Abstract
The APOBEC-AID family of cytidine deaminase prefers single-stranded nucleic acids for cytidine-to-uridine deamination. Single-stranded nucleic acids are commonly involved in the DNA repair system for breaks generated by CRISPR–Cas9. Here, we show in human cells that APOBEC3 can trigger cytidine deamination of single-stranded oligodeoxynucleotides, which ultimately results in base substitution mutations in genomic DNA through homology-directed repair (HDR) of Cas9-generated double-strand breaks. In addition, the APOBEC3-catalyzed deamination in genomic single-stranded DNA formed during the repair of Cas9 nickase-generated single-strand breaks in human cells can be further processed to yield mutations mainly involving insertions or deletions (indels). Both APOBEC3-mediated deamination and DNA-repair proteins play important roles in the generation of these indels. Therefore, optimizing conditions for the repair of CRISPR–Cas9-generated DNA breaks, such as using double-stranded donors in HDR or temporarily suppressing endogenous APOBEC3s, can repress these unwanted mutations in genomic DNA.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Harris, R. S. & Liddament, M. T. Retroviral restriction by APOBEC proteins. Nat. Rev. Immunol. 4, 868–877 (2004).
Henderson, S. & Fenton, T. APOBEC3 genes: retroviral restriction factors to cancer drivers. Trends Mol. Med. 21, 274–284 (2015).
Salter, J. D., Bennett, R. P. & Smith, H. C. The APOBEC protein family: united by structure, divergent in function. Trends Biochem. Sci. 41, 578–594 (2016).
Yang, B., Li, X., Lei, L. & Chen, J. APOBEC: From mutator to editor. J. Genet. Genomics 44, 423–437 (2017).
Chen, J., Miller, B. F. & Furano, A. V. Repair of naturally occurring mismatches can induce mutations in flanking DNA. eLife 3, e02001 (2014).
Chen, J. & Furano, A. V. Breaking bad: the mutagenic effect of DNA repair. DNA Repair (Amst.) 32, 43–51 (2015).
Roberts, S. A. et al. Clustered mutations in yeast and in human cancers can arise from damaged long single-strand DNA regions. Mol. Cell 46, 424–435 (2012).
Taylor, B. J. et al. DNA deaminases induce break-associated mutation showers with implication of APOBEC3B and 3 A in breast cancer kataegis. eLife 2, e00534 (2013).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Burns, M. B., Temiz, N. A. & Harris, R. S. Evidence for APOBEC3B mutagenesis in multiple human cancers. Nat. Genet. 45, 977–983 (2013).
Roberts, S. A. et al. An APOBEC cytidine deaminase mutagenesis pattern is widespread in human cancers. Nat. Genet. 45, 970–976 (2013).
Helleday, T., Eshtad, S. & Nik-Zainal, S. Mechanisms underlying mutational signatures in human cancers. Nat. Rev. Genet. 15, 585–598 (2014).
Chan, K. & Gordenin, D. A. Clusters of multiple mutations: incidence and molecular mechanisms. Annu. Rev. Genet. 49, 243–267 (2015).
Mali, P., Esvelt, K. M. & Church, G. M. Cas9 as a versatile tool for engineering biology. Nat. Methods 10, 957–963 (2013).
Doudna, J. A. & Charpentier, E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science 346, 1258096 (2014).
Sander, J. D. & Joung, J. K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32, 347–355 (2014).
Cox, D. B., Platt, R. J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015).
Komor, A. C., Badran, A. H. & Liu, D. R. CRISPR-based technologies for the manipulation of eukaryotic genomes. Cell 168, 20–36 (2017).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Wu, Y. et al. Correction of a genetic disease in mouse via use of CRISPR-Cas9. Cell Stem Cell 13, 659–662 (2013).
Yin, H. et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat. Biotechnol. 32, 551–553 (2014).
Symington, L. S. & Gautier, J. Double-strand break end resection and repair pathway choice. Annu. Rev. Genet. 45, 247–271 (2011).
Myler, L. R. et al. Single-molecule imaging reveals the mechanism of Exo1 regulation by single-stranded DNA binding proteins. Proc. Natl. Acad. Sci. USA 113, E1170–E1179 (2016).
Burns, M. B. et al. APOBEC3B is an enzymatic source of mutation in breast cancer. Nature 494, 366–370 (2013).
Refsland, E. W. et al. Quantitative profiling of the full APOBEC3 mRNA repertoire in lymphocytes and tissues: implications for HIV-1 restriction. Nucleic Acids Res. 38, 4274–4284 (2010).
