Abstract
Iron metabolism is critical for sustaining life and maintaining human health. Here, we find that iron homeostasis is linked to facultative heterochromatin assembly and regulation of gene expression during adaptive genome control. We show that the fission yeast Clr4/Suv39h histone methyltransferase is part of a rheostat-like mechanism in which transcriptional upregulation of mRNAs in response to environmental change provides feedback to prevent their uncontrolled expression through heterochromatin assembly. Interestingly, proper iron homeostasis is required, as iron depletion or downregulation of iron transporters causes defects in heterochromatin assembly and unrestrained upregulation of gene expression. Remarkably, an unbiased genetic screen revealed that restoration of iron homeostasis is sufficient to re-establish facultative heterochromatin and proper gene control genome-wide. These results establish a role for iron homeostasis in facultative heterochromatin assembly and reveal a dynamic mechanism for reprogramming the genome in response to environmental changes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Berger, S. L. & Sassone-Corsi, P. Metabolic signaling to chromatin. Cold Spring Harb. Perspect. Biol. 8, a019463 (2016).
Berry, S. & Dean, C. Environmental perception and epigenetic memory: mechanistic insight through FLC. Plant J. 83, 133–148 (2015).
Feng, S., Jacobsen, S. E. & Reik, W. Epigenetic reprogramming in plant and animal development. Science 330, 622–627 (2010).
Rando, O. J. & Simmons, R. A. I’m eating for two: parental dietary effects on offspring metabolism. Cell 161, 93–105 (2015).
Grewal, S. I. & Elgin, S. C. Transcription and RNA interference in the formation of heterochromatin. Nature 447, 399–406 (2007).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Trojer, P. & Reinberg, D. Facultative heterochromatin: is there a distinctive molecular signature? Mol. Cell 28, 1–13 (2007).
Wang, J., Jia, S. T. & Jia, S. New insights into the regulation of heterochromatin. Trends Genet. 32, 284–294 (2016).
Kilchert, C., Wittmann, S. & Vasiljeva, L. The regulation and functions of the nuclear RNA exosome complex. Nat. Rev. Mol. Cell Biol. 17, 227–239 (2016).
Lykke-Andersen, S. & Jensen, T. H. Nonsense-mediated mRNA decay: an intricate machinery that shapes transcriptomes. Nat. Rev. Mol. Cell Biol. 16, 665–677 (2015).
Rinn, J. L. & Chang, H. Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 81, 145–166 (2012).
Yamamoto, M. The selective elimination of messenger RNA underlies the mitosis-meiosis switch in fission yeast. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 86, 788–797 (2010).
Nakayama, J., Rice, J. C., Strahl, B. D., Allis, C. D. & Grewal, S. I. Role of histone H3 lysine 9 methylation in epigenetic control of heterochromatin assembly. Science 292, 110–113 (2001).
Zofall, M. et al. RNA elimination machinery targeting meiotic mRNAs promotes facultative heterochromatin formation. Science 335, 96–100 (2012).
Ekwall, K. Genome-wide analysis of HDAC function. Trends Genet. 21, 608–615 (2005).
Nicolas, E. et al. Distinct roles of HDAC complexes in promoter silencing, antisense suppression and DNA damage protection. Nat. Struct. Mol. Biol. 14, 372–380 (2007).
Lee, N. N. et al. Mtr4-like protein coordinates nuclear RNA processing for heterochromatin assembly and for telomere maintenance. Cell 155, 1061–1074 (2013).
Sugiyama, T. & Sugioka-Sugiyama, R. Red1 promotes the elimination of meiosis-specific mRNAs in vegetatively growing fission yeast. EMBO J. 30, 1027–1039 (2011).
Zhou, Y. et al. The fission yeast MTREC complex targets CUTs and unspliced pre-mRNAs to the nuclear exosome. Nat. Commun. 6, 7050 (2015).
Meola, N. et al. Identification of a nuclear exosome decay pathway for processed transcripts. Mol. Cell 64, 520–533 (2016).
Miller, J. E. & Reese, J. C. Ccr4-Not complex: the control freak of eukaryotic cells. Crit. Rev. Biochem. Mol. Biol. 47, 315–333 (2012).
Brönner, C., Salvi, L., Zocco, M., Ugolini, I. & Halic, M. Accumulation of RNA on chromatin disrupts heterochromatic silencing. Genome Res. 27, 1174–1183 (2017).
Folco, H. D. et al. Untimely expression of gametogenic genes in vegetative cells causes uniparental disomy. Nature 543, 126–130 (2017).
