Abstract
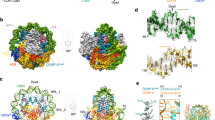
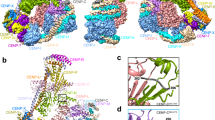
Kinetochores are multicomponent complexes responsible for coordinating the attachment of centromeric DNA to mitotic-spindle microtubules. The point centromeres of budding yeast are organized into three centromeric determining elements (CDEs), and are associated with the centromere-specific nucleosome Cse4. Deposition of Cse4 at CEN loci is dependent on the CBF3 complex that engages CDEIII to direct Cse4 nucleosomes to CDEII. To understand how CBF3 recognizes CDEIII and positions Cse4, we determined a cryo-EM structure of a CBF3−CEN complex. CBF3 interacts with CEN DNA as a head-to-head dimer that includes the whole of CDEIII and immediate 3' regions. Specific CEN-binding of CBF3 is mediated by a Cep3 subunit of one of the CBF3 protomers that forms major groove interactions with the conserved and essential CCG and TGT motifs of CDEIII. We propose a model for a CBF3−Cse4−CEN complex with implications for understanding CBF3-directed deposition of the Cse4 nucleosome at CEN loci.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bloom, K. & Costanzo, V. centromere structure and function. Prog. Mol. Subcell. Biol. 56, 515–539 (2017).
Musacchio, A. & Desai, A. A molecular view of kinetochore assembly and function. Biology (Basel) 6, 5 (2017).
Jenni, S., Dimitrova, Y. N., Valverde, R., Hinshaw, S. M. & Harrison, S. C. Molecular structures of yeast kinetochore subcomplexes and their roles in chromosome segregation. Cold Spring Harb. Symp. Quant. Biol. 82, 83–89 (2017).
Musacchio, A. The molecular biology of spindle assembly checkpoint signaling dynamics. Curr. Biol. 25, R1002–R1018 (2015).
Joglekar, A. P. A cell biological perspective on past, present and future investigations of the spindle assembly checkpoint. Biology (Basel) 5, 44 (2016).
Alfieri, C., Zhang, S. & Barford, D. Visualizing the complex functions and mechanisms of the anaphase promoting complex/cyclosome (APC/C). Open Biol. 7, 170204 (2017).
Clarke, L. & Carbon, J. Isolation of a yeast centromere and construction of functional small circular chromosomes. Nature 287, 504–509 (1980).
Fitzgerald-Hayes, M., Clarke, L. & Carbon, J. Nucleotide sequence comparisons and functional analysis of yeast centromere DNAs. Cell 29, 235–244 (1982).
Hieter, P. et al. Functional selection and analysis of yeast centromeric DNA. Cell 42, 913–921 (1985).
Peterson, J. B. & Ris, H. Electron-microscopic study of the spindle and chromosome movement in the yeast Saccharomyces cerevisiae. J. Cell Sci. 22, 219–242 (1976).
Cottarel, G., Shero, J. H., Hieter, P. & Hegemann, J. H. A 125-base-pair CEN6 DNA fragment is sufficient for complete meiotic and mitotic centromere functions in Saccharomyces cerevisiae. Mol. Cell. Biol. 9, 3342–3349 (1989).
Kingsbury, J. & Koshland, D. Centromere-dependent binding of yeast minichromosomes to microtubules in vitro. Cell 66, 483–495 (1991).
McGrew, J., Diehl, B. & Fitzgerald-Hayes, M. Single base-pair mutations in centromere element III cause aberrant chromosome segregation in Saccharomyces cerevisiae. Mol. Cell. Biol. 6, 530–538 (1986).
Ng, R. & Carbon, J. Mutational and in vitro protein-binding studies on centromere DNA from Saccharomyces cerevisiae. Mol. Cell. Biol. 7, 4522–4534 (1987).
Hegemann, J. H., Shero, J. H., Cottarel, G., Philippsen, P. & Hieter, P. Mutational analysis of centromere DNA from chromosome VI of Saccharomyces cerevisiae. Mol. Cell. Biol. 8, 2523–2535 (1988).
Meluh, P. B., Yang, P., Glowczewski, L., Koshland, D. & Smith, M. M. Cse4p is a component of the core centromere of Saccharomyces cerevisiae. Cell 94, 607–613 (1998).