Harris, R. S. et al. DNA deamination mediates innate immunity to retroviral infection. Cell 113, 803–809 (2003).
Anand, R., Beach, A., Li, K. & Haber, J. Rad51-mediated double-strand break repair and mismatch correction of divergent substrates. Nature 544, 377–380 (2017).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Cho, S. W. et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 24, 132–141 (2014).
Fu, Y., Sander, J. D., Reyon, D., Cascio, V. M. & Joung, J. K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Tsai, S. Q. et al. Dimeric CRISPR RNA-guided FokI nucleases for highly specific genome editing. Nat. Biotechnol. 32, 569–576 (2014).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Nishida, K. et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353, aaf8729 (2016).
Kim, K. et al. Highly efficient RNA-guided base editing in mouse embryos. Nat. Biotechnol. 35, 435–437 (2017).
Zhou, C. et al. Highly efficient base editing in human tripronuclear zygotes. Protein Cell (2017).
Li, G. et al. Highly efficient and precise base editing in discarded human tripronuclear embryos. Protein Cell 8, 772–775 (2017).
Paquet, D. et al. Efficient introduction of specific homozygous and heterozygous mutations using CRISPR/Cas9. Nature 533, 125–129 (2016).
Carter, R. J. & Parsons, J. L. Base excision repair, a pathway regulated by posttranslational modifications. Mol. Cell. Biol. 36, 1426–1437 (2016).
Daley, J. M., Niu, H., Miller, A. S. & Sung, P. Biochemical mechanism of DSB end resection and its regulation. DNA Repair (Amst.) 32, 66–74 (2015).
Starrett, G. J. et al. The DNA cytosine deaminase APOBEC3H haplotype I likely contributes to breast and lung cancer mutagenesis. Nat. Commun. 7, 12918 (2016).
Long, C. et al. Postnatal genome editing partially restores dystrophin expression in a mouse model of muscular dystrophy. Science 351, 400–403 (2016).
Nelson, C. E. et al. In vivo genome editing improves muscle function in a mouse model of Duchenne muscular dystrophy. Science 351, 403–407 (2016).
Tabebordbar, M. et al. In vivo gene editing in dystrophic mouse muscle and muscle stem cells. Science 351, 407–411 (2016).
Bonvin, M. et al. Interferon-inducible expression of APOBEC3 editing enzymes in human hepatocytes and inhibition of hepatitis B virus replication. Hepatology 43, 1364–1374 (2006).
Kim, Y. B. et al. Increasing the genome-targeting scope and precision of base editing with engineered Cas9-cytidine deaminase fusions. Nat. Biotechnol. 35, 371–376 (2017).
Li, J., Sun, Y., Du, J., Zhao, Y. & Xia, L. Generation of targeted point mutations in rice by a modified CRISPR/Cas9 system. Mol. Plant 10, 526–529 (2017).
Liang, P. et al. Correction of β-thalassemia mutant by base editor in human embryos. Protein Cell 8, 811–822 (2017).
Lu, Y. & Zhu, J. K. Precise editing of a target base in the rice genome using a modified CRISPR/Cas9 system. Mol. Plant 10, 523–525 (2017).
Rees, H. A. et al. Improving the DNA specificity and applicability of base editing through protein engineering and protein delivery. Nat. Commun. 8, 15790 (2017).
Shimatani, Z. et al. Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat. Biotechnol. 35, 441–443 (2017).
Zhang, Y. et al. Programmable base editing of zebrafish genome using a modified CRISPR-Cas9 system. Nat. Commun. 8, 118 (2017).
Zong, Y. et al. Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat. Biotechnol. 35, 438–440 (2017).
Hess, G. T., Tycko, J., Yao, D. & Bassik, M. C. Methods and applications of CRISPR-mediated base editing in eukaryotic genomes. Mol. Cell 68, 26–43 (2017).
Mitsunobu, H., Teramoto, J., Nishida, K. & Kondo, A. Beyond native Cas9: manipulating genomic information and function. Trends Biotechnol. 35, 983–996 (2017).
Komor, A. C. et al. Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T: Abase editors with higher efficiency and product purity. Sci. Adv. 3, eaao4774 (2017).
Wang, L. et al. Enhanced base editing by co-expression of free uracil DNA glycosylase inhibitor. Cell Res. 27, 1289–1292 (2017).
Shen, B. et al. Efficient genome modification by CRISPR-Cas9 nickase with minimal off-target effects. Nat. Methods. 11, 399–402 (2014).
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 31, 833–838 (2013).
