Abstract
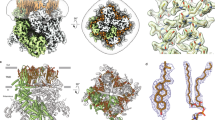
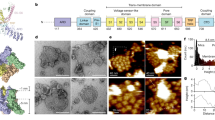
The transient receptor potential (TRP) channel TRPV4 participates in multiple biological processes, and numerous TRPV4 mutations underlie several distinct and devastating diseases. Here we present the cryo-EM structure of Xenopus tropicalis TRPV4 at 3.8-Å resolution. The ion-conduction pore contains an intracellular gate formed by the inner helices, but lacks any extracellular gate in the selectivity filter, as observed in other TRPV channels. Anomalous X-ray diffraction analyses identify a single ion-binding site in the selectivity filter, thus explaining TRPV4 nonselectivity. Structural comparisons with other TRP channels and distantly related voltage-gated cation channels reveal an unprecedented, unique packing interface between the voltage-sensor-like domain and the pore domain, suggesting distinct gating mechanisms. Moreover, our structure begins to provide mechanistic insights to the large set of pathogenic mutations, offering potential opportunities for drug development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Clapham, D. E. TRP channels as cellular sensors. Nature 426, 517–524 (2003).
Ramsey, I. S., Delling, M. & Clapham, D. E. An introduction to TRP channels. Annu. Rev. Physiol. 68, 619–647 (2006).
Venkatachalam, K. & Montell, C. TRP channels. Annu. Rev. Biochem. 76, 387–417 (2007).
Nilius, B. & Owsianik, G. The transient receptor potential family of ion channels. Genome Biol. 12, 218 (2011).
Julius, D. TRP channels and pain. Annu. Rev. Cell Dev. Biol. 29, 355–384 (2013).
Mizuno, A., Matsumoto, N., Imai, M. & Suzuki, M. Impaired osmotic sensation in mice lacking TRPV4. Am. J. Physiol. Cell Physiol. 285, C96–C101 (2003).
Liedtke, W. & Friedman, J. M. Abnormal osmotic regulation in trpv4-/- mice. Proc. Natl Acad. Sci. USA 100, 13698–13703 (2003).
Strotmann, R., Harteneck, C., Nunnenmacher, K., Schultz, G. & Plant, T. D. OTRPC4, a nonselective cation channel that confers sensitivity to extracellular osmolarity. Nat. Cell Biol. 2, 695–702 (2000).
Liedtke, W. et al. Vanilloid receptor-related osmotically activated channel (VR-OAC), a candidate vertebrate osmoreceptor. Cell 103, 525–535 (2000).
Wissenbach, U., Bödding, M., Freichel, M. & Flockerzi, V. Trp12, a novel Trp related protein from kidney. FEBS Lett. 485, 127–134 (2000).
Masuyama, R. et al. TRPV4-mediated calcium influx regulates terminal differentiation of osteoclasts. Cell Metab. 8, 257–265 (2008).
Sonkusare, S. K. et al. Elementary Ca2+ signals through endothelial TRPV4 channels regulate vascular function. Science 336, 597–601 (2012).
Ye, L. et al. TRPV4 is a regulator of adipose oxidative metabolism, inflammation, and energy homeostasis. Cell 151, 96–110 (2012).
Alessandri-Haber, N. et al. Hypotonicity induces TRPV4-mediated nociception in rat. Neuron 39, 497–511 (2003).
Alessandri-Haber, N. et al. Transient receptor potential vanilloid 4 is essential in chemotherapy-induced neuropathic pain in the rat. J. Neurosci. 24, 4444–4452 (2004).
Gao, X., Wu, L. & O’Neil, R. G. Temperature-modulated diversity of TRPV4 channel gating: activation by physical stresses and phorbol ester derivatives through protein kinase C-dependent and -independent pathways. J. Biol. Chem. 278, 27129–27137 (2003).
Köhler, R. et al. Evidence for a functional role of endothelial transient receptor potential V4 in shear stress-induced vasodilatation. Arterioscler. Thromb. Vasc. Biol. 26, 1495–1502 (2006).
Güler, A. D. et al. Heat-evoked activation of the ion channel, TRPV4. J. Neurosci. 22, 6408–6414 (2002).
Watanabe, H. et al. Heat-evoked activation of TRPV4 channels in a HEK293 cell expression system and in native mouse aorta endothelial cells. J. Biol. Chem. 277, 47044–47051 (2002).
