Abstract
Activating mutations in KRAS are among the most common tumor driver mutations. Until recently, KRAS had been considered undruggable with small molecules; the discovery of the covalent KRASG12C inhibitors ARS-853 and ARS-1620 has demonstrated that it is feasible to inhibit KRAS with high potency in cells and animals. Although the biological activity of these inhibitors has been described, the biochemical mechanism of how the compounds achieve potent inhibition remained incompletely understood. We now show that the activity of ARS-853 and ARS-1620 is primarily driven by KRAS-mediated catalysis of the chemical reaction with Cys12 in human KRASG12C, while the reversible binding affinity is weak, in the hundreds of micromolar or higher range. The mechanism resolves how an induced, shallow and dynamic pocket not expected to support high-affinity binding of small molecules can nevertheless be targeted with potent inhibitors and may be applicable to other targets conventionally considered undruggable.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Hunter, J. C. et al. Biochemical and structural analysis of common cancer-associated KRAS mutations. Mol. Cancer Res. 13, 1325–1335 (2015).
Ostrem, J. M. & Shokat, K. M. Direct small-molecule inhibitors of KRAS: from structural insights to mechanism-based design. Nat. Rev. Drug Discov. 15, 771–785 (2016).
Downward, J. Targeting RAS signalling pathways in cancer therapy. Nat. Rev. Cancer 3, 11–22 (2003).
Downward, J. RAS’s cloak of invincibility slips at last? Cancer Cell 25, 5–6 (2014).
van Hattum, H. & Waldmann, H. Chemical biology tools for regulating RAS signaling complexity in space and time. Chem. Biol. 21, 1185–1195 (2014).
Papke, B. & Der, C. J. Drugging RAS: know the enemy. Science 355, 1158–1163 (2017).
Wang, W., Fang, G. & Rudolph, J. Ras inhibition via direct Ras binding—is there a path forward? Bioorg. Med. Chem. Lett. 22, 5766–5776 (2012).
Mao, Y., Yao, H., Wang, H., Cheng, P. & Long, D. Microsecond timescale dynamics of GDP-bound Ras underlies the formation of novel inhibitor-binding pockets. Angew. Chem. Int. Edn Engl. 55, 15629–15632 (2016).
Müller, M. P., Jeganathan, S., Heidrich, A., Campos, J. & Goody, R. S. Nucleotide based covalent inhibitors of KRas can only be efficient in vivo if they bind reversibly with GTP-like affinity. Sci. Rep. 7, 3687 (2017).
Welsch, M. E. et al. Multivalent small-molecule pan-RAS inhibitors. Cell 168, 878–889.e29 (2017).
Ostrem, J. M., Peters, U., Sos, M. L., Wells, J. A. & Shokat, K. M. K-Ras(G12C) inhibitors allosterically control GTP affinity and effector interactions. Nature 503, 548–551 (2013).
Taveras, A. G. et al. Ras oncoprotein inhibitors: the discovery of potent, ras nucleotide exchange inhibitors and the structural determination of a drug-protein complex. Bioorg. Med. Chem. 5, 125–133 (1997).
Lito, P., Solomon, M., Li, L. S., Hansen, R. & Rosen, N. Allele-specific inhibitors inactivate mutant KRAS G12C by a trapping mechanism. Science 351, 604–608 (2016).
Patricelli, M. P. et al. Selective inhibition of oncogenic KRAS output with small molecules targeting the inactive state. Cancer Discov. 6, 316–329 (2016).
Janes, M. R. et al. Targeting KRAS mutant cancers with a covalent G12C-specific inhibitor. Cell 172, 578–589.e17 (2018).
Copeland, R.A. Evaluation of Enzyme Inhibitors in Drug Discovery: A Guide for Medicinal Chemists and Pharmacologists (Wiley, Hoboken, NJ, 2013).
Schwartz, P. A. et al. Covalent EGFR inhibitor analysis reveals importance of reversible interactions to potency and mechanisms of drug resistance. Proc. Natl Acad. Sci. USA 111, 173–178 (2014).
Hunter, J. C. et al. In situ selectivity profiling and crystal structure of SML-8-73-1, an active site inhibitor of oncogenic K-Ras G12C. Proc. Natl Acad. Sci. USA 111, 8895–8900 (2014).
Huynh, M. V. & Campbell, S. L. Getting a handle on RAS-targeted therapies: cysteine directed inhibitors. Mini Rev. Med. Chem. 16, 383–390 (2016).
Lu, J. et al. KRAS G12C drug development: discrimination between switch II pocket configurations using hydrogen/deuterium-exchange mass spectrometry. Structure 25, 1442–1448.e3 (2017).
Zeng, M. et al. Potent and selective covalent quinazoline inhibitors of KRAS G12C. Cell Chem Biol. 24, 1005–1016.e3 (2017).
