Abstract
The function of actin is coupled to the nucleotide bound to its active site. ATP hydrolysis is activated during polymerization; a delay between hydrolysis and inorganic phosphate (Pi) release results in a gradient of ATP, ADP–Pi and ADP along actin filaments (F-actin). Actin-binding proteins can recognize F-actin’s nucleotide state, using it as a local ‘age’ tag. The underlying mechanism is complex and poorly understood. Here we report six high-resolution cryo-EM structures of F-actin from rabbit skeletal muscle in different nucleotide states. The structures reveal that actin polymerization repositions the proposed catalytic base, His161, closer to the γ-phosphate. Nucleotide hydrolysis and Pi release modulate the conformational ensemble at the periphery of the filament, thus resulting in open and closed states, which can be sensed by coronin-1B. The drug-like toxin jasplakinolide locks F-actin in an open state. Our results demonstrate in detail how ATP hydrolysis links to F-actin’s conformational dynamics and protein interaction.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Straub, F. B. & Feuer, G. Adenosine triphosphate, the functional group of actin. Kiserl. Orvostud. 2, 141–151 (1950).
Laki, K., Bowen, W. J. & Clark, A. The polymerization of proteins; adenosine triphosphate and the polymerization of actin. J. Gen. Physiol. 33, 437–443 (1950).
Combeau, C. & Carlier, M. F. Probing the mechanism of ATP hydrolysis on F-actin using vanadate and the structural analogs of phosphate BeF-3 and A1F-4. J. Biol. Chem. 263, 17429–17436 (1988).
Cai, L., Makhov, A. M. & Bear, J. E. F-actin binding is essential for coronin 1B function in vivo. J. Cell Sci. 120, 1779–1790 (2007).
Blanchoin, L. & Pollard, T. D. Mechanism of interaction of Acanthamoeba actophorin (ADF/Cofilin) with actin filaments. J. Biol. Chem. 274, 15538–15546 (1999).
Suarez, C. et al. Cofilin tunes the nucleotide state of actin filaments and severs at bare and decorated segment boundaries. Curr. Biol. 21, 862–868 (2011).
Cai, L., Marshall, T. W., Uetrecht, A. C., Schafer, D. A. & Bear, J. E. Coronin 1B coordinates Arp2/3 complex and cofilin activities at the leading edge. Cell 128, 915–929 (2007).
Pollard, T. D. & Borisy, G. G. Cellular motility driven by assembly and disassembly of actin filaments. Cell 112, 453–465 (2003).
Blanchoin, L., Pollard, T. D. & Mullins, R. D. Interactions of ADF/cofilin, Arp2/3 complex, capping protein and profilin in remodeling of branched actin filament networks. Curr. Biol. 10, 1273–1282 (2000).
Kudryashov, D. S. & Reisler, E. ATP and ADP actin states. Biopolymers 99, 245–256 (2013).
Oztug Durer, Z. A., Diraviyam, K., Sept, D., Kudryashov, D. S. & Reisler, E. F-actin structure destabilization and DNase I binding loop: fluctuations mutational cross-linking and electron microscopy analysis of loop states and effects on F-actin. J. Mol. Biol. 395, 544–557 (2010).
Mannherz, H. G., Brehme, H. & Lamp, U. Depolymerisation of F-actin to G-actin and its repolymerisation in the presence of analogs of adenosine triphosphate. Eur. J. Biochem. 60, 109–116 (1975).
Graceffa, P. & Dominguez, R. Crystal structure of monomeric actin in the ATP state: structural basis of nucleotide-dependent actin dynamics. J. Biol. Chem. 278, 34172–34180 (2003).
Cooke, R. The role of the bound nucleotide in the polymerization of actin. Biochemistry 14, 3250–3256 (1975).
Courtemanche, N. & Pollard, T. D. Interaction of profilin with the barbed end of actin filaments. Biochemistry 52, 6456–6466 (2013). 34, 8960–8972.
