Abstract
Membrane oxidoreductase CcdA plays a central role in supplying reducing equivalents from the bacterial cytoplasm to the envelope. It transports electrons across the membrane using a single pair of cysteines by a mechanism that has not yet been elucidated. Here we report an NMR structure of the Thermus thermophilus CcdA (TtCcdA) in an oxidized and outward-facing state. CcdA consists of two inverted structural repeats of three transmembrane helices (2 × 3-TM). We computationally modeled and experimentally validated an inward-facing state, which suggests that CcdA uses an elevator-type movement to shuttle the reactive cysteines across the membrane. CcdA belongs to the LysE superfamily, and thus its structure may be relevant to other LysE clan transporters. Structure comparisons of CcdA, semiSWEET, Pnu, and major facilitator superfamily (MFS) transporters provide insights into membrane transporter architecture and mechanism.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Stewart, E. J., Katzen, F. & Beckwith, J. Six conserved cysteines of the membrane protein DsbD are required for the transfer of electrons from the cytoplasm to the periplasm of Escherichia coli. EMBO J. 18, 5963–5971 (1999).
Katzen, F. & Beckwith, J. Transmembrane electron transfer by the membrane protein DsbD occurs via a disulfide bond cascade. Cell 103, 769–779 (2000).
Collet, J. F., Riemer, J., Bader, M. W. & Bardwell, J. C. A. Reconstitution of a disulfide isomerization system. J. Biol. Chem. 277, 26886–26892 (2002).
Malojčić, G., Geertsma, E. R., Brozzo, M. S. & Glockshuber, R. Mechanism of the prokaryotic transmembrane disulfide reduction pathway and its in vitro reconstitution from purified components. Angew. Chem. Int. Ed. Engl. 51, 6900–6903 (2012).
Katzen, F., Deshmukh, M., Daldal, F. & Beckwith, J. Evolutionary domain fusion expanded the substrate specificity of the transmembrane electron transporter DsbD. EMBO J. 21, 3960–3969 (2002).
Cho, S. H. et al. A new family of membrane electron transporters and its substrates, including a new cell envelope peroxiredoxin, reveal a broadened reductive capacity of the oxidative bacterial cell envelope. MBio 3, e00291–11 (2012).
Cho, S.-H. & Collet, J.-F. Many roles of the bacterial envelope reducing pathways. Antioxid. Redox Signal. 18, 1690–1698 (2013).
Simon, J. & Hederstedt, L. Composition and function of cytochrome c biogenesis System II. FEBS J. 278, 4179–4188 (2011).
Bardischewsky, F., Fischer, J., Höller, B. & Friedrich, C. G. SoxV transfers electrons to the periplasm of Paracoccus pantotrophus - an essential reaction for chemotrophic sulfur oxidation. Microbiology 152, 465–472 (2006).
Page, M. L. D. et al. A homolog of prokaryotic thiol disulfide transporter CcdA is required for the assembly of the cytochrome b6f complex in Arabidopsis chloroplasts. J. Biol. Chem. 279, 32474–32482 (2004).
Kimball, R. A., Martin, L. & Saier, M. H. Jr. Reversing transmembrane electron flow: the DsbD and DsbB protein families. J. Mol. Microbiol. Biotechnol. 5, 133–149 (2003).
Cho, S. H. & Beckwith, J. Mutations of the membrane-bound disulfide reductase DsbD that block electron transfer steps from cytoplasm to periplasm in Escherichia coli. J. Bacteriol. 188, 5066–5076 (2006).
Hiniker, A., Vertommen, D., Bardwell, J. C. A. & Collet, J. F. Evidence for conformational changes within DsbD: possible role for membrane-embedded proline residues. J. Bacteriol. 188, 7317–7320 (2006).
Cho, S.-H., Porat, A., Ye, J. & Beckwith, J. Redox-active cysteines of a membrane electron transporter DsbD show dual compartment accessibility. EMBO J. 26, 3509–3520 (2007).
Cho, S.-H. & Beckwith, J. Two snapshots of electron transport across the membrane: insights into the structure and function of DsbD. J. Biol. Chem. 284, 11416–11424 (2009).