Lechner, J. & Carbon, J. A 240 kd multisubunit protein complex, CBF3, is a major component of the budding yeast centromere. Cell 64, 717–725 (1991).
Furuyama, S. & Biggins, S. Centromere identity is specified by a single centromeric nucleosome in budding yeast. Proc. Natl Acad. Sci. USA 104, 14706–14711 (2007).
Cole, H. A., Howard, B. H. & Clark, D. J. The centromeric nucleosome of budding yeast is perfectly positioned and covers the entire centromere. Proc. Natl Acad. Sci. USA 108, 12687–12692 (2011).
Krassovsky, K., Henikoff, J. G. & Henikoff, S. Tripartite organization of centromeric chromatin in budding yeast. Proc. Natl Acad. Sci. USA 109, 243–248 (2012).
Bloom, K. S. & Carbon, J. Yeast centromere DNA is in a unique and highly ordered structure in chromosomes and small circular minichromosomes. Cell 29, 305–317 (1982).
Bloom, K. S. et al. Chromatin conformation of yeast centromeres. J. Cell Biol. 99, 1559–1568 (1984).
Saunders, M., Fitzgerald-Hayes, M. & Bloom, K. Chromatin structure of altered yeast centromeres. Proc. Natl Acad. Sci. USA 85, 175–179 (1988).
Funk, M., Hegemann, J. H. & Philippsen, P. Chromatin digestion with restriction endonucleases reveals 150-160 bp of protected DNA in the centromere of chromosome XIV in Saccharomyces cerevisiae. Mol. Gen. Genet. 219, 153–160 (1989).
Lechner, J. A zinc finger protein, essential for chromosome segregation, constitutes a putative DNA binding subunit of the Saccharomyces cerevisiae kinetochore complex, Cbf3. EMBO J. 13, 5203–5211 (1994).
Sorger, P. K. et al. Two genes required for the binding of an essential Saccharomyces cerevisiae kinetochore complex to DNA. Proc. Natl Acad. Sci. USA 92, 12026–12030 (1995).
Espelin, C. W., Kaplan, K. B. & Sorger, P. K. Probing the architecture of a simple kinetochore using DNA-protein crosslinking. J. Cell Biol. 139, 1383–1396 (1997).
Russell, I. D., Grancell, A. S. & Sorger, P. K. The unstable F-box protein p58-Ctf13 forms the structural core of the CBF3 kinetochore complex. J. Cell Biol. 145, 933–950 (1999).
Dechassa, M. L. et al. Structure and Scm3-mediated assembly of budding yeast centromeric nucleosomes. Nat. Commun. 2, 313 (2011).
Nakane, T., Kimanius, D., Lindahl, E. & Scheres, S. H. Characterisation of molecular motions in cryo-EM single-particle data by multi-body refinement in RELION. eLife 7, e36861 (2018).
Cho, U. S. & Harrison, S. C. Ndc10 is a platform for inner kinetochore assembly in budding yeast. Nat. Struct. Mol. Biol. 19, 48–55 (2011).
Perriches, T. & Singleton, M. R. Structure of yeast kinetochore Ndc10 DNA-binding domain reveals unexpected evolutionary relationship to tyrosine recombinases. J. Biol. Chem. 287, 5173–5179 (2012).
Leber, V., Nans, A. & Singleton, M. R. Structural basis for assembly of the CBF3 kinetochore complex. EMBO J. 37, 269–281 (2018).
Kaplan, K. B., Hyman, A. A. & Sorger, P. K. Regulating the yeast kinetochore by ubiquitin-dependent degradation and Skp1p-mediated phosphorylation. Cell 91, 491–500 (1997).
Bellizzi, J. J. 3rd, Sorger, P. K. & Harrison, S. C. Crystal structure of the yeast inner kinetochore subunit Cep3p. Structure 15, 1422–1430 (2007).
Purvis, A. & Singleton, M. R. Insights into kinetochore-DNA interactions from the structure of Cep3Delta. EMBO Rep. 9, 56–62 (2008).
King, D. A., Zhang, L., Guarente, L. & Marmorstein, R. Structure of HAP1-18-DNA implicates direct allosteric effect of protein-DNA interactions on transcriptional activation. Nat. Struct. Biol. 6, 22–27 (1999).