Patricelli, M. P., Peters, U., Li, L., Ren, P. & Liu, Y. Method for screening inhibitors of Ras. US Patent 9, 810690 (2017).
Bandaru, P. et al. Deconstruction of the Ras switching cycle through saturation mutagenesis. eLife 6, e27810 (2017).
Isom, D. G., Castañeda, C. A., Cannon, B. R. & García-Moreno, B. Large shifts in pKa values of lysine residues buried inside a protein. Proc. Natl Acad. Sci. USA 108, 5260–5265 (2011).
Hagel, M. et al. First selective small molecule inhibitor of FGFR4 for the treatment of hepatocellular carcinomas with an activated FGFR4 signaling pathway. Cancer Discov. 5, 424–437 (2015).
Harling, J. D. et al. Discovery of novel irreversible inhibitors of interleukin (IL)-2-inducible tyrosine kinase (Itk) by targeting cysteine 442 in the ATP pocket. J. Biol. Chem. 288, 28195–28206 (2013).
Cross, D. A. et al. AZD9291, an irreversible EGFR TKI, overcomes T790M-mediated resistance to EGFR inhibitors in lung cancer. Cancer Discov. 4, 1046–1061 (2014).
Solca, F. et al. Target binding properties and cellular activity of afatinib (BIBW 2992), an irreversible ErbB family blocker. J. Pharmacol. Exp. Ther. 343, 342–350 (2012).
Gay, N. J. & Walker, J. E. Homology between human bladder carcinoma oncogene product and mitochondrial ATP-synthase. Nature 301, 262–264 (1983).
Walker, J. E., Saraste, M., Runswick, M. J. & Gay, N. J. Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J. 1, 945–951 (1982).
Pai, E. F. et al. Refined crystal structure of the triphosphate conformation of H-ras p21 at 1.35 A resolution: implications for the mechanism of GTP hydrolysis. EMBO J. 9, 2351–2359 (1990).
Privé, G. G. et al. X-ray crystal structures of transforming p21 ras mutants suggest a transition-state stabilization mechanism for GTP hydrolysis. Proc. Natl Acad. Sci. USA 89, 3649–3653 (1992).
Zhang, T. et al. Discovery of potent and selective covalent inhibitors of JNK. Chem. Biol. 19, 140–154 (2012).
Bulaj, G., Kortemme, T. & Goldenberg, D. P. Ionization-reactivity relationships for cysteine thiols in polypeptides. Biochemistry 37, 8965–8972 (1998).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
We thank J. Edwards, K. Shokat and D. Dhanak for critical reading of the manuscript and helpful discussions, A. Borum for technical assistance with chemistry and compound management and Shanghai Langtze Biomedical Technology for chemistry support on compound synthesis. Use of the IMCA-CAT beamline 17-ID at the Advanced Photon Source was supported by the companies of the Industrial Macromolecular Crystallography Association through a contract with Hauptman-Woodward Medical Research Institute. This research used resources of the Advanced Photon Source, a US Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357 and of the Advanced Light Source (beamline 8.2.2), which is a DOE Office of Science User Facility under contract no. DE-AC02-05CH11231. We would like to thank R. Alexander and J. C. Spurlino for acquiring the crystallographic dataset for ARS-917. The KRASG12C project is financially supported by Janssen Biotech, Inc. under a collaboration between Wellspring Biosciences, Inc. and Janssen Biotech, Inc.
Author information
Authors and Affiliations
Contributions
R.H. and U.P. contributed equally as co-first authors, and A.B. and Y.C. contributed equally as co-second authors. R.H. directed biochemical experiments, performed the MS and analyzed MS data. U.P. performed and coordinated the crystallography, solved the crystal structures, developed and performed the high protein concentration kinetic assays and expressed and purified recombinant KRAS proteins. A.B., Y.C. and U.P. performed the biochemical kinetics experiments and analyzed experimental data. J.F. synthesized compounds. L.-S. L. and P.R. designed compounds and directed the chemistry. M.R.J. directed experiments and edited the manuscript. Y.L. suggested the study, supervised the work and edited the manuscript. P.P.Z. conceptualized, designed and directed the study and analyzed data. R.H., U.P., L.-S. L. and P.P.Z. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
All authors are employees of Wellspring Biosciences, Inc. and shareholders of Araxes Pharma LLC, which holds the rights to the inhibitors characterized in the paper.
Additional information
Publisherʼs note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Time courses of KRASG12C covalent engagement.