Hiriart, E. et al. Mmi1 RNA surveillance machinery directs RNAi complex RITS to specific meiotic genes in fission yeast. EMBO J. 31, 2296–2308 (2012).
Sugiyama, T. et al. Enhancer of rudimentary cooperates with conserved RNA-processing factors to promote meiotic mRNA decay and facultative heterochromatin assembly. Mol. Cell 61, 747–759 (2016).
Yamanaka, S. et al. RNAi triggered by specialized machinery silences developmental genes and retrotransposons. Nature 493, 557–560 (2013).
Volpe, T. A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002).
Cotobal, C. et al. Role of Ccr4-Not complex in heterochromatin formation at meiotic genes and subtelomeres in fission yeast. Epigenetics Chromatin 8, 28 (2015).
Shah, S., Wittmann, S., Kilchert, C. & Vasiljeva, L. lncRNA recruits RNAi and the exosome to dynamically regulate pho1 expression in response to phosphate levels in fission yeast. Genes Dev. 28, 231–244 (2014).
Touat-Todeschini, L. et al. Selective termination of lncRNA transcription promotes heterochromatin silencing and cell differentiation. EMBO J. 36, 2626–2641 (2017).
Joh, R. I. et al. Survival in quiescence requires the euchromatic deployment of Clr4/SUV39H by Argonaute-associated small RNAs. Mol. Cell 64, 1088–1101 (2016).
Hall, I. M. et al. Establishment and maintenance of a heterochromatin domain. Science 297, 2232–2237 (2002).
Tashiro, S., Asano, T., Kanoh, J. & Ishikawa, F. Transcription-induced chromatin association of RNA surveillance factors mediates facultative heterochromatin formation in fission yeast. Genes Cells 18, 327–339 (2013).
Chen, D. et al. Global transcriptional responses of fission yeast to environmental stress. Mol. Biol. Cell 14, 214–229 (2003).
Chalamcharla, V. R., Folco, H. D., Dhakshnamoorthy, J. & Grewal, S. I. Conserved factor Dhp1/Rat1/Xrn2 triggers premature transcription termination and nucleates heterochromatin to promote gene silencing. Proc. Natl Acad. Sci. USA 112, 15548–15555 (2015).
Tucker, J. F. et al. A novel epigenetic silencing pathway involving the highly conserved 5′-3′ exoribonuclease Dhp1/Rat1/Xrn2 in Schizosaccharomyces pombe. PLoS Genet. 12, e1005873 (2016).
Labbé, S., Khan, M. G. & Jacques, J. F. Iron uptake and regulation in Schizosaccharomyces pombe. Curr. Opin. Microbiol. 16, 669–676 (2013).
Pelletier, B., Beaudoin, J., Mukai, Y. & Labbé, S. Fep1, an iron sensor regulating iron transporter gene expression in Schizosaccharomyces pombe. J. Biol. Chem. 277, 22950–22958 (2002).
Pelletier, B., Beaudoin, J., Philpott, C. C. & Labbé, S. Fep1 represses expression of the fission yeast Schizosaccharomyces pombe siderophore-iron transport system. Nucleic Acids Res. 31, 4332–4344 (2003).
Encinar del Dedo, J., Gabrielli, N., Carmona, M., Ayté, J. & Hidalgo, E. A cascade of iron-containing proteins governs the genetic iron starvation response to promote iron uptake and inhibit iron storage in fission yeast. PLoS Genet. 11, e1005106 (2015).
Fagerström-Billai, F. & Wright, A. P. Functional comparison of the Tup11 and Tup12 transcriptional corepressors in fission yeast. Mol. Cell. Biol. 25, 716–727 (2005).
Watson, A. D. et al. Ssn6-Tup1 interacts with class I histone deacetylases required for repression. Genes Dev. 14, 2737–2744 (2000).
Znaidi, S., Pelletier, B., Mukai, Y. & Labbé, S. The Schizosaccharomyces pombe corepressor Tup11 interacts with the iron-responsive transcription factor Fep1. J. Biol. Chem. 279, 9462–9474 (2004).
Grewal, S. I. & Jia, S. Heterochromatin revisited. Nat. Rev. Genet. 8, 35–46 (2007).
Churchman, L. S. & Weissman, J. S. Nascent transcript sequencing visualizes transcription at nucleotide resolution. Nature 469, 368–373 (2011).
Chen, E. S. et al. Cell cycle control of centromeric repeat transcription and heterochromatin assembly. Nature 451, 734–737 (2008).
Cam, H. P. et al. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat. Genet. 37, 809–819 (2005).
Wagschal, A. et al. Microprocessor, Setx, Xrn2, and Rrp6 co-operate to induce premature termination of transcription by RNAPII. Cell 150, 1147–1157 (2012).