Bogerd, H. P., Wiegand, H. L., Doehle, B. P. & Cullen, B. R. The intrinsic antiretroviral factor APOBEC3B contains two enzymatically active cytidine deaminase domains. Virology 364, 486–493 (2007).
Acknowledgements
We are grateful to A. Furano, H. Lin and H. Wang for discussing and commenting on this manuscript, L.-L. Chen and N. Jing for technical support, X. Li and Y. Pan for participating in the examination of APOBEC expression, J. Wu for maintaining cell lines and H. Fang for participating in deep-sequencing library preparation. Next-generation deep sequencing was performed at the CAS-MPG PICB Omics Core, Shanghai, China. This work is supported by a MOST grant (2014CB910600 to L. Yang), NSFC grants (91540115 to L. Yang, 31571372 to B.S., 31471241 to L. Yang, 31600619 to B.Y. and 31600654 to J.C.), the Shanghai Pujiang program (16PJ1407000 to J.C. and 16PJ1407500 to B.Y.) and CAS Key Laboratory of Computational Biology grants (2015KLCB01 and 2016KLCB01 to L. Yang and J.C.).
Author information
Authors and Affiliations
Contributions
J.C., L. Yang and B.S. conceived, designed and supervised the project. L.L., H.C., B.Y. and B.H. performed most of the experiments with the help of L.W., Y.C., W.L. and J. Wang on RT–qPCR, plasmid construction and in vitro transcription and W.S. and L. Yan on Cas9 protein purification. J. Wei prepared samples for deep sequencing, and W.X. performed the deep-sequencing data analyses and bioinformatics analysis, supervised by L. Yang. J.G., J.S., M.Z. and X.H. provided critical technical assistance. B.Y., J.C., L. Yang and B.S. wrote the paper with inputs from all authors. J.C. managed the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated Supplementary Information
Supplementary Figure 1 Examination of APOBEC3’s effect on the base substitution frequency of sgRNA and ODNs.
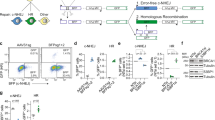
(a) Endogenous expression of APOBEC-AID family members in 293FT and HeLa cells. mRNA levels of APOBEC-AID family members were determined by RT-qPCR and normalized against TATA binding protein (TBP) mRNA levels. (b) Western blots of the whole-cell lysates from wild type 293FT cells (293FT) and 293FT cells stably expressing the C-terminal HA tagged APOBEC3B (A3B) or the catalytically inactive one (A3Bm). Western blots were performed using anti-HA antibody. Alpha tubulin served as loading control. Uncropped blot images are shown in Supplementary Data Set 1. The blots shown are representative of three experiments. (c) Relative APOBEC3B mRNA level in 293FT and A3B cells. APOBEC3B mRNA levels were determined by RT-qPCR and normalized against TATA binding protein (TBP) mRNA levels. (d) Schematic diagram illustrating the procedures to examine the base substitutions in the sgRNAs transfected into cells. (e) Base substitution frequency of each base in the spacer regions of indicated sgRNAs that are either non-transfected or transfected into 293FT or A3B cells. The data were from two independent experiments. (f) Base substitution frequency of each base in ssODNs or dsODNs that are transfected into A3B or A3Bm cells. (g) Comparison of the theoretical base substitution fractions at each dinucleotide calculated from the base content of ssODNs and the experimentally determined ones in 293FT and A3B cells. The base substations on C of CpC or TpC reflected APOBEC3 mutational signatures. Red arrows highlight the same base substitutions as observed in the ODN-cognate genomic regions in Fig. 2. (a, c, f) The means (±s.d.) were from three independent experiments.
Supplementary Figure 2 Amplification of ODN-cognate genomic region derived from HDR.
(a) Schematic diagrams illustrating the procedures to amplify the ODN-cognate genomic region from HDR products. (b), (c) Tag-specific genomic DNA PCR was performed using 3' end phosphorothioate modified primers, which prevent the 3' to 5' exonuclease activity of Hi-Fi DNA polymerases, to amplify the ODN-cognate genomic region from 293FT, A3B and A3Bm cells that are treated as indicated. The gels shown in (b, c) are representative of three experiments. (d), (e) Tag-specific genomic DNA PCR was performed to amplify the ODN-cognate genomic region from primary human T cells that are treated as indicated. The gels shown in (d, e) are representative of two experiments. Uncropped gel images for (b-e) are shown in Supplementary Data Set 1.
Supplementary Figure 3 APOBEC3s expression in primary human T cells.