Rozhkova, A. & Glockshuber, R. Thermodynamic aspects of DsbD-mediated electron transport. J. Mol. Biol. 380, 783–788 (2008).
Zhou, Y. et al. NMR solution structure of the integral membrane enzyme DsbB: functional insights into DsbB-catalyzed disulfide bond formation. Mol. Cell. 31, 896–908 (2008).
Cierpicki, T. & Bushweller, J. H. Charged gels as orienting media for measurement of residual dipolar couplings in soluble and integral membrane proteins. J. Am. Chem. Soc. 126, 16259–16266 (2004).
Liang, B., Bushweller, J. H. & Tamm, L. K. Site-directed parallel spin-labeling and paramagnetic relaxation enhancement in structure determination of membrane proteins by solution NMR spectroscopy. J. Am. Chem. Soc. 128, 4389–4397 (2006).
Fitzkee, N. C. & Bax, A. Facile measurement of 1H-15N residual dipolar couplings in larger perdeuterated proteins. J. Biomol. NMR 48, 65–70 (2010).
Lomize, M. A., Pogozheva, I. D., Joo, H., Mosberg, H. I. & Lomize, A. L. OPM database and PPM web server: resources for positioning of proteins in membranes. Nucl. Acids Res. 40, D370–D376 (2012).
Deshmukh, M., Brasseur, G. & Daldal, F. Novel Rhodobacter capsulatus genes required for the biogenesis of various c-type cytochromes. Mol. Microbiol. 35, 123–138 (2000).
Williamson, J. A. et al. Structure and multistate function of the transmembrane electron transporter CcdA. Nat. Struct. Mol. Biol. 22, 809–814 (2015).
Vergara-Jaque, A., Fenollar-Ferrer, C., Kaufmann, D. & Forrest, L. R. Repeat-swap homology modeling of secondary active transporters: updated protocol and prediction of elevator-type mechanisms. Front. Pharmacol. 6, 183 (2015).
Mulligan, C. et al. The bacterial dicarboxylate transporter VcINDY uses a two-domain elevator-type mechanism. Nat. Struct. Mol. Biol. 23, 256–263 (2016).
Drew, D. & Boudker, O. Shared molecular mechanisms of membrane transporters. Annu. Rev. Biochem. 85, 543–572 (2016).
Senes, A., Engel, D. E. & DeGrado, W. F. Folding of helical membrane proteins: the role of polar, GxxxG-like and proline motifs. Curr. Opin. Struct. Biol. 14, 465–479 (2004).
Mackenzie, K. R., Prestegard, J. H. & Engelman, D. M. A transmembrane helix dimer: structure and implications. Science 276, 131–133 (1997).
Tsu, B. V. & Saier, M. H. Jr. The LysE superfamily of transport proteins involved in cell physiology and pathogenesis. PLoS. One 10, e0137184 (2015).
Colinet, A. S. et al. Acidic and uncharged polar residues in the consensus motifs of the yeast Ca2+ transporter Gdt1p are required for calcium transport. Cell. Microbiol. 19, e12729 (2017).
Xu, Y. et al. Structures of bacterial homologues of SWEET transporters in two distinct conformations. Nature 515, 448–452 (2014).
Jaehme, M., Guskov, A. & Slotboom, D. J. Crystal structure of the vitamin B3 transporter PnuC, a full-length SWEET homolog. Nat. Struct. Mol. Biol. 21, 1013–1015 (2014).
Jaehme, M., Guskov, A. & Slotboom, D. J. The twisted relation between Pnu and SWEET transporters. Trends Biochem. Sci. 40, 183–188 (2015).
Lee, Y., Nishizawa, T., Yamashita, K., Ishitani, R. & Nureki, O. Structural basis for the facilitative diffusion mechanism by SemiSWEET transporter. Nat. Commun. 6, 6112 (2015).
Joh, N. H. et al. De novo design of a transmembrane Zn2+-transporting four-helix bundle. Science 346, 1520–1524 (2014).
