Abstract
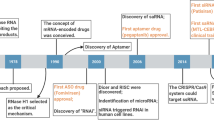
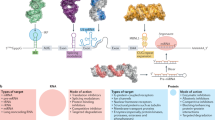
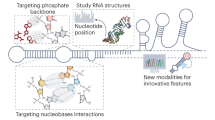
A recent revolution in RNA biology has led to the identification of new RNA classes with unanticipated functions, new types of RNA modifications, an unexpected multiplicity of alternative transcripts and widespread transcription of extragenic regions. This development in basic RNA biology has spawned a corresponding revolution in RNA-based strategies to generate new types of therapeutics. Here, I review RNA-based drug design and discuss barriers to broader applications and possible ways to overcome them. Because they target nucleic acids rather than proteins, RNA-based drugs promise to greatly extend the domain of ‘druggable’ targets beyond what can be achieved with small molecules and biologics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Debbie Maizels/Springer Nature

Debbie Maizels/Springer Nature

Debbie Maizels/Springer Nature
Similar content being viewed by others
References
Cech, T. R. & Steitz, J. A. The noncoding RNA revolution—trashing old rules to forge new ones. Cell 157, 77–94 (2014).
Deveson, I. W., Hardwick, S. A., Mercer, T. R. & Mattick, J. S. The dimensions, dynamics, and relevance of the mammalian noncoding transcriptome. Trends Genet. 33, 464–478 (2017).
Schonrock, N., Jonkhout, N. & Mattick, J. S. Seq and you will find. Curr. Gene Ther. 16, 220–229 (2016).
Brockdorff, N. et al. The product of the mouse Xist gene is a 15 kb inactive X-specific transcript containing no conserved ORF and located in the nucleus. Cell 71, 515–526 (1992).
Brown, C. J. et al. The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 71, 527–542 (1992).
Lee, R. C., Feinbaum, R. L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 (1993).
Wightman, B., Ha, I. & Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75, 855–862 (1993).
Ebert, M. S. & Sharp, P. A. Roles for microRNAs in conferring robustness to biological processes. Cell 149, 515–524 (2012).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Thomas, M., Lieberman, J. & Lal, A. Desperately seeking microRNA targets. Nat. Struct. Mol. Biol. 17, 1169–1174 (2010).
Volpe, T. A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002).
Verdel, A. et al. RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303, 672–676 (2004).
Watanabe, T. & Lin, H. Posttranscriptional regulation of gene expression by Piwi proteins and piRNAs. Mol. Cell 56, 18–27 (2014).
Holoch, D. & Moazed, D. RNA-mediated epigenetic regulation of gene expression. Nat. Rev. Genet. 16, 71–84 (2015).
Iwasaki, Y. W., Siomi, M. C. & Siomi, H. PIWI-interacting RNA: its biogenesis and functions. Annu. Rev. Biochem. 84, 405–433 (2015).
Janowski, B. A. et al. Activating gene expression in mammalian cells with promoter-targeted duplex RNAs. Nat. Chem. Biol. 3, 166–173 (2007).
Li, L. C. et al. Small dsRNAs induce transcriptional activation in human cells. Proc. Natl Acad. Sci. USA 103, 17337–17342 (2006).
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature 435, 834–838 (2005).
Breaker, R. R. Riboswitches and the RNA world. Cold Spring Harb. Perspect. Biol. 4, a003566 (2012).
Gelinas, A. D., Davies, D. R. & Janjic, N. Embracing proteins: structural themes in aptamer–protein complexes. Curr. Opin. Struct. Biol. 36, 122–132 (2016).
Sharp, P. A. On the origin of RNA splicing and introns. Cell 42, 397–400 (1985).
Gilbert, W. Origin of life: the RNA world. Nature 319, 618 (1986).
Holbrook, S. R. RNA structure: the long and the short of it. Curr. Opin. Struct. Biol. 15, 302–308 (2005).
Ellington, A. D. & Szostak, J. W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990).
Serganov, A. & Nudler, E. A decade of riboswitches. Cell 152, 17–24 (2013).
Quinn, J. J. & Chang, H. Y. Unique features of long non-coding RNA biogenesis and function. Nat. Rev. Genet. 17, 47–62 (2016).
Chen, K., Zhao, B. S. & He, C. Nucleic acid modifications in regulation of gene expression. Cell Chem. Biol. 23, 74–85 (2016).
Kopp, F. & Mendell, J. T. Functional classification and experimental dissection of long noncoding RNAs. Cell 172, 393–407 (2018).