Horn, P. J., Bastie, J. N. & Peterson, C. L. A. A Rik1-associated, cullin-dependent E3 ubiquitin ligase is essential for heterochromatin formation. Genes Dev. 19, 1705–1714 (2005).
Jia, S., Kobayashi, R. & Grewal, S. I. Ubiquitin ligase component Cul4 associates with Clr4 histone methyltransferase to assemble heterochromatin. Nat. Cell Biol. 7, 1007–1013 (2005).
Hong, E. J., Villén, J., Gerace, E. L., Gygi, S. P. & Moazed, D. A cullin E3 ubiquitin ligase complex associates with Rik1 and the Clr4 histone H3-K9 methyltransferase and is required for RNAi-mediated heterochromatin formation. RNA Biol. 2, 106–111 (2005).
Greil, F. et al. Distinct HP1 and Su(var)3-9 complexes bind to sets of developmentally coexpressed genes depending on chromosomal location. Genes Dev. 17, 2825–2838 (2003).
Piacentini, L., Fanti, L., Berloco, M., Perrini, B. & Pimpinelli, S. Heterochromatin protein 1 (HP1) is associated with induced gene expression in Drosophila euchromatin. J. Cell Biol. 161, 707–714 (2003).
Vakoc, C. R., Mandat, S. A., Olenchock, B. A. & Blobel, G. A. Histone H3 lysine 9 methylation and HP1gamma are associated with transcription elongation through mammalian chromatin. Mol. Cell 19, 381–391 (2005).
Tsukada, Y. et al. Histone demethylation by a family of JmjC domain-containing proteins. Nature 439, 811–816 (2006).
Kimura, S. & Suzuki, T. Iron-sulfur proteins responsible for RNA modifications. Biochim. Biophys. Acta 1853, 1272–1283 (2015).
Roignant, J. Y. & Soller, M. m(6)A in mRNA: an ancient mechanism for fine-tuning gene expression. Trends Genet. 33, 380–390 (2017).
Andrews, N. C. Disorders of iron metabolism. N. Engl. J. Med. 341, 1986–1995 (1999).
Hentze, M. W., Muckenthaler, M. U., Galy, B. & Camaschella, C. Two to tango: regulation of Mammalian iron metabolism. Cell 142, 24–38 (2010).
Moreno, S., Klar, A. & Nurse, P. Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol. 194, 795–823 (1991).
Wood, V. et al. The genome sequence of Schizosaccharomyces pombe. Nature 415, 871–880 (2002).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
R Development Core Team. A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, Austria, 2016).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, New York, 2009).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012).
Pierik, A. J., Netz, D. J. & Lill, R. Analysis of iron-sulfur protein maturation in eukaryotes. Nat. Protoc. 4, 753–766 (2009).
Churchman, L. S. & Weissman, J. S. Native elongating transcript sequencing (NET-seq). Curr. Protoc. Mol. Biol. 98, 14.4 (2012).
Acknowledgements
We thank E. Hidalgo (Universitat Pompeu Fabra, Spain) for yeast strains, J. Zhu, V. Bliskovsky and S. Shema for valuable technical advice, S. Holla for helpful contributions, J. Barrowman for editing the manuscript, and members of the Laboratory of Biochemistry and Molecular Biology, in particular the Grewal laboratory, for discussions. This study used the Helix Systems and Biowulf Linux cluster at the National Institutes of Health. This work was supported by the Intramural Research Program of the National Institutes of Health, National Cancer Institute.
Author information
Authors and Affiliations
Contributions
S.I.S.G. and P.S.G. conceived and supervised the project. P.S.G., M.L., J.D., V.B., H.X., and C.W. performed experiments and analyzed data. R.C., G.T., and D.W. performed bioinformatics analyses of genomic datasets. S.I.S.G. and P.S.G. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Current model showing the roles of RNA processing factors in the degradation of transcripts and facultative heterochromatin assembly.
RNA binding proteins engage a network of nuclear RNA processing factors such as MTREC and CCR4-NOT, which in turn act along with the termination factor Dhp1/Xrn2 to promote degradation of RNAs by the exosome and/or RNAi machinery. In addition to RNA degradation, these factors are believed to mediate recruitment of histone H3 lysine 9 methyltransferase Clr4 to promote the assembly of facultative heterochromatin islands.
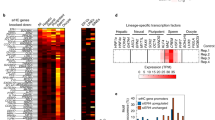
Supplementary Figure 2 Loci showing expression changes and formation of new heterochromatin islands in wild-type cells grown at 18 °C are enriched in stress response genes.