Law, C. J., Maloney, P. C. & Wang, D.-N. Ins and outs of major facilitator superfamily antiporters. Annu. Rev. Microbiol. 62, 289–305 (2008).
Abramson, J. et al. Structure and mechanism of the lactose permease of Escherichia coli. Science 301, 610–615 (2003).
Yan, N. Structural advances for the major facilitator superfamily (MFS) transporters. Trends Biochem. Sci. 38, 151–159 (2013).
Jardetzky, O. Simple allosteric model for membrane pumps. Nature 211, 969–970 (1966).
Yan, N. Structural biology of the major facilitator superfamily transporters. Annu. Rev. Biophys. 44, 257–283 (2015).
Shen, Y. & Bax, A. Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J. Biomol. NMR 56, 227–241 (2013).
Tian, Y., Schwieters, C. D., Opella, S. J. & Marassi, F. M. A practical implicit membrane potential for NMR structure calculations of membrane proteins. Biophys. J. 109, 574–585 (2015).
Acknowledgements
We thank J. F. Ellena for support and advice on NMR. We thank D. S. Cafiso for scientific discussions. This work was supported by grant R01GM078296 to J.H.B. from the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Y.Z. and J.H.B. designed the research; Y.Z. performed the experiments and analyzed the data; Y.Z. and J.H.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 TtCcdA is monodisperse, active, and shows internal sequence repeats
a, Alignment of the internal sequence repeats of TtCcdA (30% identity and 57% similarity). The PCxxP motifs are colored yellow. b, Elution profiles of oxidized TtCcdA in 4.2 mM n-decyl-β-D-maltoside (DM) and 1.5 mM n-dodecyl phosphocholine (DPC) and Bio-Rad gel filtration standard run on S200 size exclusion column (GE Healthcare). Based on the protein standard, the estimated molecular weight of TtCcdA-detergent complex is 70 kDa. c, Time course of the thiol-disulfide exchange reactions between 50 μM reduced TtTrx and 10 μM oxidized TtCcdA in DM and in DPC.
Supplementary Figure 2 Structural restraints, calculation, and validation
a, Paramagnetic relaxation enhancement restraints were recorded as the signal intensity reduction comparing the nitroxide spin labeled paramagnetic sample (right, in this case label on V78C) and the ascorbic acid reduced diamagnetic sample (left). Residues in the red circles show significant signal intensity reductions due to line-broadening. b, Superimposition of the 10 lowest energy structures of oxidized TtCcdA. Green spheres show the locations of the 8 paramagnetic labels. c, 1DHN residual dipolar couplings from neutral gel (black) and positively charged gel (red) are plotted against the TtCcdA sequence. For a straight helix, residual dipolar couplings oscillate periodically in a certain range. The kinks and bends of the helices can be seen in the shift of the data range. d, The backbone RMSD vs. Xplor-NIH energy scatter plot of the fold and refine steps of structural calculation. RMSD is to the lowest energy structure of each calculation step. The green dots represent structures of the correct fold. The dark green dots represent the 10 lowest energy structures of the 200 structures calculated. The red dots represent structures of the mirror image fold. e, The structure of TtCcdA was determined using redundant restraints. Four structural ensembles (10 structures each) were calculated using partial long-range PRE and/or RDC restraints (see methods for the restraints used in each case). Each ensemble was compared with the reported representative structure. The Qfree factors were calculated using the withheld RDC data from the neutral gel. The mean and standard deviation of the pairwise RMSD and Qfree factors are shown as blue bars and red dots, respectively. The structures calculated using seven sets of PRE restraints without RDC restraints (PRE x 7, using ~70% the long-range restraints) are close to the reported structure. The structures calculated using six sets of PRE restraints without RDC restraints (PRE x 6, using ~60% the long-range restraints) could be separated into two clustered groups of equal populations. One has the correct fold and the other has the mirror image fold.