Fisher, A.J. et al. X-ray structures of the myosin motor domain of Dictyostelium discoideum complexed with MgADP·BeFx and MgADP·AlF4. Biochemistry (1995).
Gulick, A. M., Bauer, C. B., Thoden, J. B. & Rayment, I. X-ray structures of the MgADP, MgATPgammaS, and MgAMPPNP complexes of the Dictyostelium discoideum myosin motor domain. Biochemistry 36, 11619–11628 (1997).
Rould, M. A., Wan, Q., Joel, P. B., Lowey, S. & Trybus, K. M. Crystal structures of expressed nonpolymerizable monomeric actin in the ADP and ATP states. J. Biol. Chem. 281, 31909–31919 (2006).
Vorobiev, S. et al. The structure of nonvertebrate actin: implications for the ATP hydrolytic mechanism. Proc. Natl. Acad. Sci. USA 100, 5760–5765 (2003).
McCullagh, M., Saunders, M. G. & Voth, G. A. Unraveling the mystery of ATP hydrolysis in actin filaments. J. Am. Chem. Soc. 136, 13053–13058 (2014).
Cooke, R. & Murdoch, L. Interaction of actin with analogs of adenosine triphosphate. Biochemistry 12, 3927–3932 (1973).
Nolen, B. J. & Pollard, T. D. Insights into the influence of nucleotides on actin family proteins from seven structures of Arp2/3 complex. Mol. Cell 26, 449–457 (2007).
Murakami, K. et al. Structural basis for actin assembly, activation of ATP hydrolysis, and delayed phosphate release. Cell 143, 275–287 (2010).
Crews, P., Manes, L. V. & Boehler, M. Jasplakinolide, a cyclodepsipeptide from the marine sponge, SP. Tetrahedr. Lett. 27, 2797–2800 (1986).
Bubb, M. R., Spector, I., Beyer, B. B. & Fosen, K. M. Effects of jasplakinolide on the kinetics of actin polymerization: an explanation for certain in vivo observations. J. Biol. Chem. 275, 5163–5170 (2000).
Vig, A. et al. The effect of toxins on inorganic phosphate release during actin polymerization. Eur. Biophys. J. 40, 619–626 (2011).
Tannert, R. et al. Synthesis and structure-activity correlation of natural-product inspired cyclodepsipeptides stabilizing F-actin. J. Am. Chem. Soc. 132, 3063–3077 (2010).
Milroy, L.-G. et al. Selective chemical imaging of static actin in live cells. J. Am. Chem. Soc. 134, 8480–8486 (2012).
Lukinavičius, G. et al. Fluorogenic probes for live-cell imaging of the cytoskeleton. Nat. Methods 11, 731–733 (2014).
Pospich, S. et al. Near-atomic structure of jasplakinolide-stabilized malaria parasite F-actin reveals the structural basis of filament instability. Proc. Natl. Acad. Sci. USA https://doi.org/10.1073/pnas.1707506114 (2017).
Galkin, V. E., Orlova, A., Vos, M. R., Schröder, G. F. & Egelman, E. H. Near-atomic resolution for one state of F-actin. Structure 23, 173–182 (2015).
von der Ecken, J. et al. Structure of the F-actin–tropomyosin complex. Nature 519, 114–117 (2015).
von der Ecken, J., Heissler, S. M., Pathan-Chhatbar, S., Manstein, D. J. & Raunser, S. Cryo-EM structure of a human cytoplasmic actomyosin complex at near-atomic resolution. Nature 534, 724–728 (2016).
Otterbein, L. R., Graceffa, P. & Dominguez, R. The crystal structure of uncomplexed actin in the ADP state. Science 293, 708–711 (2001).
Zheng, X., Diraviyam, K. & Sept, D. Nucleotide effects on the structure and dynamics of actin. Biophys. J. 93, 1277–1283 (2007).
Isambert, H. et al. Flexibility of actin filaments derived from thermal fluctuations: effect of bound nucleotide, phalloidin, and muscle regulatory proteins. J. Biol. Chem. 270, 11437–11444 (1995).