Zamecnik, P. C. & Stephenson, M. L. Inhibition of Rous sarcoma virus replication and cell transformation by a specific oligodeoxynucleotide. Proc. Natl Acad. Sci. USA 75, 280–284 (1978).
Stephenson, M. L. & Zamecnik, P. C. Inhibition of Rous sarcoma viral RNA translation by a specific oligodeoxyribonucleotide. Proc. Natl Acad. Sci. USA 75, 285–288 (1978).
Dixon, S. J. & Stockwell, B. R. Identifying druggable disease-modifying gene products. Curr. Opin. Chem. Biol. 13, 549–555 (2009).
Fauman, E. B., Rai, B. K. & Huang, E. S. Structure-based druggability assessment—identifying suitable targets for small molecule therapeutics. Curr. Opin. Chem. Biol. 15, 463–468 (2011).
Cirak, S. et al. Exon skipping and dystrophin restoration in patients with Duchenne muscular dystrophy after systemic phosphorodiamidate morpholino oligomer treatment: an open-label, phase 2, dose-escalation study. Lancet 378, 595–605 (2011).
Finkel, R. S. et al. Nusinersen versus sham control in infantile-onset spinal muscular atrophy. N. Engl. J. Med. 377, 1723–1732 (2017).
Gorski, S. A., Vogel, J. & Doudna, J. A. RNA-based recognition and targeting: sowing the seeds of specificity. Nat. Rev. Mol. Cell Biol. 18, 215–228 (2017).
Kole, R., Krainer, A. R. & Altman, S. RNA therapeutics: beyond RNA interference and antisense oligonucleotides. Nat. Rev. Drug Discov. 11, 125–140 (2012).
Bennett, C. F., Baker, B. F., Pham, N., Swayze, E. & Geary, R. S. Pharmacology of antisense drugs. Annu. Rev. Pharmacol. Toxicol. 57, 81–105 (2017).
Crooke, S. T. Molecular mechanisms of antisense oligonucleotides. Nucleic Acid Ther. 27, 70–77 (2017).
Crooke, S. T., Wang, S., Vickers, T. A., Shen, W. & Liang, X. H. Cellular uptake and trafficking of antisense oligonucleotides. Nat. Biotechnol. 35, 230–237 (2017).
Khvorova, A. & Watts, J. K. The chemical evolution of oligonucleotide therapies of clinical utility. Nat. Biotechnol. 35, 238–248 (2017).
Raal, F. J. et al. Mipomersen, an apolipoprotein B synthesis inhibitor, for lowering of LDL cholesterol concentrations in patients with homozygous familial hypercholesterolaemia: a randomised, double-blind, placebo-controlled trial. Lancet 375, 998–1006 (2010).
Liang, X. H. et al. Translation efficiency of mRNAs is increased by antisense oligonucleotides targeting upstream open reading frames. Nat. Biotechnol. 34, 875–880 (2016).
Krützfeldt, J. et al. Silencing of microRNAs in vivo with ‘antagomirs’. Nature 438, 685–689 (2005).
Janssen, H. L. et al. Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 368, 1685–1694 (2013).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Wittrup, A. & Lieberman, J. Knocking down disease: a progress report on siRNA therapeutics. Nat. Rev. Genet. 16, 543–552 (2015).
Elbashir, S. M. et al. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494–498 (2001).
Lum, L. et al. Identification of Hedgehog pathway components by RNAi in Drosophila cultured cells. Science 299, 2039–2045 (2003).
Caplen, N. J., Fleenor, J., Fire, A. & Morgan, R. A. dsRNA-mediated gene silencing in cultured Drosophila cells: a tissue culture model for the analysis of RNA interference. Gene 252, 95–105 (2000).
Fraser, A. G. et al. Functional genomic analysis of C. elegans chromosome I by systematic RNA interference. Nature 408, 325–330 (2000).
Boutros, M. et al. Genome-wide RNAi analysis of growth and viability in Drosophila cells. Science 303, 832–835 (2004).
Luo, J. et al. A genome-wide RNAi screen identifies multiple synthetic lethal interactions with the Ras oncogene. Cell 137, 835–848 (2009).
Silva, J. M. et al. Profiling essential genes in human mammary cells by multiplex RNAi screening. Science 319, 617–620 (2008).
Schlabach, M. R. et al. Cancer proliferation gene discovery through functional genomics. Science 319, 620–624 (2008).
Brass, A. L. et al. Identification of host proteins required for HIV infection through a functional genomic screen. Science 319, 921–926 (2008).