Supplementary Figure 3 Solvent accessibility, Gx6G motifs, dynamics, and membrane topology of TtCcdA
a, The ratio of peak intensity changes upon titration of 20 mM gadolinium paramagnetic relaxation reagent. b, Mapping the intensity change onto the structure of TtCcdA. Residues showing large intensity changes (Ipara/Idia < 0.2) are colored green. Residues showing small changes (Ipara/Idia > 0.5) are colored purple. c, The Gx6G motifs between TM3/TM6 and TM2/TM5. Cartoon representations of the antiparallel left-handed TM3/TM6 (left) and TM2/TM5 (right) pairs, viewed in the membrane from the outside of the protein. “G” is glycine or other small amino acids such as alanine or serine (red). They form complementary surfaces with the large hydrophobic residues (green) from the opposite TM helix. The Gx6G motifs form tight packing between TM helices. d, The random coil index derived order parameters (RCI-S2) are plotted against the TtCcdA sequence. Higher values up to 1 indicate restricted motion on the ns-ps time scale. e, TtCcdA assumes an Nout-Cout membrane topology. The residues of positive charge (arginine and lysine) and negative charge (aspartic acid and glutamic acid) are shown as blue and red sticks, respectively.
Supplementary Figure 4 The structure, sequence alignments, and mapped solvent accessibility of AfCcdA
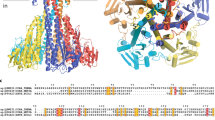
a, Cartoon representations of AfCcdA (PDB 2N4X) as viewed in the membrane plane (left) and from the periplasmic side (right). The two sequence repeats, the non-repeating region, and the two PCxxP motifs are colored red/blue, white, and yellow, respectively. The TM helices are numbered and the horizontal helices are labeled h and h’. The yellow spheres show the locations that correspond to the two essential cysteines. b, The solvent accessibility profile of EcDsbD is mapped onto the structure of AfCcdA (green: solvent accessible, purple: solvent inaccessible). c, AfCcdA displays the internal sequence repeats (red versus blue, 23% identity, 55% similarity). d, Multiple sequence alignment of CcdA. The sequences are from Thermus thermophilus HB8 (YP_144675.1, wild-type TtCcdA in this study), Meiothermus rufus (WP_027881202.1), Marinithermus hydrothermalis (WP_013703256.1), Oceanithermus profundus (WP_013457257.1), Rhodobacter capsulatus (WP_013067872.1), and Archaeoglobus fulgidus DSM 4304 (AAB90190.1, wild-type AfCcdA). The conserved PCxxP, Gx6G and other glycine-containing motifs are colored yellow, cyan, and green, respectively. The six TM helices of AfCcdA are labeled using red letters.
Supplementary Figure 5 Sequence alignment for the repeat-swap homology modeling
a, Schematic of model and template alignment: TM 1-3 (blue) is modeled using the TM 4–6 (red) structure as template, L3 (grey) is modeled de novo, TM 4–6 is modeled using the TM 1–3 structure as template, and H1-H2 (grey) is modeled by itself. b, Model and template alignment used by the MODELLER program to generate the homology models. Blue and red indicate the repeats modeled by each other. Grey indicates the non-repeating sequences modeled using their own structures as templates. The rest are modeled de novo. Open boxes indicate the helices of the model and the template.
Supplementary Figure 6 TROSY-HSQC spectra of the mutants of oxidized TtCcdA
a, 0.27 mM F45C/A113C mutant. b, 0.27 mM F45C/A113C mutant with 0.30 mM HgCl2. c, Overlay of the two previous spectra plotted at the same contour level. d, G154A mutant.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Note
Supplementary Dataset 1
Original gel image of Figure 3b showing the cross-linking of the TtCcdA F45C A113C mutant.
Supplementary Video 1
Video showing conformational change of CcdA going to and from the inward- and outward-facing states.
Rights and permissions
About this article
Cite this article
Zhou, Y., Bushweller, J.H. Solution structure and elevator mechanism of the membrane electron transporter CcdA. Nat Struct Mol Biol 25, 163–169 (2018). https://doi.org/10.1038/s41594-018-0022-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0022-z
This article is cited by
-
Structural and mechanistic analysis of a tripartite ATP-independent periplasmic TRAP transporter
Nature Communications (2022)