Watanabe, H. et al. Activation of TRPV4 channels (hVRL-2/mTRP12) by phorbol derivatives. J. Biol. Chem. 277, 13569–13577 (2002).
Watanabe, H. et al. Anandamide and arachidonic acid use epoxyeicosatrienoic acids to activate TRPV4 channels. Nature 424, 434–438 (2003).
Nilius, B. & Voets, T. The puzzle of TRPV4 channelopathies. EMBO Rep. 14, 152–163 (2013).
White, J. P. M. et al. TRPV4: molecular conductor of a diverse orchestra. Physiol. Rev. 96, 911–973 (2016).
Liao, M., Cao, E., Julius, D. & Cheng, Y. Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504, 107–112 (2013).
Cao, E., Liao, M., Cheng, Y. & Julius, D. TRPV1 structures in distinct conformations reveal activation mechanisms. Nature 504, 113–118 (2013).
Gao, Y., Cao, E., Julius, D. & Cheng, Y. TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. Nature 534, 347–351 (2016).
Zubcevic, L. et al. Cryo-electron microscopy structure of the TRPV2 ion channel. Nat. Struct. Mol. Biol. 23, 180–186 (2016).
Huynh, K. W. et al. Structure of the full-length TRPV2 channel by cryo-EM. Nat. Commun. 7, 11130 (2016).
Saotome, K., Singh, A. K., Yelshanskaya, M. V. & Sobolevsky, A. I. Crystal structure of the epithelial calcium channel TRPV6. Nature 534, 506–511 (2016).
Singh, A. K., Saotome, K. & Sobolevsky, A. I. Swapping of transmembrane domains in the epithelial calcium channel TRPV6. Sci. Rep. 7, 10669 (2017).
Whicher, J. R. & MacKinnon, R. Structure of the voltage-gated K+ channel Eag1 reveals an alternative voltage sensing mechanism. Science 353, 664–669 (2016).
Tao, X., Hite, R. K. & MacKinnon, R. Cryo-EM structure of the open high-conductance Ca2+-activated K+ channel. Nature 541, 46–51 (2017).
Lee, C. H. & MacKinnon, R. Structures of the Human HCN1 Hyperpolarization-Activated Channel. Cell 168, 111–120.e11 (2017).
Inada, H., Procko, E., Sotomayor, M. & Gaudet, R. Structural and biochemical consequences of disease-causing mutations in the ankyrin repeat domain of the human TRPV4 channel. Biochemistry 51, 6195–6206 (2012).
Takahashi, N. et al. TRPV4 channel activity is modulated by direct interaction of the ankyrin domain to PI(4,5)P2. Nat. Commun. 5, 4994 (2014).
Shigematsu, H., Sokabe, T., Danev, R., Tominaga, M. & Nagayama, K. A 3.5-nm structure of rat TRPV4 cation channel revealed by Zernike phase-contrast cryoelectron microscopy. J. Biol. Chem. 285, 11210–11218 (2010).
Becker, D., Müller, M., Leuner, K. & Jendrach, M. The C-terminal domain of TRPV4 is essential for plasma membrane localization. Mol. Membr. Biol. 25, 139–151 (2008).
Lei, L. et al. A TRPV4 channel C-terminal folding recognition domain critical for trafficking and function. J. Biol. Chem. 288, 10427–10439 (2013).
Long, S. B., Campbell, E. B. & Mackinnon, R. Crystal structure of a mammalian voltage-dependent Shaker family K+ channel. Science 309, 897–903 (2005).
Long, S. B., Campbell, E. B. & Mackinnon, R. Voltage sensor of Kv1.2: structural basis of electromechanical coupling. Science 309, 903–908 (2005).
Long, S. B., Tao, X., Campbell, E. B. & MacKinnon, R. Atomic structure of a voltage-dependent K+ channel in a lipid membrane-like environment. Nature 450, 376–382 (2007).
Gregorio-Teruel, L. et al. The integrity of the TRP domain is pivotal for correct TRPV1 channel gating. Biophys. J. 109, 529–541 (2015).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360, 376 (1996).
Jordt, S.-E., Tominaga, M. & Julius, D. Acid potentiation of the capsaicin receptor determined by a key extracellular site. Proc. Natl Acad. Sci. USA 97, 8134–8139 (2000).
Cui, Y. et al. Selective disruption of high sensitivity heat activation but not capsaicin activation of TRPV1 channels by pore turret mutations. J. Gen. Physiol. 139, 273–283 (2012).