Paddison, P. J. et al. A resource for large-scale RNA-interference-based screens in mammals. Nature 428, 427–431 (2004).
Novina, C. D. et al. siRNA-directed inhibition of HIV-1 infection. Nat. Med. 8, 681–686 (2002).
Jacque, J. M., Triques, K. & Stevenson, M. Modulation of HIV-1 replication by RNA interference. Nature 418, 435–438 (2002).
Martínez, M. A. et al. Suppression of chemokine receptor expression by RNA interference allows for inhibition of HIV-1 replication. AIDS 16, 2385–2390 (2002).
Park, W. S. et al. Prevention of HIV-1 infection in human peripheral blood mononuclear cells by specific RNA interference. Nucleic Acids Res. 30, 4830–4835 (2002).
Surabhi, R. M. & Gaynor, R. B. RNA interference directed against viral and cellular targets inhibits human immunodeficiency virus type 1 replication. J. Virol. 76, 12963–12973 (2002).
Song, E. et al. RNA interference targeting Fas protects mice from fulminant hepatitis. Nat. Med. 9, 347–351 (2003).
Morrissey, D. V. et al. Potent and persistent in vivo anti-HBV activity of chemically modified siRNAs. Nat. Biotechnol. 23, 1002–1007 (2005).
Judge, A. D. et al. Sequence-dependent stimulation of the mammalian innate immune response by synthetic siRNA. Nat. Biotechnol. 23, 457–462 (2005).
Jackson, A. L. et al. Position-specific chemical modification of siRNAs reduces “off-target” transcript silencing. RNA 12, 1197–1205 (2006).
Gilleron, J. et al. Image-based analysis of lipid nanoparticle-mediated siRNA delivery, intracellular trafficking and endosomal escape. Nat. Biotechnol. 31, 638–646 (2013).
Wittrup, A. et al. Visualizing lipid-formulated siRNA release from endosomes and target gene knockdown. Nat. Biotechnol. 33, 870–876 (2015).
Fitzgerald, K. et al. A highly durable RNAi therapeutic inhibitor of PCSK9. N. Engl. J. Med. 376, 41–51 (2017).
Nair, J. K. et al. Multivalent N-acetylgalactosamine-conjugated siRNA localizes in hepatocytes and elicits robust RNAi-mediated gene silencing. J. Am. Chem. Soc. 136, 16958–16961 (2014).
Rajeev, K. G. et al. Hepatocyte-specific delivery of siRNAs conjugated to novel non-nucleosidic trivalent N-acetylgalactosamine elicits robust gene silencing in vivo. ChemBioChem 16, 903–908 (2015).
Matsuda, S. et al. siRNA conjugates carrying sequentially assembled trivalent N-acetylgalactosamine linked through nucleosides elicit robust gene silencing in vivo in hepatocytes. ACS Chem. Biol. 10, 1181–1187 (2015).
Biessen, E. A. et al. Design of a targeted peptide nucleic acid prodrug to inhibit hepatic human microsomal triglyceride transfer protein expression in hepatocytes. Bioconjug. Chem. 13, 295–302 (2002).
Zimmermann, T. S. et al. Clinical proof of concept for a novel hepatocyte-targeting GalNAc-siRNA conjugate. Mol. Ther. 25, 71–78 (2017).
Garber, K. Worth the RISC? Nat. Biotechnol. 35, 198–202 (2017).
Pasi, K. J. et al. Targeting of antithrombin in hemophilia A or B with RNAi therapy. N. Engl. J. Med. 377, 819–828 (2017).
Yasuda, M. et al. RNAi-mediated silencing of hepatic Alas1 effectively prevents and treats the induced acute attacks in acute intermittent porphyria mice. Proc. Natl Acad. Sci. USA 111, 7777–7782 (2014).
Dutta, C. et al. Inhibition of glycolate oxidase with Dicer-substrate siRNA reduces calcium oxalate deposition in a mouse model of primary hyperoxaluria type 1. Mol. Ther. 24, 770–778 (2016).
Liebow, A. et al. An investigational RNAi therapeutic targeting glycolate oxidase reduces oxalate production in models of primary hyperoxaluria. J. Am. Soc. Nephrol. 28, 494–503 (2017).
Huang, Y. Preclinical and clinical advances of GalNAc-decorated nucleic acid therapeutics. Mol. Ther. Nucleic Acids 6, 116–132 (2017).
Prakash, T. P. et al. Targeted delivery of antisense oligonucleotides to hepatocytes using triantennary N-acetyl galactosamine improves potency 10-fold in mice. Nucleic Acids Res. 42, 8796–8807 (2014).