Kardos, R. et al. The effect of jasplakinolide on the thermodynamic properties of ADP.BeF(x) bound actin filaments. Thermochim. Acta 463, 77–80 (2007).
Alushin, G. M. et al. High-resolution microtubule structures reveal the structural transitions in αβ-tubulin upon GTP hydrolysis. Cell 157, 1117–1129 (2014).
Bharat, T. A. M., Murshudov, G. N., Sachse, C. & Löwe, J. Structures of actin-like ParM filaments show architecture of plasmid-segregating spindles. Nature 523, 106–110 (2015).
Ge, P., Durer, Z. A. O., Kudryashov, D., Zhou, Z. H. & Reisler, E. Cryo-EM reveals different coronin binding modes for ADP– and ADP–BeFx actin filaments. Nat. Struct. Mol. Biol. 21, 1075–1081 (2014).
Oda, T., Iwasa, M., Aihara, T., Maéda, Y. & Narita, A. The nature of the globular- to fibrous-actin transition. Nature 457, 441–445 (2009).
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015).
Allegretti, M., Mills, D. J., McMullan, G., Kühlbrandt, W. & Vonck, J. Atomic model of the F420-reducing [NiFe] hydrogenase by electron cryo-microscopy using a direct electron detector. eLife 3, e01963 (2014).
Wriggers, W. & Schulten, K. Investigating a back door mechanism of actin phosphate release by steered molecular dynamics. Proteins 35, 262–273 (1999).
Bubb, M. R., Senderowicz, A. M., Sausville, E. A., Duncan, K. L. & Korn, E. D. Jasplakinolide, a cytotoxic natural product, induces actin polymerization and competitively inhibits the binding of phalloidin to F-actin. J. Biol. Chem. 269, 14869–14871 (1994).
Papp, G. et al. Conformational changes in actin filaments induced by formin binding to the barbed end. Biophys. J. 91, 2564–2572 (2006).
Strzelecka-Gołaszewska, H., Mossakowska, M., Woźniak, A., Moraczewska, J. & Nakayama, H. Long-range conformational effects of proteolytic removal of the last three residues of actin. Biochem. J. 307, 527–534 (1995).
Zimmermann, D., Santos, A., Kovar, D. R. & Rock, R. S. Actin age orchestrates myosin-5 and myosin-6 run lengths. Curr. Biol. 25, 2057–2062 (2015).
Mentes, A. et al. High-resolution cryo-EM structures of actin-bound myosin states reveal the mechanism of myosin force sensing. Proc. Natl. Acad. Sci. USA 115, 1292–1297 (2018).
Galkin, V. E. et al. Remodeling of actin filaments by ADF/cofilin proteins. Proc. Natl. Acad. Sci. USA 108, 20568–20572 (2011).
Muhlrad, A., Pavlov, D., Peyser, Y. M. & Reisler, E. Inorganic phosphate regulates the binding of cofilin to actin filaments. FEBS J. 273, 1488–1496 (2006).
Moriyama, K. & Yahara, I. The actin-severing activity of cofilin is exerted by the interplay of three distinct sites on cofilin and essential for cell viability. Biochem. J. 365, 147–155 (2002).
Kabsch, W., Mannherz, H. G., Suck, D., Pai, E. F. & Holmes, K. C. Atomic structure of the actin:DNase I complex. Nature 347, 37–44 (1990).
Aitken, C. E., Marshall, R. A. & Puglisi, J. D. An oxygen scavenging system for improvement of dye stability in single-molecule fluorescence experiments. Biophys. J. 94, 1826–1835 (2008).
Rasnik, I., McKinney, S. A. & Ha, T. Nonblinking and long-lasting single-molecule fluorescence imaging: it’s ProQuest. Nat. Methods 3, 891–893 (2006).
Bieling, P., Telley, I. A., Hentrich, C., Piehler, J. & Surrey, T. Fluorescence microscopy assays on chemically functionalized surfaces for quantitative imaging of microtubule, motor, and +TIP dynamics. Methods Cell Biol. 95, 555–580 (2010).
