Abstract
The precise regulation of gene expression is fundamental to neurodevelopment, plasticity and cognitive function. Although several studies have profiled transcription in the developing human brain, there is a gap in understanding of accompanying translational regulation. In this study, we performed ribosome profiling on 73 human prenatal and adult cortex samples. We characterized the translational regulation of annotated open reading frames (ORFs) and identified thousands of previously unknown translation events, including small ORFs that give rise to human-specific and/or brain-specific microproteins, many of which we independently verified using proteomics. Ribosome profiling in stem-cell-derived human neuronal cultures corroborated these findings and revealed that several neuronal activity-induced non-coding RNAs encode previously undescribed microproteins. Physicochemical analysis of brain microproteins identified a class of proteins that contain arginine-glycine-glycine (RGG) repeats and, thus, may be regulators of RNA metabolism. This resource expands the known translational landscape of the human brain and illuminates previously unknown brain-specific protein products.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Data reprocessed from Wang et al.70 were accessed from ArrayExpress with accession number E-MTAB-7247. Human brain primary tissue RNA-seq and Ribo-seq data have been submitted to the database of Genotypes and Phenotypes (dbGaP) under accession number phs002489. NGN2 RNA-seq and Ribo-seq data have been submitted to the Gene Expression Omnibus under accession number GSE180240. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD035950. Our web-based searchable database is available from https://greenberg.hms.harvard.edu/project/human-brain-orf-database/. Source data are provided with this paper.
Code availability
Custom Python code for physicochemical analysis is available at https://github.com/IPritisanac/idr.mol.feats. All other code used in this study was previously published and is cited in the Methods section.
References
Kim, M.-S. et al. A draft map of the human proteome. Nature 509, 575–581 (2014).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. S. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Guydosh, N. R. & Green, R. Dom34 rescues ribosomes in 3′ untranslated regions. Cell 156, 950–962 (2014).
van Heesch, S. et al. The translational landscape of the human heart. Cell 178, 242–260.e29 (2019).
Prensner, J. R. et al. Noncanonical open reading frames encode functional proteins essential for cancer cell survival. Nat. Biotechnol. 39, 697–704 (2021).
Makarewich, C. A. et al. The DWORF micropeptide enhances contractility and prevents heart failure in a mouse model of dilated cardiomyopathy. eLife 7, e38319 (2018).
Makarewich, C. A. et al. MOXI is a mitochondrial micropeptide that enhances fatty acid β-oxidation. Cell Rep. 23, 3701–3709 (2018).
D’Lima, N. G. et al. A human microprotein that interacts with the mRNA decapping complex. Nat. Chem. Biol. 13, 174–180 (2017).
Chen, J. et al. Pervasive functional translation of noncanonical human open reading frames. Science 367, 1140–1146 (2020).
Ji, Z. RibORF: identifying genome-wide translated open reading frames using ribosome profiling. Curr. Protoc. Mol. Biol. 124, e67 (2018).
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Ji, Z., Song, R., Regev, A. & Struhl, K. Many lncRNAs, 5′ UTRs, and pseudogenes are translated and some are likely to express functional proteins. eLife 4, e08890 (2015).
Kalish, B. T. et al. Maternal immune activation in mice disrupts proteostasis in the fetal brain. Nat. Neurosci. 24, 204–213 (2021).
Slavoff, S. A., Heo, J., Budnik, B. A., Hanakahi, L. A. & Saghatelian, A. A human short open reading frame (sORF)-encoded polypeptide that stimulates DNA end joining. J. Biol. Chem. 289, 10950–10957 (2014).
Miller, J. A. et al. Transcriptional landscape of the prenatal human brain. Nature 508, 199–206 (2014).
Kang, H. J. et al. Spatio-temporal transcriptome of the human brain. Nature 478, 483–489 (2011).
Chothani, S. et al. deltaTE: detection of translationally regulated genes by integrative analysis of Ribo-seq and RNA-seq data. Curr. Protoc. Mol. Biol. 129, e108 (2019).