Pietrasanta, L. I. et al. Probing the Saccharomyces cerevisiae centromeric DNA (CEN DNA)-binding factor 3 (CBF3) kinetochore complex by using atomic force microscopy. Proc. Natl Acad. Sci. USA 96, 3757–3762 (1999).
Zhang, W., Lukoynova, N., Miah, S., Lucas, J. & Vaughan, C. K. Insights into centromere dna bending revealed by the cryo-em structure of the core centromere binding factor 3 with ndc10. Cell Rep. 24, 744–754 (2018).
Tachiwana, H. et al. Crystal structure of the human centromeric nucleosome containing CENP-A. Nature 476, 232–235 (2011).
Kingston, I. J., Yung, J. S. & Singleton, M. R. Biophysical characterization of the centromere-specific nucleosome from budding yeast. J. Biol. Chem. 286, 4021–4026 (2011).
Espelin, C. W., Simons, K. T., Harrison, S. C. & Sorger, P. K. Binding of the essential Saccharomyces cerevisiae kinetochore protein Ndc10p to CDEII. Mol. Biol. Cell. 14, 4557–4568 (2003).
Morey, L., Barnes, K., Chen, Y., Fitzgerald-Hayes, M. & Baker, R. E. The histone fold domain of Cse4 is sufficient for CEN targeting and propagation of active centromeres in budding yeast. Eukaryot. Cell 3, 1533–1543 (2004).
Henikoff, S. et al. The budding yeast centromere DNA Element II wraps a stable Cse4 hemisome in either orientation in vivo. eLife 3, e01861 (2014).
Nechemia-Arbely, Y. et al. Human centromeric CENP-A chromatin is a homotypic, octameric nucleosome at all cell cycle points. J. Cell Biol. 216, 607–621 (2017).
Henikoff, S. & Henikoff, J. G. “Point” centromeres of Saccharomyces harbor single centromere-specific nucleosomes. Genetics 190, 1575–1577 (2012).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 6, e21856 (2017).
Lawrimore, J., Bloom, K. S. & Salmon, E. D. Point centromeres contain more than a single centromere-specific Cse4 (CENP-A) nucleosome. J. Cell Biol. 195, 573–582 (2011).
Fitzgerald, D. J. et al. Multiprotein expression strategy for structural biology of eukaryotic complexes. Structure 15, 275–279 (2007).
Zhang, Z., Yang, J. & Barford, D. Recombinant expression and reconstitution of multiprotein complexes by the USER cloning method in the insect cell-baculovirus expression system. Methods 95, 13–25 (2016).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Fernandez-Leiro, R. & Scheres, S. H. W. A pipeline approach to single-particle processing in RELION. Acta Crystallogr. D Struct. Biol. 73, 496–502 (2017).
Elmlund, H., Elmlund, D. & Bengio, S. PRIME: probabilistic initial 3D model generation for single-particle cryo-electron microscopy. Structure 21, 1299–1306 (2013).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Schulman, B. A. et al. Insights into SCF ubiquitin ligases from the structure of the Skp1–Skp2 complex. Nature 408, 381–386 (2000).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Buchan, D. W., Minneci, F., Nugent, T. C., Bryson, K. & Jones, D. T. Scalable web services for the PSIPRED Protein Analysis Workbench. Nucleic Acids Res. 41, W349–W357 (2013).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Luger, K., Mader, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389, 251–260 (1997).
Chittori, S. et al. Structural mechanisms of centromeric nucleosome recognition by the kinetochore protein CENP-N. Science 359, 339–343 (2018).
Pentakota, S. et al. Decoding the centromeric nucleosome through CENP-N. eLife 6, e33442 (2017).
Bao, Y., White, C. L. & Luger, K. Nucleosome core particles containing a poly(dA.dT) sequence element exhibit a locally distorted DNA structure. J. Mol. Biol. 361, 617–624 (2006).
Acknowledgements
This work was funded by a MRC grant (no. MC_UP_1201/6) and a CR-UK grant (no. C576/A14109) to D.B. We thank S. Chen, G. Cannone and G. McMullan for help with EM data collection, J. Grimmett and T. Darling for computing and J. Shi for help with insect cell expression.