Pardee, J. D. & Spudich, J. A. Purification of muscle actin. Methods Enzymol. 85, 164–181 (1982).
Hansen, S. D., Zuchero, J. B. & Mullins, R. D. Cytoplasmic actin: purification and single molecule assembly assays. Methods Mol. Biol. 1046, 145–170 (2013).
Margossian, S. S. & Lowey, S. Preparation of myosin and its subfragments from rabbit skeletal muscle. Methods Enzymol. 85, 55–71 (1982).
Pollard, T. D. Myosin purification and characterization. Methods Cell Biol. 24, 333–371 (1982).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Behrmann, E. et al. Real-space processing of helical filaments in SPARX. J. Struct. Biol. 177, 302–313 (2012).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Sachse, C. et al. High-resolution electron microscopy of helical specimens: a fresh look at tobacco mosaic virus. J. Mol. Biol. 371, 812–835 (2007).
Moriya, T. et al. High-resolution single particle analysis from electron cryo-microscopy images using SPHIRE. J. Vis. Exp. e55448 (2017).
Song, Y. et al. High-resolution comparative modeling with RosettaCM. Structure 21, 1735–1742 (2013).
Trabuco, L. G., Villa, E., Mitra, K., Frank, J. & Schulten, K. Flexible fitting of atomic structures into electron microscopy maps using molecular dynamics. Structure 16, 673–683 (2008).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
DiMaio, F. et al. Atomic-accuracy models from 4.5-Å cryo-electron microscopy data with density-guided iterative local refinement. Nat. Methods 12, 361–365 (2015).
Fleishman, S. J. et al. RosettaScripts: a scripting language interface to the Rosetta macromolecular modeling suite. PLoS One 6, e20161 (2011).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Kortemme, T., Kim, D.E. & Baker, D. Computational alanine scanning of protein-protein interfaces. Sci. STKE 2004, pl2 (2004).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Appleton, B. A., Wu, P. & Wiesmann, C. The crystal structure of murine coronin-1: a regulator of actin cytoskeletal dynamics in lymphocytes. Structure 14, 87–96 (2006).
Eswar, N., Eramian, D., Webb, B., Shen, M.-Y. & Sali, A. Protein structure modeling with MODELLER. Methods Mol. Biol. 426, 145–159 (2008).
Acknowledgements
We thank O. Hofnagel and D. Prumbaum for assistance with data collection. We thank W. Linke and A. Unger (Ruhr-Universität Bochum, Germany) for providing us with muscle acetone powder. This work was supported by the Max Planck Society (to S.R.), the state of Thuringia (to H.-D.A., grant 43-5572-321-12040-12) and the European Council under the European Union’s Seventh Framework Programme (FP7/ 2007–2013) (grant 615984) (to S.R.). S.P. is supported as a fellow of Studienstiftung des deutschen Volkes.
Author information
Authors and Affiliations
Contributions
S.R. designed the project. F.M. and S.P. prepared and processed cryo-EM specimens, and built the atomic models. T.W. designed and implemented the filament autopicking tool. J.F. and P.B. designed and analyzed the TIRF microscopy experiments. J.F. produced recombinant coronin-1B and performed the TIRF microscopy experiments. F.K. and H.-D.A. synthesized JASP. F.M., S.P. and J.F. prepared figures and videos. F.M. and S.R. wrote the manuscript. All authors reviewed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated Supplementary Information
Supplementary Figure 1 Schematic representation of the modified single-particle analysis method used to obtain the F-actin models.
For each step, possible filament segment alignments are represented along with the development of the Fourier shell correlation curves and the maps with their corresponding distribution of angular assignments. For details see Methods.
Supplementary Figure 2 Overview of the cryo-EM data and resolution for all six datasets.