Levy, S., Avni, D., Hariharan, N., Perry, R. P. & Meyuhas, O. Oligopyrimidine tract at the 5′ end of mammalian ribosomal protein mRNAs is required for their translational control. Proc. Natl Acad. Sci. USA 88, 3319–3323 (1991).
Jeong, H. et al. Evolution of DNA methylation in the human brain. Nat. Commun. 12, 2021 (2021).
Huh, G. S. et al. Functional requirement for class I MHC in CNS development and plasticity. Science 290, 2155–2159 (2000).
Stevens, B. et al. The classical complement cascade mediates CNS synapse elimination. Cell 131, 1164–1178 (2007).
Klaudiny, J., von der Kammer, H. & Scheit, K. H. Characterization by cDNA cloning of the mRNA of a highly basic human protein homologous to the yeast ribosomal protein YL41. Biochem. Biophys. Res. Commun. 187, 901–906 (1992).
Odermatt, A. et al. Characterization of the gene encoding human sarcolipin (SLN), a proteolipid associated with SERCA1: absence of structural mutations in five patients with Brody disease. Genomics 45, 541–553 (1997).
Khitun, A. & Slavoff, S. A. Proteomic detection and validation of translated small open reading frames. Curr. Protoc. Chem. Biol. 11, e77 (2019).
Johnson, E. C. B. et al. Large-scale proteomic analysis of Alzheimer’s disease brain and cerebrospinal fluid reveals early changes in energy metabolism associated with microglia and astrocyte activation. Nat. Med 26, 769–780 (2020).
Frigerio, F. et al. Deletion of glutamate dehydrogenase 1 (Glud1) in the central nervous system affects glutamate handling without altering synaptic transmission. J. Neurochem 123, 342–348 (2012).
Lander, S. S. et al. Glutamate dehydrogenase-deficient mice display schizophrenia-like behavioral abnormalities and CA1-specific hippocampal dysfunction. Schizophr. Bull. 45, 127–137 (2019).
Sinvani, H. et al. Translational tolerance of mitochondrial genes to metabolic energy stress involves TISU and eIF1-eIF4GI cooperation in start codon selection. Cell Metab. 21, 479–492 (2015).
Nehme, R. et al. Combining NGN2 programming with developmental patterning generates human excitatory neurons with NMDAR-mediated synaptic transmission. Cell Rep. 23, 2509–2523 (2018).
Sanchez-Priego, C. et al. Mapping cis-regulatory elements in human neurons links psychiatric disease heritability and activity-regulated transcriptional programs. Cell Rep. 39, 110877 (2022).
Zhang, P. et al. Genome-wide identification and differential analysis of translational initiation. Nat. Commun. 8, 1749 (2017).
Ataman, B. et al. Evolution of Osteocrin as an activity-regulated factor in the primate brain. Nature 539, 242–247 (2016).
Issler, O. et al. Sex-specific role for the long non-coding RNA LINC00473 in depression. Neuron 106, 912–926 (2020).
Domazet-Loso, T., Brajković, J. & Tautz, D. A phylostratigraphy approach to uncover the genomic history of major adaptations in metazoan lineages. Trends Genet. 23, 533–539 (2007).
Calviello, L. et al. Detecting actively translated open reading frames in ribosome profiling data. Nat. Methods 13, 165–170 (2016).
Playfoot, C. J. et al. Transposable elements and their KZFP controllers are drivers of transcriptional innovation in the developing human brain. Genome Res. 31, 1531–1545 (2021).
Carlevaro-Fita, J. et al. Ancient exapted transposable elements promote nuclear enrichment of human long noncoding RNAs. Genome Res. 29, 208–222 (2019).
Kapusta, A. et al. Transposable elements are major contributors to the origin, diversification, and regulation of vertebrate long noncoding RNAs. PLoS Genet. 9, e1003470 (2013).
Johnson, R. & Guigó, R. The RIDL hypothesis: transposable elements as functional domains of long noncoding RNAs. RNA 20, 959–976 (2014).
Uhlén, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Brar, G. A. et al. High-resolution view of the yeast meiotic program revealed by ribosome profiling. Science 335, 552–557 (2012).