Ryu, S., Liu, B., Yao, J., Fu, Q. & Qin, F. Uncoupling proton activation of vanilloid receptor TRPV1. J. Neurosci. 27, 12797–12807 (2007).
Voets, T. et al. Molecular determinants of permeation through the cation channel TRPV4. J. Biol. Chem. 277, 33704–33710 (2002).
Shi, N., Ye, S., Alam, A., Chen, L. & Jiang, Y. Atomic structure of a Na+- and K+-conducting channel. Nature 440, 570–574 (2006).
Derebe, M. G. et al. Tuning the ion selectivity of tetrameric cation channels by changing the number of ion binding sites. Proc. Natl Acad. Sci. USA 108, 598–602 (2011).
Sauer, D. B., Zeng, W., Canty, J., Lam, Y. & Jiang, Y. Sodium and potassium competition in potassium-selective and non-selective channels. Nat. Commun. 4, 2721 (2013).
Lockless, S. W. Determinants of cation transport selectivity: equilibrium binding and transport kinetics. J. Gen. Physiol. 146, 3–13 (2015).
van der Cruijsen, E. A. W. et al. Importance of lipid-pore loop interface for potassium channel structure and function. Proc. Natl Acad. Sci. USA 110, 13008–13013 (2013).
Brohawn, S. G., Campbell, E. B. & MacKinnon, R. Physical mechanism for gating and mechanosensitivity of the human TRAAK K+ channel. Nature 516, 126–130 (2014).
Yang, H. et al. Pore architecture of TRIC channels and insights into their gating mechanism. Nature 538, 537–541 (2016).
Vriens, J., Owsianik, G., Janssens, A., Voets, T. & Nilius, B. Determinants of 4 alpha-phorbol sensitivity in transmembrane domains 3 and 4 of the cation channel TRPV4. J. Biol. Chem. 282, 12796–12803 (2007).
Lamandé, S. R. et al. Mutations in TRPV4 cause an inherited arthropathy of hands and feet. Nat. Genet. 43, 1142–1146 (2011).
Nishimura, G. et al. TRPV4-associated skeletal dysplasias. Am. J. Med. Genet. C. Semin. Med. Genet. 160C, 190–204 (2012).
Krakow, D. et al. Mutations in the gene encoding the calcium-permeable ion channel TRPV4 produce spondylometaphyseal dysplasia, Kozlowski type and metatropic dysplasia. Am. J. Hum. Genet. 84, 307–315 (2009).
Camacho, N. et al. Dominant TRPV4 mutations in nonlethal and lethal metatropic dysplasia. Am. J. Med. Genet. A. 152A, 1169–1177 (2010).
Dai, J. et al. Novel and recurrent TRPV4 mutations and their association with distinct phenotypes within the TRPV4 dysplasia family. J. Med. Genet. 47, 704–709 (2010).
Nishimura, G. et al. Spondylo-epiphyseal dysplasia, Maroteaux type (pseudo-Morquio syndrome type 2), and parastremmatic dysplasia are caused by TRPV4 mutations. Am. J. Med. Genet. A. 152A, 1443–1449 (2010).
Landouré, G. et al. Mutations in TRPV4 cause Charcot–Marie–Tooth disease type 2C. Nat. Genet. 42, 170–174 (2010).
Auer-Grumbach, M. et al. Alterations in the ankyrin domain of TRPV4 cause congenital distal SMA, scapuloperoneal SMA and HMSN2C. Nat. Genet. 42, 160–164 (2010).
Fiorillo, C. et al. TRPV4 mutations in children with congenital distal spinal muscular atrophy. Neurogenetics 13, 195–203 (2012).
Deng, H.-X. et al. Scapuloperoneal spinal muscular atrophy and CMT2C are allelic disorders caused by alterations in TRPV4. Nat. Genet. 42, 165–169 (2010).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Scheres, S. H. W. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Ludtke, S. J. Single-particle refinement and variability analysis in EMAN2.1. Methods Enzymol. 579, 159–189 (2016).