Author information
Authors and Affiliations
Contributions
D.B. directed the project. Z.Z. cloned all constructs. J.Y and Z.Z. purified complexes. Z.Z. performed the mitotic stability assay. J.Y. and S.M. performed SEC-MALS experiments. K.Y. prepared EM grids, collected and analyzed EM data. K.Y. and D.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Biochemical characterization of CBF3−CEN3 complexes.
a, Left, Coomassie blue–stained SDS–PAGE of CBF3−CEN3 complex. Right, ethidium bromide staining of the same gel. Below, SEC chromatogram of (i) CBF3−CEN3, (ii) CEN3 and (iii) CBF3. b, Left, Coomassie blue–stained SDS–PAGE of CBF3core−Ndc10DBD−CEN3 complex. Right, ethidium bromide staining of the same gel. Below, SEC chromatogram of (i) CBF3core−Ndc10DBD−CEN3, (ii) CEN3 and (iii) CBF3core−Ndc10DBD. c, Mutations of I76 and Y79 of Ctf13 disrupt interactions between CBF3core and Ndc10. The experiments shown were performed in triplicate with similar results.
Supplementary Figure 2 Cryo-EM analysis and resolution of CBF3−CEN3 complex.
a, A typical cryo-EM micrograph of CBF3–CEN3 representative of 3,537 micrographs. b, Gallery of 2D class averages of CBF3−CEN3 complex showing different views representative of 100 2D classes. c, Angular distribution plot of CBF3−CEN3 particles. d, Workflow of 3D classification of CBF3−CEN3 showing a 4.4-Å map of dimeric CBF3−CEN3 complex and a 3.0-Å map of CBF3core−Ndc10DBD (lower right) (masked EM density). Red boxes, dimeric CBF3−CEN3; black box, CBF3core; blue box, monomeric CBF3−CEN3 (CBF3core and Ndc10 dimer); black box, CBF3core. The dashed-line red box shows different DNA conformations. A HEAT map representing the structural variation of the three 3D classes of the dimeric CBF3–CEN3 complex is shown in Supplementary Fig. 4c. e, Local-resolution map of multi-body segment CBF3core of CBF3A. f, Local-resolution map of multi-body segment CBF3core of CBF3B. g, Local-resolution map of multi-body segment of Ndc10DBD−CEN3−DNA. h, Cryo-EM density of the 4.4-Å CBF3−CEN3 complex (left) and local-resolution map calculated with RESMAP 56 (right). i, Fourier shell correlation (FSC) curves are shown for the cryo-EM reconstructions of CBF3–CEN3 and three multi-body segments CBF3core-A, CBF3core-B and Ndc10DBD−CEN3−DNA. j, Fourier shell correlation (FSC) curves are shown for CBF3msk (gold-standard FSC, two half maps, and merged map). The resolution at FSC = 0.5 for the model–map comparison is indicated. k, Representative EM densities of regions of the 3.0-Å map of CBF3msk.
Supplementary Figure 3 Cryo-EM analysis and resolution of CBF3core−Ndc10DBD−CEN3 complex.
a, A typical cryo-EM micrograph of CBF3core−Ndc10DBD−CEN3 complex representative of 1,171 micrographs. b, A typical cryo-EM micrograph of CBF3core–Ndc10DBD representative of 724 micrographs. These two datasets were combined for subsequent processing steps to determine 3D reconstructions of DNA-free CBF3core and CBF3core–Ndc10DBD at 3.9 Å and 3.6 Å, respectively. c, Gallery of 2D class averages of CBF3core–Ndc10DBD showing different views representative of 100 2D classes. d, Angular distribution plot of CBF3core–Ndc10DBD particles. e, Workflow of 3D classification of combined datasets from CBF3core–Ndc10DBD and CBF3core–Ndc10DBD–CEN3. Orange boxes, DNA-free CBF3core–Ndc10DBD; black boxes, CBF3core; green box, CBF3core–Ndc10DBD–CEN3 DNA complex; red boxes, dimeric CBF3core–Ndc10DBD–CEN3 complex; blue box, monomeric CBF3core–Ndc10DBD–CEN3 complex. A HEAT map representing the structural variation of the three 3D classes of DNA-free CBF3core–Ndc10DBD is shown in Supplementary Fig 4b. f, Cryo-EM density of the 3.6-Å CBF3core–Ndc10DBD complex (no DNA) (left) and local-resolution map calculated with RESMAP56 (right). g, Fourier shell correlation (FSC) curves are shown for the cryo-EM reconstructions CBF3core–Ndc10DBD and CBF3core.