(Left) Histogram showing the numbers of genes in broad categories with increased expression (fold change value ≥ 1 and P value < 0.05) or decreased expression (fold change value ≤ 1 and P value < 0.05) in wild-type cells grown at 18 °C relative to wild-type cells grown at 30 °C as determined by RNA-seq (n = 2 independent experiments per condition). Genes with increased expression are shown in red and genes with decreased expression are shown in blue. Statistical significance (right-hand, single-tailed P value of binomial distribution, n=6380 transcripts) of the number of genes with increased expression is listed on the right. (Right) List of categories for genes within heterochromatin islands in wild-type cells grown at 18 °C as determined by ChIP-chip. Statistical significance (P value) of the number of genes within 18 °C heterochromatin islands is listed for each category.
Supplementary Figure 3 Clr6 histone deacetylase complex regulates expression of iron transporter genes.
(a) Area proportional Venn diagrams representing the numbers of genes with increased expression (fold change value ≥ 1 and P value < 0.05) in clr6-1 mutant cells grown at 30 °C and wild-type cells grown at 18 °C (top) or clr6-1 cells grown at 30 °C and clr4Δ cells grown at 18 °C (bottom). Statistical significance (right-hand, single-tailed P value of binomial distribution, n=6380 transcripts) of the overlap between the two groups is shown above. (b) Expression levels of six representative iron transporter genes determined by RNA-seq are shown in wild-type and clr6-1 cells grown at 30 °C. (c) Pst1-HA and Clr6-HA relative enrichment in cells grown at 30 °C was determined by ChIP-chip and is shown for iron transporter genes.
Supplementary Figure 4 Clr4 represses expression of meiotic genes at low temperature.
(a) Expression levels of four representative meiotic genes determined by RNA-seq are shown in wild-type and clr4Δ cells grown at 30 °C or 18 °C. (b) Wild-type and clr4Δ cells were grown at 18 °C in YEA rich medium and optical densities (ODs) of the cultures are shown as the mean ±s.d. (n = 2 independent experiments).
Supplementary Figure 5 RNAi is dispensable for the assembly of heterochromatin islands at 18 °C.
(a) Genome-wide H3K9me distribution at 18 °C in wild-type, ago1Δ, and ccr4Δ cells was determined by ChIP-chip and is plotted on all three chromosomes of S. pombe. New 18 °C facultative heterochromatin peaks are indicated. Wild-type ChIP-chip data is also presented in Fig. 2a and ccr4Δ ChIP-chip data is also presented in Fig. 6a. (b) H3K9me enrichments at 18 °C in wild-type and ago1Δ cells were determined by ChIP-qPCR. H3K9me fold enrichments at SPAPB1A11.02, SPBC428.10, and SPBC1289.14 relative to the control leu1 gene are shown as the mean +s.d. (n = 3 independent experiments). (c) H3K9me enrichments at 18 °C in wild-type, ccr4Δ, and red1Δ cells were determined by ChIP-qPCR. H3K9me fold enrichments at SPAPB1A11.02, SPBC428.10, and SPBC1289.14 relative to the control leu1 gene are shown as the mean +s.d. (n = 3 independent experiments). Wild-type ChIP-qPCR data is also presented in Supplementary Fig. 5b.
Supplementary Figure 6 Fep1 colocalizes with Ssn6 and Clr6 at various loci, including iron-transporter gene promoters, and its loss restores iron homeostasis in ccr4Δ cells.
(a) ChIP-chip analyses of Fep1, Ssn6 and Clr6 histone deacetylase complex-I subunit Pst1. Fep1-GFP, Ssn6-Myc, and Pst1-HA relative enrichment in cells grown at 30 °C. Iron transporter loci and additional peaks are indicated. (b) Fep1-GFP, Ssn6-Myc, and Pst1-HA relative enrichments are shown for representative iron transporter genes. (c) Mutation in fep1 in ccr4Δ cells causes up-regulation of iron transporter genes that are otherwise defective in their expression at 18 °C. Expression levels of frp1, str1, and str3 relative to act1 in the indicated strains grown at 18 °C were determined by RT-qPCR and are shown as the mean +s.d. (n = 2 independent experiments). Expression levels are normalized to wild-type cells grown at 30 °C. (d) Iron-55 uptake in wild-type, ccr4Δ, and ccr4Δ fep1Δ cells grown at 18 °C. (e) Mutation in ssn6 restores facultative heterochromatin assembly in ccr4Δ cells grown at low temperature. H3K9me enrichments at 18 °C in wild-type, ccr4Δ, and ccr4Δ ssn6-1 cells were determined by ChIP-qPCR. H3K9me fold enrichments at indicated loci relative to the control leu1 gene are shown as the mean +s.d. (n = 3 independent experiments).