Chew, G.-L., Pauli, A. & Schier, A. F. Conservation of uORF repressiveness and sequence features in mouse, human and zebrafish. Nat. Commun. 7, 11663 (2016).
Tresenrider, A. et al. Integrated genomic analysis reveals key features of long undecoded transcript isoform-based gene repression. Mol. Cell 81, 2231–2245.e11 (2021).
Aspden, J. L. et al. Extensive translation of small open reading frames revealed by Poly-Ribo-Seq. eLife 3, e03528 (2014).
Rasmussen, A. H., Rasmussen, H. B. & Silahtaroglu, A. The DLGAP family: neuronal expression, function and role in brain disorders. Mol. Brain 10, 43 (2017).
Xing, J. et al. Resequencing and association analysis of six PSD-95-related genes as possible susceptibility genes for schizophrenia and autism spectrum disorders. Sci. Rep. 6, 27491 (2016).
Prilusky, J. et al. FoldIndex: a simple tool to predict whether a given protein sequence is intrinsically unfolded. Bioinformatics 21, 3435–3438 (2005).
Tsang, B. et al. Phosphoregulated FMRP phase separation models activity-dependent translation through bidirectional control of mRNA granule formation. Proc. Natl Acad. Sci. USA 116, 4218–4227 (2019).
Gueroussov, S. et al. Regulatory expansion in mammals of multivalent hnRNP assemblies that globally control alternative splicing. Cell 170, 324–339 (2017).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Zarin, T. et al. Proteome-wide signatures of function in highly diverged intrinsically disordered regions. eLife 8, e46883 (2019).
Chong, P. A., Vernon, R. M. & Forman-Kay, J. D. RGG/RG motif regions in RNA binding and phase separation. J. Mol. Biol. 430, 4650–4665 (2018).
Jishi, A., Qi, X. & Miranda, H. C. Implications of mRNA translation dysregulation for neurological disorders. Semin. Cell Dev. Biol. 114, 11–19 (2021).
Chen, Y.-C., Chang, Y.-W. & Huang, Y.-S. Dysregulated translation in neurodevelopmental disorders: an overview of autism-risk genes involved in translation. Dev. Neurobiol. 79, 60–74 (2019).
Kelleher, R. J. & Bear, M. F. The autistic neuron: troubled translation? Cell 135, 401–406 (2008).
Kapur, M., Monaghan, C. E. & Ackerman, S. L. Regulation of mRNA translation in neurons-a matter of life and death. Neuron 96, 616–637 (2017).
de Goede, O. M. et al. Population-scale tissue transcriptomics maps long non-coding RNAs to complex disease. Cell 184, 2633–2648.e19 (2021).
McGlincy, N. J. & Ingolia, N. T. Transcriptome-wide measurement of translation by ribosome profiling. Methods 126, 112–129 (2017).
Ordureau, A. et al. Global landscape and dynamics of Parkin and USP30-dependent ubiquitylomes in iNeurons during mitophagic signaling. Mol. Cell 77, 1124–1142.e10 (2020).
Seifuddin, F. et al. lncRNAKB, a knowledgebase of tissue-specific functional annotation and trait association of long noncoding RNA. Sci. Data 7, 326 (2020).
Dong, M. et al. SCDC: bulk gene expression deconvolution by multiple single-cell RNA sequencing references. Brief. Bioinform. 22, 416–427 (2021).
Nagy, C. et al. Single-nucleus transcriptomics of the prefrontal cortex in major depressive disorder implicates oligodendrocyte precursor cells and excitatory neurons. Nat. Neurosci. 23, 771–781 (2020).
Fields, A. P. et al. A regression-based analysis of ribosome-profiling data reveals a conserved complexity to mammalian translation. Mol. Cell 60, 816–827 (2015).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
de Hoon, M. J. L., Imoto, S., Nolan, J. & Miyano, S. Open source clustering software. Bioinformatics 20, 1453–1454 (2004).
Saldanha, A. J. Java Treeview—extensible visualization of microarray data. Bioinformatics 20, 3246–3248 (2004).
Kumar, S., Stecher, G., Suleski, M. & Hedges, S. B. TimeTree: a resource for timelines, timetrees, and divergence times. Mol. Biol. Evol. 34, 1812–1819 (2017).