(a,b) Representative micrographs at ~ -1.5 μm defocus (a) and their power spectrum (b). (c) The local resolution of all reconstructions decreases gradually towards the ends of the filament, reflecting the intrinsic flexibility of F-actin. In addition, SD1 has systematically lower local resolution than the rest of the monomer, suggesting this area is flexible in the filament. The same color scale was used for all volumes. (d) Fourier shell correlation (FSC) curves for the masked (black) and unmasked (green) maps. The FSC plots for the unmasked maps clearly show that there is no over-refinement in the reconstructions. FSCs were calculated for a 120-Å-long central section of the filament. The red curves show the correlation for phase-randomized maps, proving that the masking has no effect on the estimated resolution.
Supplementary Figure 3 Density at the nucleotide-binding site.
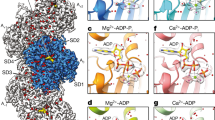
Stereo pair images for the nucleotide-binding site of F-actin in complex with AppNHp (a), ADP–BeFx (b), ADP–Pi (c), ADP–Pi JASP (d), ADP JASP (e), and ADP (f). The images show the density for the metal-nucleotide complex (yellow mesh) and protein (transparent gray). In addition, we included several key amino acids at the nucleotide-binding pocket. For further orientation and residue labeling please see Fig. 2.
Supplementary Figure 4 G-to F transition of AppNHp-actin.
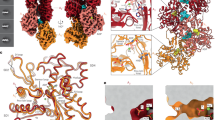
The superposition of the models of AppNHp-actin in the G- (blue, bound Ca2+, PDB 1NWK (Graceffa, P., and Dominguez, R., J Biol Chem. 278, 34172–34180, 2003)) and F-state (red, bound Mg2+) at the center illustrates the conformational change upon polymerization. Insets show the nucleotide-binding pockets of G-actin (left) and F-actin (right) including possible hydrogen bonds between the nucleotide and the protein, highlighted by green dashed lines. The interactions of the γ-phosphate with the protein backbone found in G-actin are probably preserved in the F-state. Both models are shown as side (upper panel) and top (lower panel) views (see also Supplementary Video 3).
Supplementary Figure 5 Evaluation of the nucleotide composition of the AppNHp filaments by reverse-phase ion-pair chromatography.
Before grid preparation, filaments were collected and resuspended in F-buffer without nucleotides. Samples were heat-denatured and loaded directly onto the column. Shaded regions represent peaks coming from AppNHp and AppNH2 (blue), ADP (turquoise), or the buffer (gray). The fractions of total nucleotides in the sample are indicated for each species, demonstrating that the AppNHp filaments contained a mixture of AppNHp and its hydrolysis products. Buffer control represents a run with F-buffer, necessary to identify the signal of the buffer components.
Supplementary Figure 6 Comparison between bare F-actin–ADP and its complex with tropomyosin.
(a) Superposition of the atomic models of F-actin–ADP (turquoise) and F-actin–ADP–tropomyosin (beige and green, PDB 5JLF, EMDB 8162 (von der Ecken, J., et al., Nature. 534, 724–728, 2016; von der Ecken, J., et al., Nature. 519, 114–117, 2015). The C terminal tails of all F-actin subunits are highlighted in red. (b, c) Close-up views of the C termini and corresponding densities in the absence (b) and presence (c) of tropomyosin, illustrating that tropomyosin binding results in the disordering of the C terminal tail.
Supplementary Figure 7 In silico alanine scanning for F-actin in the open and closed D loop conformations.
The interface was defined based on the central actin subunit of each model filament. (a) Predicted change of free energy upon alanine mutations in the closed (represented by F-actin–ADP, top, turquoise) and open (represented by F-actin–ADP–BeFx, bottom, orange) conformations of the intrastrand interface. The dotted lines indicate the threshold over which alanine mutations are considered destabilizing. (b) Predicted effect of the alanine mutations mapped in the filament structures of F-actin–ADP (closed state, top) and F-actin–ADP–BeFx (open state, bottom). In the open state, the effect is generally bigger than in the closed state, explaining the lower stability of F-actin–ADP. For guidance, regions forming the filament interfaces are highlighted and labeled, these include the D loop (a.a. 38 – 52), proline rich loop (P loop, a.a. 108 – 112), W loop (a.a. 165 – 172), plug (H plug, a.a. 263 – 273), and the C terminus. Subunits are shown as surfaces calculated from backbone and Cβ-atoms, except the central one, which is shown in ribbons. The color scale represents the predicted change of free energy from destabilizing (red) to stabilizing (blue).