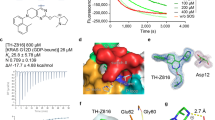
Data are shown for the three or four independent experiments for each compound used to calculate the values shown in Table 1 and in the main text and shown in Fig. 1. (a) ARS-853. (b) ARS-1620. For the experiments in (a) and (b) the KRASG12C concentration was 2 μM and the inhibitor concentration was varied from 8 to 64 μM, as shown. For each compound, the percent KRASG12C covalently labeled by inhibitor over time, as assessed by mass spectrometry, is shown at multiple compound concentrations, along with the kobs obtained by fitting each time course to a single exponential, with the percent engagement at time zero fixed at zero percent. (c) ARS-853 under conditions where the inhibitor concentration was held at 8 μM and the protein concentration varied from 20 to 320 μM. The percent compound covalently bound to KRASG12C over time, as assessed by the decrease in free compound concentration, is shown at multiple protein concentrations, along with kobs obtained by fitting each time course to a single exponential, with the percent engagement at time zero fixed at zero percent.
Supplementary Figure 2 Chemical structures of compound 12 and noncovalent analogs of ARS-1620 and ARS-853.
Chemical structures of compound 12, four analogs of ARS-1620 and one analog of ARS-853 in which the electrophile warhead has been modified or deleted such that the analogs cannot react covalently. The ARS-1620 analogs ARS-1372, ARS-1408, ARS-1440 and ARS-1448 were synthesized as 50:50 mixtures of atropisomers. For each of these, the isomer corresponding to the active isomer of ARS-1620 is as shown for ARS-1620 in Fig. 3.
Supplementary Figure 3 pKa determination.
(a) The pKa of cysteine 12 in native and denatured KRASG12C, cysteine in the model peptide RFAACAA and the KRASG12C-derived peptide VGACGVGKS. The rate constant kobs of reaction of 1 mM iodoacetamide with each cysteine substrate was determined at a range of pH values in three independent experiments for each substrate. Individual rate constants were normalized to the highest value measured in each experiment, and the mean relative kobs for each pH is plotted. The error bars represent the standard error of the normalized kobs values. The pKa values are shown in Table 2, and the individual normalized kobs values in Supplementary Table 1. (b) Engagement reaction of 8 μM ARS-1620 with KRASG12C at pH 7.5 and pH 10. Reaction progress was assessed by mass spectrometry as described in the Online Methods. This experiment was repeated independently two additional times, with similar results. (c) The pKa of cysteine 12 in native KRASG12C and cysteine in the RFAACAA peptide, determined by measuring the kobs of the reaction of 5 mM N-benzylacrylamide with KRASG12C cysteine 12, and 10 mM N-benzylacrylamide with the peptide across a range of pH values. kobs was normalized as described above. This experiment was performed once.
Supplementary Figure 4 kinact/Ki determination for ARS-107, ARS-917, and compound 12.
(a) The rate of covalent engagement was measured at multiple concentrations for each compound at 2 μM KRASG12C, and the rate constants plotted versus inhibitor concentration. The insert shows the plot for compound 12 with an adjusted y-axis range to facilitate visualization. (b) The rate of covalent engagement was measured for 8 μM ARS-107 at multiple concentrations of KRASG12C. The mean of kobs from three independent experiments is shown in black for each point, and individual replicate values in red. The error bars correspond to the standard error. Data were fit to a straight line with intercept fixed at the origin. kinact/Ki corresponds to the slope of the line.
Supplementary Figure 5 Determination of Ki and kinact for the interaction of ARS-853 with KRASD92C and at pH 9.5.
(a) Determination for KRASD92C in standard pH 7.5 conditions. (b) Determination for KRASG12C at pH 9.5. The rate of covalent engagement was measured at multiple concentrations of KRASD92C or KRASG12C, as indicated, at 8 μM ARS-853 in two independent experiments, and the observed rate constants plotted versus protein concentration. The data were fit to a one site binding function as in Fig. 1b. The values for Ki, kinact and kinact/Ki shown represent the averages of the values obtained in the two independent experiments, with the error encompassing the range of values observed.
Supplementary information
Rights and permissions
About this article
Cite this article
Hansen, R., Peters, U., Babbar, A. et al. The reactivity-driven biochemical mechanism of covalent KRASG12C inhibitors. Nat Struct Mol Biol 25, 454–462 (2018). https://doi.org/10.1038/s41594-018-0061-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0061-5
This article is cited by
-
Strain-release alkylation of Asp12 enables mutant selective targeting of K-Ras-G12D
Nature Chemical Biology (2024)
-
Real-time monitoring of the reaction of KRAS G12C mutant specific covalent inhibitor by in vitro and in-cell NMR spectroscopy
Scientific Reports (2023)
-
Hydrophobic interactions dominate the recognition of a KRAS G12V neoantigen
Nature Communications (2023)
-
Chemical acylation of an acquired serine suppresses oncogenic signaling of K-Ras(G12S)
Nature Chemical Biology (2022)
-
KRAS(G12D) can be targeted by potent inhibitors via formation of salt bridge
Cell Discovery (2022)