Tanowitz, M. et al. Asialoglycoprotein receptor 1 mediates productive uptake of N-acetylgalactosamine-conjugated and unconjugated phosphorothioate antisense oligonucleotides into liver hepatocytes. Nucleic Acids Res. 45, 12388–12400 (2017).
Dassie, J. P. et al. Systemic administration of optimized aptamer-siRNA chimeras promotes regression of PSMA-expressing tumors. Nat. Biotechnol. 27, 839–846 (2009).
McNamara, J. O. II et al. Cell type-specific delivery of siRNAs with aptamer-siRNA chimeras. Nat. Biotechnol. 24, 1005–1015 (2006).
Zhou, J. & Rossi, J. Aptamers as targeted therapeutics: current potential and challenges. Nat. Rev. Drug Discov. 16, 181–202 (2017).
Kortylewski, M. et al. In vivo delivery of siRNA to immune cells by conjugation to a TLR9 agonist enhances antitumor immune responses. Nat. Biotechnol. 27, 925–932 (2009).
Peer, D., Zhu, P., Carman, C. V., Lieberman, J. & Shimaoka, M. Selective gene silencing in activated leukocytes by targeting siRNAs to the integrin lymphocyte function-associated antigen-1. Proc. Natl Acad. Sci. USA 104, 4095–4100 (2007).
Song, E. et al. Antibody mediated in vivo delivery of small interfering RNAs via cell-surface receptors. Nat. Biotechnol. 23, 709–717 (2005).
Peer, D., Park, E. J., Morishita, Y., Carman, C. V. & Shimaoka, M. Systemic leukocyte-directed siRNA delivery revealing cyclin D1 as an anti-inflammatory target. Science 319, 627–630 (2008).
Ramishetti, S., Landesman-Milo, D. & Peer, D. Advances in RNAi therapeutic delivery to leukocytes using lipid nanoparticles. J. Drug Target. 24, 780–786 (2016).
Weinstein, S. et al. Harnessing RNAi-based nanomedicines for therapeutic gene silencing in B-cell malignancies. Proc. Natl Acad. Sci. USA 113, E16–E22 (2016).
Nikan, M. et al. Docosahexaenoic acid conjugation enhances distribution and safety of siRNA upon local administration in mouse brain. Mol. Ther. Nucleic Acids 5, e344 (2016).
Kedmi, R. et al. A modular platform for targeted RNAi therapeutics. Nat. Nanotechnol. 13, 214–219 (2018).
Gilboa-Geffen, A. et al. Gene knockdown by EpCAM aptamer-siRNA chimeras suppresses epithelial breast cancers and their tumor-initiating cells. Mol. Cancer Ther. 14, 2279–2291 (2015).
Esposito, C. L. et al. Multifunctional aptamer-miRNA conjugates for targeted cancer therapy. Mol. Ther. 22, 1151–1163 (2014).
Warren, L. et al. Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7, 618–630 (2010).
Sasaki, S. & Guo, S. Nucleic acid therapies for cystic fibrosis. Nucleic Acid Ther. 28, 1–9 (2018).
Yin, H. et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat. Biotechnol. 32, 551–553 (2014).
Nelson, C. E. et al. In vivo genome editing improves muscle function in a mouse model of Duchenne muscular dystrophy. Science 351, 403–407 (2016).
Tabebordbar, M. et al. In vivo gene editing in dystrophic mouse muscle and muscle stem cells. Science 351, 407–411 (2016).
Komor, A. C., Badran, A. H. & Liu, D. R. CRISPR-based technologies for the manipulation of eukaryotic genomes. Cell 168, 20–36 (2017).
Fellmann, C., Gowen, B. G., Lin, P. C., Doudna, J. A. & Corn, J. E. Cornerstones of CRISPR–Cas in drug discovery and therapy. Nat. Rev. Drug Discov. 16, 89–100 (2017).
Dowdy, S. F. Overcoming cellular barriers for RNA therapeutics. Nat. Biotechnol. 35, 222–229 (2017).
Wang, H. X. et al. CRISPR/Cas9-based genome editing for disease modeling and therapy: challenges and opportunities for nonviral delivery. Chem. Rev. 117, 9874–9906 (2017).
Graham, M. J. et al. Antisense inhibition of proprotein convertase subtilisin/kexin type 9 reduces serum LDL in hyperlipidemic mice. J. Lipid Res. 48, 763–767 (2007).