Supplementary Figure 8 Proposed effect of the nucleotide state of actin on the binding of different actin-binding proteins.
(a-c) Molecular dynamics flexible fitting models of the complex between yeast coronin and F-actin. Ribbon representation of F-actin in complex with ADP–BeFx (red, a) or ADP states (green, b) complexed with coronin (blue). The insets show possible contacts between the D loop and coronin residues, illustrating that coronin could directly sense the changes occurring in this region. (c) To exclude a possible direct interaction of JASP with coronin, we superimposed the density found in the JASP complexes (yellow) with the ADP–BeFx filament. The results highlight the lack of overlap between the coronin and JASP binding sites. The molecular dynamics flexible fitting was performed using previously published electron density maps (EMDB accession codes 6101 and 6100, respectively (Ge, P., et al., Nat Struct Mol Biol. 21, 1075–1081, 2014.)). (d,e) Superposition between the F-actin–cofilin (gray/purple) complex (PDB 3J0S (Galkin, V.E., et al., Proc Nat Acad Sci. 108, 20568–20572, 2011)) and our ADP–BeFx (red, d) and ADP (green, e) models. The first 5 residues of cofilin, shown to be sensitive to the nucleotide state of F-actin (Moriyama, K., and Yahara, I., Biochem J. 365, 147–155, 2002) are highlighted in yellow. The figure illustrates how the N terminus of cofilin binds to the same pocket as the C terminus of actin in the F-actin–ADP–BeFx structure (d). In addition, the position of S3 is highlighted, which is known to be phosphorylated to regulate the function of cofilin.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Note
Supplementary Video 1
Agreement between models and maps. (a–f) Model-map agreement of the (a) F-actin–AppNHp, (b) F-actin–ADP–BeFx, (c) F-actin–ADP–Pi, (d) F-actin–ADP–Pi JASP, (e) F-actin–ADP JASP, and (f) F-actin–ADP reconstructions. For each map the same representative α-helical and β-sheet regions as well as the nucleotide-binding pocket are shown. Ribbons are colored according to the nucleotide state represented in the model: ATP-like: AppNHp/ADP–BeFx (blue), ADP–Pi-like: ADP–Pi/ADP–Pi JASP (red) and ADP-like: ADP/ADP JASP (green). The central subunits are colored in a lighter shade to highlight the dimensions of a single subunit. Maps are depicted at comparable thresholds.
Supplementary Video 2
Density at the ligand binding sites. (a–f) The movie shows the density for the ligand and key amino acids at the nucleotide-binding sites of F-actin–AppNHp (a), F-actin–ADP–BeFx (b), F-actin–ADP–Pi (c), F-actin–ADP–Pi JASP (d), F-actin–ADP JASP (e), and F-actin–ADP (f). For F-actin–ADP–BeFx, the density corresponding to BeFx is highlighted in magenta. In addition, the density for JASP and key binding residues are also shown for F-actin–ADP–Pi JASP (d) and F-actin–ADP JASP (e).
Supplementary Video 3
G- to F-actin transition at the active center of AppNHp-bound actin. (a) Whole monomer conformational change. Residues Q137, D154, and H161 as well as the AppNHp and Mg2+ are shown in balls and sticks. Only residues present in the G- (PDB-ID: 1NWK) and F-actin model can be included in the morph, therefore residues 40 – 51 (D loop) and 372-375 (C terminus) are not shown. (b) Close-up view of the conformational change. The density corresponds to the AppNHp map. The morph demonstrates that the new conformation of His161 is supported by the data. (c) Overall fit of the atomic model to the sharpened cryo-EM density.
