box boundaries represent 25th and 75th percentiles; the center line represents the median; whiskers indicate ± 1.5 times the interquartile range (IQR), and points are the actual values of outliers. Right, profiles presenting mean ATAC-seq and mean SMC1a ChIP-seq signals around ATAC-seq peak center. b, Representative examples of Xi-restored ATAC-seq peak regions displaying restoration of SMC1a binding (highlighted). c, SMC1a peak-centered heat map of ATAC-seq signal and profiles presenting mean ATAC-seq signal around SMC1a ChIP-seq peak center in siCTL- and siBRG1-treated WT cells (using composite reads). Only 25% of SMC1a peaks displaying a significant ATAC-seq peak in a ± 1.5-kb flanking region are displayed. SMC1a peaks display ATAC-seq peaks at the same position (‘accessible peaks’ ~0 bp) (left) or in fewer cases at distal regions (‘distally accessible peaks’ ~400 bp) (right)
Supplementary Figure 10 Integrative analysis of re-accessibility, BRG1 and SMC1 re-occupancy sites and Hi-C data in WT and Xist-deleted cells.
Allele-specific Hi-C analysis map showing the results of Pearson correlation analysis of the Xi chromosome with chromosomal locations of restored ATAC, BRG1, cohesins and TAD regions across the Xi chromosome in WT (top) and Xist deletion (bottom) cells
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 supplementary notes
Supplementary Dataset 1
Uncropped gels
Supplementary Dataset 2
Data processing matrices
Supplementary Dataset 3
ATAC-seq accessibility classes
Supplementary Dataset 4
RNA-seq gene expression classes
Supplementary Dataset 5
PWMs of de novo motifs identified in accessibility classes
Supplementary Dataset 6
Accessibility differences in siRNAs
Supplementary Dataset 7
BRG1 ChIP-seq deposition classes
Supplementary Dataset 8
Sequences of primers used in this study (5ʹ–3ʹ)
Rights and permissions
About this article
Cite this article
Jégu, T., Blum, R., Cochrane, J.C. et al. Xist RNA antagonizes the SWI/SNF chromatin remodeler BRG1 on the inactive X chromosome. Nat Struct Mol Biol 26, 96–109 (2019). https://doi.org/10.1038/s41594-018-0176-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0176-8
This article is cited by
-
Long non-coding RNAs as the critical regulators of PI3K/AKT, TGF-β, and MAPK signaling pathways during breast tumor progression
Journal of Translational Medicine (2023)
-
Unconventional roles of chromatin remodelers and long non-coding RNAs in cell division
Cellular and Molecular Life Sciences (2023)
-
Long non-coding RNAs: definitions, functions, challenges and recommendations
Nature Reviews Molecular Cell Biology (2023)
-
The endothelial-enriched lncRNA LINC00607 mediates angiogenic function
Basic Research in Cardiology (2023)
-
Xist exerts gene-specific silencing during XCI maintenance and impacts lineage-specific cell differentiation and proliferation during hematopoiesis
Nature Communications (2022)