Turelli, P. et al. Primate-restricted KRAB zinc finger proteins and target retrotransposons control gene expression in human neurons. Sci. Adv. 6, eaba3200 (2020).
Hubley, R. et al. The Dfam database of repetitive DNA families. Nucleic Acids Res. 44, D81–D89 (2016).
Wang, Z.-Y. et al. Transcriptome and translatome co-evolution in mammals. Nature 588, 642–647 (2020).
Boulting, G. L. et al. Activity-dependent regulome of human GABAergic neurons reveals new patterns of gene regulation and neurological disease heritability. Nat. Neurosci. 24, 437–448 (2021).
Acknowledgements
This research was supported by the Allen Discovery Center program, a Paul G. Allen Frontiers Group advised program of the Paul G. Allen Family Foundation. E.E.D. was supported by the Damon Runyon Cancer Research Foundation (DRG-2397-20). A.C.C. was supported by the Hanna H. Gray Fellowship through the Howard Hughes Medical Institute. V.L. was supported by a Boston Children’s Hospital Career Development Award (to A.O.D.L.), and V.L. and N.S. were supported by National Human Genomic Research Institute (NHGRI) R01HG010898 and NIH 1R01HG010898-01A1. E.J.H. was supported by NIH P01 NS083513. A.O.D.L. and W.P. were supported by a Manton Center Endowed Scholar Award and NHGRI U01HG008900. M.A.S. was supported by National Institute of Mental Health (NIMH) F31MH124393. A.M.M. and J.D.F.-K. acknowledge funding from the Canadian Institutes of Health Research (CIHR PJT-148532). B.T.K. was supported by National Institute of Neurological Diseases and Stroke (NINDS) K08 NS112338. M.E.G. was supported by funding from NINDS R01 NS115965. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. We thank members of the Greenberg laboratory for helpful discussions on the manuscript. We thank the Taplin Mass Spectrometry Facility at Harvard Medical School for their technical expertise and analysis of proteomics samples and S. Slavoff for advice on size-selection proteomics. We thank the Broad Institute Genomics Program for next-generation sequencing of ribosome profiling libraries. We are grateful to the NIH NeuroBioBank and the Human Developmental Biology Resource for providing human adult and prenatal brain tissue, respectively. We are grateful to the laboratory of D. Trono for sharing the human transposable element annotation and to W. Harper for reagents and technical advice related to iPSC-derived human neurons. We thank the Neurobiology Department and the Neurobiology Imaging Facility for consultation and instrument availability that supported this work. This facility is supported, in part, by the Neural Imaging Center as part of an NINDS P30 Core Center grant (NS072030). Figures 3d, 4a and 7d were made with BioRender.
Author information
Authors and Affiliations
Contributions
E.E.D., B.T.K. and M.E.G. conceptualized the study and designed the experiments. E.E.D. and B.T.K. performed ribosome profiling and RNA-seq. E.E.D., B.T.K. and B.F. analyzed ribosome profiling and RNA-seq. E.E.D. and J.C. analyzed harringtonine-treated Ribo-seq datasets. G.C. prepared human embryonic stem-cell-derived neurons for ribosome profiling. A.C.C. analyzed the insertion of TEs at ORF translation start sites. I.P., A.A., J.F.-K. and A.M.M. performed physiochemical analysis. V.L., A.O.D.L, A. Karger, W.P. and N.S. performed phylostratigraphy analysis. B.T.K. prepared samples for proteomics, and B.T.K., B.F. and A. Khitun analyzed proteomics data. E.E.D., E.G.A. and N.P. performed microprotein validation experiments in 293T cells. M.S. and B.B. performed disease heritability analysis. E.E.C. and E.J.H. provided prenatal brain tissue samples and technical advice for tissue processing. S.G. assisted with Ribo-seq quality control analyses. E.E.D., B.T.K., B.F., E.C.G. and M.E.G. drafted the manuscript, with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Linnaea Ostroff and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Ribosome profiling captures active translation in the human adult and prenatal brain, Related to Fig. 1.