Supplementary Video 4
Intrastrand interactions at the D loop/C terminus interface. (a–f) Visualization of side chains and their corresponding densities at the D loop/C terminus interface of the (a) F-actin–AppNHp, (b) F-actin–ADP–BeFx, (c) F-actin–ADP–Pi, (d) F-actin–ADP–Pi JASP, (e) F-actin–ADP JASP, and (f) F-actin–ADP reconstructions. The video highlights the different intrastrand interactions in association to the nucleotide states. All maps were filtered to their local resolution. The interface is located near the center of the reconstruction, with the central subunit shown in a lighter shade.
Supplementary Video 5
Mixed conformations at the D loop/C terminus interface of the F-actin–AppNHp, ADP–BeFx and ADP–Pi map. (a) The sharpened map of F-actin–AppNHp (gray) is superimposed with the corresponding atomic model (shades of blue). (b) Close-up view of the D loop and C terminus interface of F-actin–AppNHp map filtered to 4.1 Å. The D loop is not included in the model, as it is disordered when the map is filtered to the nominal resolution. The density of the C terminus splits, resembling two distinct conformations. Superimposition of the F-actin–AppNHp map (gray) with the model and map of F-actin–ADP (close D loop state, shades of green, c) and F-actin–ADP–Pi JASP (open D loop state, shades of red, d) illustrating a mixture of conformations at the D loop/C terminus interface of F-actin–AppNHp. (e–h) and (i–l) show the same views as in (a–d) but for F-actin–ADP–BeFx (shades of blue) and F-actin–ADP–Pi (shades of red), respectively. Maps shown in (b–d, f–h and j–l) were filtered to 4.1 Å, corresponding to the nominal resolution of the F-actin–ADP map, and are shown at lower thresholds to visualize conformations with low occupancy.
Supplementary Video 6
Principal component analysis of the different actin structures. The movie shows the variation of an F-actin monomer projected on either the first or the second principal component. The dotted line in the plot (left) illustrates the position of the structure (right) along the corresponding component. For clarity, two views are provided for each projection. We used only Cα atoms present in all six structures for the analysis, so part of the D loop (a.a. 45-49) and the C terminus (a.a. 372-375) are not included in the movie. The actin monomer is colored according to its subdomain structure; SD1 is cyan, SD2 green, SD3 orange, and SD4 red.
Supplementary Video 7
Conformational change at the D-loop/C-terminus interface. (a) The sharpened map of F-actin–ADP–Pi JASP (gray) is superimposed with the corresponding atomic model (shades of red) with JASP highlighted in yellow. (b) Close-up view of the D loop and C terminal interface of F-actin–ADP–Pi JASP. (c) Morph (gray) of atomic models illustrating the conformational change of the D loop and C terminus from the open to the closed state. (d) Close-up view of the F-actin–ADP map (gray) and the corresponding model (shades of green). (e) Overview of the complete sharpened F-actin–ADP map and model. Maps shown in (b,d) were filtered to their local resolution.
Rights and permissions
About this article
Cite this article
Merino, F., Pospich, S., Funk, J. et al. Structural transitions of F-actin upon ATP hydrolysis at near-atomic resolution revealed by cryo-EM. Nat Struct Mol Biol 25, 528–537 (2018). https://doi.org/10.1038/s41594-018-0074-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0074-0
This article is cited by
-
Characterizing ATP processing by the AAA+ protein p97 at the atomic level
Nature Chemistry (2024)
-
Distinct inter-domain interactions of dimeric versus monomeric α-catenin link cell junctions to filaments
Communications Biology (2023)
-
Molecular mechanisms of inorganic-phosphate release from the core and barbed end of actin filaments
Nature Structural & Molecular Biology (2023)
-
Cyclophosphamide treatment modifies the thermal stability of profilin bound monomeric and leiomodin2 bound filamentous actin
Journal of Thermal Analysis and Calorimetry (2023)
-
Magic angle spinning NMR structure of human cofilin-2 assembled on actin filaments reveals isoform-specific conformation and binding mode
Nature Communications (2022)