Supplementary Figure 7 Iron affects gene regulation and the assembly of facultative heterochromatin islands at low temperature.
(a) Heat map of fold change values relative to wild-type 30 °C cells for up-regulated transcripts in indicated cultures grown at 18 °C. Clusters are grouped according to the analysis presented in Fig. 1f. (b) H3K9me enrichments in wild-type cells untreated or treated with 250µM Dip and grown at 18 °C for 72 hours (upper) or 24 hours (lower). H3K9me fold enrichments at indicated loci were determined by ChIP-qPCR and are shown relative to the control leu1 gene. Enrichments for the 72 hour treatments are shown as the mean +s.d. (n = 3 independent experiments). (c) H3K9me enrichments at 18 °C in ccr4Δ fep1-1 cells untreated or treated with 250µM Dip were determined by ChIP-qPCR. H3K9me fold enrichments at indicated loci are shown relative to the control leu1 gene.
Supplementary Figure 8 Iron depletion causes defects in RNA processing similar to those observed in the mtl1 mutant cells.
(a) Schematic of cryptic introns detected using RNA-seq in wild-type cells untreated or treated with 250µM Dip, and mtl1-1 mutant cells grown at 30 °C. The arcs below or above the line represent intron junction reads that map to the bottom or top DNA strands. (b) Hierarchical clustering of the indicated strains based on Pearson's correlation coefficients determined using RNA-seq (log2 fold change versus wild-type cells grown at 30 °C). Pairwise comparisons were performed (n = 6380 transcripts per comparison) and Pearson's correlation coefficients were converted into a color gradient. RNA-seq data for Dip-treated wild-type and clr4Δ cells grown at 18 °C and 30 °C were compared to various RNA processing mutants such as RNAi components (ago1Δ and dcr1Δ), nuclear RNA elimination factors (mmi1Δ, erh1Δ, red1Δ and mtl1-1), and CCR4-NOT complex (ccr4Δ) cultured at 30 °C. cwf10-1 splicing factor mutant was cultured at 26 °C. Also included is Clr6 HDAC mutant (clr6-1) grown at 30 °C that shows de-repression of genes affected by clr4Δ at 18 °C. Notice the high correlation between RNA processing mutant mtl1-1 and cells depleted for iron or lacking Clr4. (c) Density plots comparing transcripts (n = 6380) in mtl1-1 cells grown at 30 °C and wild-type cells treated with 250 µM Dip and grown at 30 °C (upper) or 18 °C (lower). Pearson's correlation coefficients (r) and the P values of the linear regressions are indicated. Source Data for Supplementary Fig. 8a are available with the paper online.
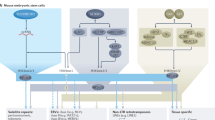
Supplementary Figure 9 Iron homeostasis is important for epigenetic genome control.
(Upper) Model showing regulation of iron transporter genes by Fep1, Ssn6-Tup11/12 and Clr6 histone deacetylase complex. In the presence of Fep1, Ssn6-Tup11/12 and Clr6 complex localize to iron transporter loci and regulate gene expression (left panel). In fep1 mutant cells, Ssn6 and likely Clr6 complex are de-localized, allowing for increased expression of iron transporter genes (right panel). The thickness of the arrows indicates expression levels of iron transporter genes. In addition to iron transporters, Clr6 regulates other loci at 30 °C that are regulated by Clr4 at 18 °C. (Lower) Model showing the requirement for intracellular iron in facultative heterochromatin assembly and proper gene regulation in adaptive genome control. In cells grown under suboptimal growth conditions, Clr4 is recruited to genomic locations in a transcription- and RNA-dependent mechanism involving RNA processing and termination factors, such as cleavage factors (CF) and Dhp1/Xrn2, for facultative heterochromatin assembly and serves as a rheostat to prevent hyper-elevation of transcripts (right panel). Cells depleted of intracellular iron show defects in facultative heterochromatin assembly and display aberrant gene expression in response to changing environmental growth conditions (left panel).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9
Supplementary Table 1
List of strains and oligos used in this study
Supplementary Dataset 1
Uncropped gel image
Rights and permissions
About this article
Cite this article
Gallagher, P.S., Larkin, M., Thillainadesan, G. et al. Iron homeostasis regulates facultative heterochromatin assembly in adaptive genome control. Nat Struct Mol Biol 25, 372–383 (2018). https://doi.org/10.1038/s41594-018-0056-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0056-2