Lyumkis, D., Brilot, A. F., Theobald, D. L. & Grigorieff, N. Likelihood-based classification of cryo-EM images using FREALIGN. J. Struct. Biol. 183, 377–388 (2013).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Brown, A. et al. Tools for macromolecular model building and refinement into electron cryo-microscopy reconstructions. Acta Crystallogr. D Biol. Crystallogr. 71, 136–153 (2015).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A. J. Solving structures of protein complexes by molecular replacement with Phaser. Acta Crystallogr. D Biol. Crystallogr. 63, 32–41 (2007).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta. Crystallogr. D. Biol. Crystallogr. 53, 240–255 (1997).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Bond, C. S. & Schüttelkopf, A. W.ALINE: a WYSIWYG protein-sequence alignment editor for publication-quality alignments.Acta Crystallogr. D Biol. Crystallogr. 65, 510–512 (2009).
Schnorf, M., Potrykus, I. & Neuhaus, G. Microinjection technique: routine system for characterization of microcapillaries by bubble pressure measurement. Exp. Cell Res. 210, 260–267 (1994).
Cooper, P. E., Sala-Rabanal, M., Lee, S. J. & Nichols, C. G. Differential mechanisms of Cantú syndrome-associated gain of function mutations in the ABCC9 (SUR2) subunit of the KATP channel. J. Gen. Physiol. 146, 527–540 (2015).
Acknowledgements
We thank staff at APS beamlines 24-ID C/E, especially K. Rajashankar, K. Perry and N. Sukumar, for assistance at the synchrotron. This work used NE-CAT beamlines (GM103403), a Pilatus detector (RR029205) and an Eiger detector (OD021527) at the APS (DE-AC02-06CH11357). We thank the staff of the Sloan Kettering cryo-EM facility and Subangstrom LLC for assistance with cryo-EM data collection. This work was supported by startup funds from Washington University School of Medicine, the Mallinckrodt Foundation grant, American Heart Association Award 17SDG33400229, National Institutes of Health Grant R01NS099341 (all to P.Y.) and by startup funds from Memorial Sloan Kettering Cancer Center (to R.K.H.).
Author information
Authors and Affiliations
Contributions
Z.D. and P.Y. performed biochemical preparations and X-ray crystallography experiments. N.P. and R.K.H. conducted cryo-EM experiments and model building. G.M. and M.S.-R. conducted rubidium-flux and electrophysiology experiments. P.Y. designed and supervised the project. Z.D., G.M., C.G.N., R.K.H. and P.Y. analyzed the results and prepared the manuscript. All authors discussed the results and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Channel activation by GSK1016790A (GSK101).
Activation for the full-length wild type Xenopus TRPV4 (black filled squares, n = 4, mean ± SEM) and the truncated construct TRPV4cryst (grey filled squares, n = 4, mean ± SEM) in CosM6 cells by GSK101 at a, 0 nM, b, 40 nM, c, 200 nM and d, 1000 nM is evaluated by relative efflux of 86Rb+, in comparison with cells transfected with an empty vector (open squares, n = 4, mean ± SEM). e, Relative, apparent GSK101 activation constant as a function of GSK101 concentration, calculated from data presented in a-d. Wild type TRPV4 (filled circles) is more sensitive to GSK101 (~ 4- to 6-fold) than TRPV4cryst (open circles). Calculation and curve fitting are described in Methods.
Supplementary Figure 2 Ion permeation and blockage of Xenopus TRPV4 and TRPV4cryst.