(a) Expression of endogenous APOBEC3s was up-regulated during primary human T cell activation. mRNA levels of APOBEC3 family in naive (day 0) and activated (day 3, 7 and 10) primary human CD8+ T cells were determined by RT-qPCR and normalized against TATA binding protein (TBP) mRNA levels. ND denotes non-detected. (b) Efficiency of endogenous APOBEC3s knockdown. (c) Schematic diagram illustrating that APOBEC could target both ssODN donor and the complementary genomic ssDNA for cytidine deamination, which finally cause base substitutions on Cs and Gs in genomic DNA. (a, b) The means (±s.d.) were from three independent experiments.
Supplementary Figure 4 APOBEC3 introduces sparse base substitutions in genomic DNA near Cas9 nickase-generated SSBs.
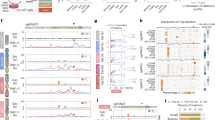
(a) Schematic diagram illustrating the procedures to determine the base substitution and indel mutations induced by Cas9 variant-generated breaks in genomic DNA. (b) Base substitution frequency induced by Cas9 variant-generated breaks in genomic DNA. 293FT cells were either left non-transfected (NT) or co-transfected with indicated sgRNAs and Cas9, D10A, H840A or dCas9. Base substitution frequency of each base in the upstream and downstream 25-bp region of Cas9 cleavage sites was measured by deep sequencing. Red arrow indicates the APOBEC3-featured base substitution. Asterisk indicates a base substitution specifically induced by sgVEGFA-3-Cas9, which however was not up-regulated by APOBEC3B over-expression (data not shown). Other mutations were all background base substitutions that remain largely unchanged in all tested situations (compare Cas9, D10A, H840A, dCas9 and NT with each other). (c) Base substitution frequency induced by sgVEGFA-D10A-generated SSB was up-regulated by A3B overexpression. 293FT, A3B or A3Bm cells were co-transfected with sgVEGFA together with Cas9, D10A, H840A or dCas9, respectively. (b, c) The means (±s.d.) were from three independent experiments.
Supplementary Figure 5 Overexpression of APOBEC3 upregulates the indel formation induced by Cas9 nickase-generated SSB in genomic DNA.
(a), (b) Indel frequencies induced by Cas9 variant-generated breaks in 293FT (a) and HeLa (b) cells. 293FT and HeLa cells were either non-transfected (NT) or co-transfected with indicated sgRNAs together with Cas9, D10A, H840A or dCas9, after which the indel frequencies were determined by deep sequencing. (c) Left: Schematic diagram illustrating that indels induced by Cas9 nickases are more distal to SSB sites, while Cas9 mainly induced indels at DSB site (at the ±3-bp region). Right: Statistical analysis of the fractions of reads containing indels at the ±3-bp region of cleavage site. The data represented are from (a, b) and Fig. 3b. The median, interquartile range (IQR), 1.5 × IQR and outliers are shown. ***: P < 0.001, one-tailed Student’s t-test. (d) Indel frequencies induced by Cas9 variant-generated breaks in 293FT, A3B or A3Bm cells. (e) Indel frequencies at indicated non-relevant genomic loci in A3B cells that are co-transfected with indicated sgRNAs and Cas9, D10A, H840A or dCas9. The data were from two independent experiments. (a, b, d) The means (±s.d.) were from three independent experiments.
Supplementary Figure 6 Examination of base substitution and indel mutations induced by Cas9-variant-generated breaks in episomal shuttle vectors.
(a) Schematic diagram illustrating the procedures to determine the base substitution and indel mutations induced by Cas9 variant-generated breaks in episomal shuttle vectors. (b) Number of colonies containing mutated shuttle vectors that are induced by Cas9 variant-generated breaks. (c) Indel frequencies induced by Cas9 variant-generated breaks in episomal shuttle vectors. The means (±s.d.) were from three independent experiments. (d) Fractions at each base and distribution curves of Cas9 variant-induced indels at the cleavage site upstream and downstream 50-bp region in episomal shuttle vectors.
Supplementary Figure 7 Distributions of indels induced by Cas9 variants.
(a) 293FT indel fraction at each base (indel counts at each base relative to the total indel counts in the cleavage site upstream and downstream 25-bp region, %) and the fitting curves of indel fractions in the same region. The indel fitting curves illustrating indel distribution are presented in Fig. 3c. (b) Left: Fractions at each base and distribution curves of Cas9 or D10A-induced indels in the cleavage site upstream and downstream 25-bp region for A3B cells. Right: statistical analysis of the distances from the cleavage site to curve peaks. The median, interquartile range (IQR) and 1.5 × IQR are shown. ***: P < 0.001, one-tailed Student’s t-test.