Supplementary Figure 4 Structural variations of CBF3 complexes.
a, Overall structure of CBF3. b, Heat map of the r.m.s. deviation between the 3D classes DNA-free CBF3core–Ndc10DBD. Coordinates were superimposed on CBF3A. The DNA-binding module of Ndc10DBD is the most structurally variable. c, Two orthogonal views showing a HEAT map of the r.m.s. deviation between the 3D classes of the dimeric CBF3–CEN3 DNA complex.
Supplementary Figure 5 Mitotic stability assay of wild-type and mutant Ctf13.
a, Agar plate culture of yeast strains: wild type and mutants in Ctf13, the CCG motif (to AGC) of the CEN3 CDEIII or a combination of Ctf13 and CEN3 mutations. Cultures were incubated for strains harboring mini-chromosome (WT or CEN3 CDEIII CCG mutation) in combination with CTF13 (WT or I76/Y79 mutation) and grown 48 h for about 20 generations of nonselective growth for mini-chromosome. Single colonies were generated on nonselective media and further streaked onto selective media. The percentage of grown colonies on selective media was determined for mitotic transmission of mini-chromosomes. b, Table showing the percentage of colony growth on selective media (% mean survival ± 1 s.d.). Differences in mitotic stability between pairs of strains were analyzed using paired t tests. There was no significant difference between the CTF13I76R/Y79R and CEN3-CDEIIICCG/AGC mutants. The data are based on five replicates.
Supplementary Figure 6 The Cep3 subunits differ in conformation in the CBF3−CEN3 complex and schematic of the modeled CBF3−Cse4−CEN3 complex.
a, EM density map from the CBF3–CEN3 DNA complex showing the visible C termini of the two Ndc10 subunits of the Ndc10DBD dimer in close proximity. b, Superimposition of Cep3A and Cep3B of CBF3A from the CBF3–CEN3 DNA complex. CEN3 DNA as bound to Cep3A of CBF3A is shown. The Zn2Cys6 cluster and αMN helix of Cep3A interact with the major grooves of the CCG and TGT motifs of CDEIII, respectively. The major differences between Cep3A and Cep3B involve these structural elements. c, Schematic of Fig. 4b showing the modeled CBF3–Cse4–CEN3 complex. The Cse4 nucleosome is depicted with left-handed chirality. d, Cse4 nucleosome with right-handed chirality. In this instance, the Cse4 nucleosome clashes with CBF3A and therefore represents a less likely scenario. CBF3 is shown transparent.
Supplementary Figure 7 SEC-MALS analysis of CBF3−CEN3 complexes.
a, SEC MALS chromatogram for CBF3 complex without CEN3–DNA shows two species of molecular mass 888 kDa and 454 kDa, corresponding to dimeric CBF3 ([Cep3]2–Ctf13–Skp1–[Ndc10]2)2 (expected mass of 453 kDa) and monomeric CBF3 ([Cep3]2–Ctf13–Skp1–[Ndc10]2) (expected mass of 906 kDa). b, In the presence of CEN3 DNA, all CBF3 migrates as a single species of molecular mass 1,018 kDa corresponding to a dimeric CBF3–CEN3 complex ([Cep3]2–Ctf13–Skp1–[Ndc10]2)2–CEN3 (expected mass of 1,010 kDa). The experiments shown were performed in triplicate with similar results.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Table 1
Rights and permissions
About this article
Cite this article
Yan, K., Zhang, Z., Yang, J. et al. Architecture of the CBF3–centromere complex of the budding yeast kinetochore. Nat Struct Mol Biol 25, 1103–1110 (2018). https://doi.org/10.1038/s41594-018-0154-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0154-1
This article is cited by
-
Structural and dynamic mechanisms of CBF3-guided centromeric nucleosome formation
Nature Communications (2021)
-
Cell-cycle phospho-regulation of the kinetochore
Current Genetics (2021)
-
Structure of the inner kinetochore CCAN complex assembled onto a centromeric nucleosome
Nature (2019)