a, Confocal images of CosM6 cells expressing full-length wild type Xenopus TRPV4 fused with EGFP (left panel) and the crystal construct TRPV4cryst fused with EGFP (right panel). TRPV4cryst shows altered expression pattern and is mainly localized to intracellular organelles. b, K+ and Cs+ conduction of wild type Xenopus TRPV4 and TRPV4cryst. Single-channel current-voltage dependence of TRPV4 in symmetrical 150 mM KCl (open circles, n = 8), in symmetrical 150 mM CsCl (open squares, n = 7), and TRPV4cryst in symmetrical 150 mM KCl (filled circles, n = 12), in symmetrical 150 mM CsCl (filled squares, n = 6). Currents were measured in inside-out excised patches upon application of GSK101 (10 nM for wild type TRPV4, 20 nM to 5 μM in KCl and 5 μM in CsCl for TRPV4cryst) from the cytoplasmic side. Wild type TRPV4 has a unitary conductance of 280 ± 11 pS and 191 ± 8 pS at −100 mV membrane potential in symmetrical 150 mM KCl and symmetrical 150 mM CsCl respectively. Under the same conditions, TRPV4cryst has a unitary conductance of 272 ± 13 pS and 185 ± 9 pS in symmetrical 150 mM KCl and symmetrical 150 mM CsCl respectively. c, Representative traces of activation of TRPV4cryst by GSK101 in symmetrical 150 mM KCl (top panel, 20 nM GSK101) and in 150 mM CsCl (bottom panel, 5 μM GSK101). Membrane potential is −120 mV. d, Ca2+ and Ba2+ permeation of wild type Xenopus TRPV4. Inside-out membrane patches were excised in symmetrical BaCl2 (75 mM) or CaCl2 (75 mM) buffers. Wild type TRPV4 currents (Ba2+, black; Ca2+, grey) were induced by perfusion of the bath (cytoplasmic side) with Kint containing 10 nM GSK101. Reversal potentials are 27 mV for BaCl2/KCl and 33 mV for CaCl2/KCl, respectively. After liquid junction potential correction, the corresponding permeability ratios were calculated to be PBa/PK = 3.4 (n = 8) and PCa/PK = 4.8 (n = 5). Basal currents in the absence of GSK101 were very low and only noticeable at extreme potentials (−100 and 100 mV). e, Gd3+ blockage of TRPV4cryst. GSK101 activation (1 μM) of TRPV4cryst (filled squares, n = 3, mean ± SEM) in CosM6 cells was assessed by relative efflux of 86Rb+. Basal 86Rb+ efflux was measured in cells without GSK101 stimulation (empty circles, n = 3, mean ± SEM). GSK101-induced 86Rb+ efflux was inhibited in the presence of 1 mM GdCl3 (empty squares, n = 3, mean ± SEM). f, GSK2193874 (GSK219) inhibition of Xenopus TRPV4 and TRPV4cryst. GSK101 activation (40 nM) of wild type TRPV4 (black diamonds, n = 3, mean ± SEM) and TRPV4cryst (black squares, n = 3, mean ± SEM) in CosM6 cells was assessed by relative efflux of 86Rb+. Basal 86Rb+ efflux was measured in cells transfected with empty vector and treated with GSK101 (40 nM) (black circles, n = 3, mean ± SEM). GSK219 (10 μM) partially inhibits GSK101-induced 86Rb+ effluxes for both wild type Xenopus TRPV4 (grey diamonds, n = 3, mean ± SEM) and TRPV4cryst (grey squares, n = 3, mean ± SEM).
Supplementary Figure 3 Cryo-EM reconstruction of Xenopus TRPV4cryst.
a, Flowchart of cryo-EM data processing. b, Fourier shell correlation plot for half-maps calculated in FREALIGN. The overall resolution is estimated to be 3.84 Å on the basis of the FSC = 0.143 cut-off criterion. c, Angular distribution plot for particles in the reconstruction. d, Sharpened cryo-EM density map colored according to local resolution using ResMap.
Supplementary Figure 4 Model building into the cryo-EM density map.
a, Representative density fragments with the refined TRPV4 model. b, Fourier shell correlation plots for refined model and half-map 1 (FSC work, red), refined model and half-map 2 (FSC free, blue) and refined model and full map (FSC sum, black).
Supplementary Figure 5 Sequence alignment of TRPV channels.
Sequences of Xenopus tropicalis TRPV4 (xTRPV4, Gene ID: 100496204), Homo sapiens TRPV4 (hTRPV4, Gene ID: 59341), Homo sapiens TRPV1 (hTRPV1, Gene ID: 7442), Homo sapiens TRPV2 (hTRPV2, Gene ID: 51393), Homo sapiens TRPV3 (hTRPV3, Gene ID: 162514), Homo sapiens TRPV5 (hTRPV5, Gene ID: 56302), and Homo sapiens TRPV6 (hTRPV6, Gene ID: 55503) were aligned. Secondary structure elements of xTRPV4 are shown above the sequence, and dashed lines indicate unresolved regions in the cryo-EM structure. Disease mutations discussed in the main text are highlighted in the same colors as in Fig. 6a.
Supplementary Figure 6 Superposition of the S1–S4 domains.
Orthogonal views of superposition of TRPV4 (green) and TRPV1 (grey) S1–S4 domains (PDB: 3J5P).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6
Rights and permissions
About this article
Cite this article
Deng, Z., Paknejad, N., Maksaev, G. et al. Cryo-EM and X-ray structures of TRPV4 reveal insight into ion permeation and gating mechanisms. Nat Struct Mol Biol 25, 252–260 (2018). https://doi.org/10.1038/s41594-018-0037-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0037-5