Supplementary Figure 8 APOBEC3 binds to genomic ssDNA regions exposed near Cas9 nickase-generated SSBs and induces indel formation.
(a) Indel frequencies induced by Cas9-generated DSB at indicated activation time points in primary human T cells that are pretreated with control siRNA (siCtrl) or siRNA against endogenous APOBECs (siA3(Mix)). (b) Normalized indel frequencies (D10A- or dCas9-induced indel frequencies relative to Cas9-induced ones) at indicated activation time points in primary human T cells. The indel frequencies induced by Cas9 increased during T cell activation (a), suggesting that the sgRNA-Cas9 RNP electroporation efficiency may increase during T cell activation. Therefore, D10A-induced indel frequencies (Fig. 3f) were normalized against Cas9-induced ones to exclude the effect of electroporation efficiency. (c) APOBEC3 knockdown suppressed the indel formation induced by Cas9 nickase-generated SSB in episomal shuttle vector. (d) A3B cells were either non-transfected (NT) or co-transfected with indicated sgRNAs and Cas9, D10A, H840A or dCas9. ChIP-qPCR assays were then performed to detect the binding capacities of histone H3 (using H3 antibody, H3 ab) at indicated genomic loci. The results from the anti-6×His-tag antibody (Ctrl ab) were included as negative controls. The means (±s.d.) were from three independent experiments. (a-c) The data were from two independent experiments.
Supplementary Figure 9 Inhibiting DNA-repair enzymes manifested their effects on indel formation.
(a)-(d) Knockdown or knockout of DNA repair enzymes MRE11, Exo1, UNG, SMUG1 and APE1. The inhibitions were examined by western blots and the knockouts were confirmed by sequencing the genomic loci of indicated gene. (e)-(h) Inhibition of various DNA repair enzymes manifested their effects on the indel formations induced by Cas9 variant-generated breaks. 293FT and the corresponding knockdown (KD) or knockout (KO) cells were co-transfected with indicated sgRNAs and Cas9, BE3, D10A, H840A or dCas9, after which the indel frequencies were determined by deep sequencing. (e-h) The means (±s.d.) were from three independent experiments for sgEMX1. The data were from two independent experiments for sgRNF2. Uncropped blot images for (a-d) are shown in Supplementary Data Set 1. All blots shown are representative of three experiments.
Supplementary Figure 10 Association of sgRNA-H840A RNP complex to single-stranded target DNA strand.
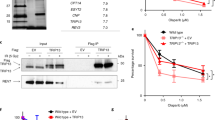
(a), (b) Purified Cas9 and Cas9 nickases showed expected nuclease or nickase activities in oligonucleotides cleavage assay. Double stranded DNA substrates were 5’ labeled in the target (a) or non-target (b) strand. (c), (d) EMSA assay results showed that sgRNA-H840A and sgRNA-dCas9 RNP complex can re-associate to the single-stranded target strand that are generated via resection of non-target strand from the SSB (right panels), while sgRNA-D10A and sgRNA-dCas9 RNP complex cannot re-associate to the single-stranded non-target strand (left panels). (e) Schematic diagram illustrating the hypothesis that different indel frequencies induced by D10A- and H840A-generated SSB result from their different strand preferences. Uncropped gel images for (a-d) are shown in Supplementary Data Set 1. All gels shown are representative of three experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Tables 1–4 and Supplementary Note 1.
Supplementary Dataset 1
Uncropped images
Supplementary Dataset 2
Base substitution frequency determined by deep sequencing
Supplementary Dataset 3
Indel frequency determined by deep sequencing
Rights and permissions
About this article
Cite this article
Lei, L., Chen, H., Xue, W. et al. APOBEC3 induces mutations during repair of CRISPR–Cas9-generated DNA breaks. Nat Struct Mol Biol 25, 45–52 (2018). https://doi.org/10.1038/s41594-017-0004-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-017-0004-6
This article is cited by
-
Enhancement of the viability of T cells electroporated with DNA via osmotic dampening of the DNA-sensing cGAS–STING pathway
Nature Biomedical Engineering (2023)
-
Design and application of the transformer base editor in mammalian cells and mice
Nature Protocols (2023)
-
The coevolution between APOBEC3 and retrotransposons in primates
Mobile DNA (2022)
-
Gene editing and its applications in biomedicine
Science China Life Sciences (2022)
-
Eliminating base-editor-induced genome-wide and transcriptome-wide off-target mutations
Nature Cell Biology (2021)