Karatasakis, A. et al. Effect of PCSK9 inhibitors on clinical outcomes in patients with hypercholesterolemia: a meta-analysis of 35 randomized controlled trials. J. Am. Heart Assoc. 6, e006910 (2017).
Benson, M. D., Dasgupta, N. R., Rissing, S. M., Smith, J. & Feigenbaum, H. Safety and efficacy of a TTR specific antisense oligonucleotide in patients with transthyretin amyloid cardiomyopathy. Amyloid 24, 219–225 (2017).
Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602–607 (2011).
Marraffini, L. A. & Sontheimer, E. J. CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322, 1843–1845 (2008).
Barrangou, R. The roles of CRISPR–Cas systems in adaptive immunity and beyond. Curr. Opin. Immunol. 32, 36–41 (2015).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Cornu, T. I., Mussolino, C. & Cathomen, T. Refining strategies to translate genome editing to the clinic. Nat. Med. 23, 415–423 (2017).
Neelapu, S. S. et al. Axicabtagene ciloleucel CAR T-cell therapy in refractory large B-cell lymphoma. N. Engl. J. Med. 377, 2531–2544 (2017).
Maude, S. L. et al. Tisagenlecleucel in children and young adults with B-cell lymphoblastic leukemia. N. Engl. J. Med. 378, 439–448 (2018).
Russell, S. et al. Efficacy and safety of voretigene neparvovec (AAV2-hRPE65v2) in patients with RPE65-mediated inherited retinal dystrophy: a randomised, controlled, open-label, phase 3 trial. Lancet 390, 849–860 (2017).
Dunbar, C. E. et al. Gene therapy comes of age. Science 359, eaan4672 (2018).
Karikó, K. et al. Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol. Ther. 16, 1833–1840 (2008).
Weissman, D. et al. HIV gag mRNA transfection of dendritic cells (DC) delivers encoded antigen to MHC class I and II molecules, causes DC maturation, and induces a potent human in vitro primary immune response. J. Immunol. 165, 4710–4717 (2000).
Weissman, D. & Karikó, K. mRNA: fulfilling the promise of gene therapy. Mol. Ther. 3, 1416–1417 (2015).
Pardi, N., Hogan, M. J., Porter, F. W. & Weissman, D. mRNA vaccines—a new era in vaccinology. Nat. Rev. Drug Discov. https://dx.doi.org/10.1038/nrd.2017.243 (2018).
Kranz, L. M. et al. Systemic RNA delivery to dendritic cells exploits antiviral defence for cancer immunotherapy. Nature 534, 396–401 (2016).
Richner, J. M. et al. Modified mRNA vaccines protect against Zika virus infection. Cell 168, 1114–1125 (2017).
Pardi, N. et al. Zika virus protection by a single low-dose nucleoside-modified mRNA vaccination. Nature 543, 248–251 (2017).
Dassie, J. P. & Giangrande, P. H. Current progress on aptamer-targeted oligonucleotide therapeutics. Ther. Deliv. 4, 1527–1546 (2013).
Keefe, A. D., Pai, S. & Ellington, A. Aptamers as therapeutics. Nat. Rev. Drug Discov. 9, 537–550 (2010).
Gold, L., Walker, J. J., Wilcox, S. K. & Williams, S. Advances in human proteomics at high scale with the SOMAscan proteomics platform. N. Biotechnol. 29, 543–549 (2012).
Lollo, B., Steele, F. & Gold, L. Beyond antibodies: new affinity reagents to unlock the proteome. Proteomics 14, 638–644 (2014).
Lieberman, J. Manipulating the in vivo immune response by targeted gene knockdown. Curr. Opin. Immunol. 35, 63–72 (2015).
Acknowledgements
This work was supported by NIH CA13944.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
J.L. is on the Scientific Advisory Board of Alnylam Pharmaceuticals.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lieberman, J. Tapping the RNA world for therapeutics. Nat Struct Mol Biol 25, 357–364 (2018). https://doi.org/10.1038/s41594-018-0054-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0054-4
This article is cited by
-
Liquid crystalline inverted lipid phases encapsulating siRNA enhance lipid nanoparticle mediated transfection
Nature Communications (2024)
-
The role and regulation of Maf proteins in cancer
Biomarker Research (2023)
-
The lncRNA HOTAIR: a pleiotropic regulator of epithelial cell plasticity
Journal of Experimental & Clinical Cancer Research (2023)
-
Failure of cartilage regeneration: emerging hypotheses and related therapeutic strategies
Nature Reviews Rheumatology (2023)
-
RNA therapeutics: updates and future potential
Science China Life Sciences (2023)