(a) Pie chart displaying the fraction of raw sequence reads derived from tRNA, ribosomal RNA (rRNA), mitochondrial RNA (mtRNA), and remaining aligned reads (clean) from human adult and prenatal brain RNA-seq and Ribo-seq. (b) Beeswarm plot of sequenced ribosome footprint lengths across all 73 adult and prenatal brain samples. Red line indicates the average percentage of Ribo-seq reads assigned to a given read length across all samples. (c) Bar plot of the percentage of reads mapping to the coding sequence (CDS) and untranslated regions (5′ and 3′ UTR) of annotated protein-coding genes (Refseq hg38). Each bar represents an individual sample. (d) Bar plot of the number of ORFs identified by RibORF in each sample after filtering (see Methods). Notably, we identify many more ORFs in the prenatal brain compared to the adult brain, which is likely at least in part a result of the longer post-mortem interval in adult compared to prenatal samples. (e) Pie chart of ORF types detected in our study as well as four previous studies4,11,12,13. Because each study defines ORF types differently, ORFs are in hues denoting similar ORF types. Blue = CDS ORFs including non-canonical, out-of-frame ORFs; pink = uORFs and overlapping uORFs; orange = dORFs; yellow = ORFs translated from previously annotated non-coding RNAs. (f-g) PCA analysis of all genes in the human brain (f) RNA-seq and (g) Ribo-seq, colored by sample type (adult vs prenatal), post-mortem interval for adult samples, adult age, prenatal age (pcw), sex, and read depth (based on DESeq2 scale factors of estimated library size). The validity of combining samples into two groups in subsequent analyses was confirmed by the finding that these two groups were well separated by PCA analysis for both the transcriptome and translatome. (h) PCA analysis of batches of samples processed for RNA-seq (left) and Ribo-seq (right) to test for batch effect. We note a mild batch effect in some cases. To address the remaining batch effects in our differential expression analysis, we employed DTEG, an algorithm that uses DESeq2 to normalize across samples and includes batch correction. (i) Linear regression between postmortem tissue RNA integrity number (RIN) and read depth for RNA-seq and Ribo-seq. There is no significant correlation between the two variables (two-sided significance test for linear regression, p > 0.05 for RNA-seq and Ribo-seq). Gray shading = 95% CI. (j) Stacked bar plot of ORF types distributed by translation probability value, as calculated by RibORF. While RibORF uses a translation probability cutoff of 0.7 to determine significantly translated ORFs, we detect novel ORFs with a large range of translation probabilities.
Extended Data Fig. 2 Transcriptional and translational regulation across human brain development, Related to Fig. 2.
While the translatome has not been previously characterized in the developing human brain, our measurements of the transcriptome are consistent with published gene expression data from the BrainSpan Atlas of the Developing Human Brain15. (a) Bulk gene expression deconvolution of adult and prenatal brain RNA-seq using multiple single-cell RNA sequencing references (SCDC)61. Differences in cell type composition are not statistically significant by two-way ANOVA (F-statistic = 0, P-value = 1), strongly suggesting that observed differences in transcription and translation between prenatal and adult brain samples are not driven by differences in cell type composition. Data are shown as median ± IQR (whiskers = 1.5*IQR), notches indicate median + /−1.58*IQR/sqrt(n), n = 43 (adult) and 30 (prenatal) biologically independent tissues. (b) We sought to benchmark the Ribo-seq data in human postmortem brain tissue in this study to a recently published dataset of Ribo-seq in the human cerebellum70. PCA analysis of human prenatal and adult Ribo-seq (this study), as well as Ribo-seq of human adult cerebellum from Wang et al70. (c) Diagram of overlapping ORFs identified from human prenatal and adult Ribo-seq (this study), as well as Ribo-seq of human adult cerebellum from Wang et al70. Overall, we found a similar level of overlap between the Wang data and our human brain tissue samples as we found with our human NGN2 neuron samples. We do not find this limited overlap surprising, however, given that the Wang data is from adult cerebellum whereas we sample dorsolateral prefrontal cortex across a variety of ages. (d) Dot plot of the top enriched GO terms in each regulatory category defined in Fig. 2B. (e) Heatmap of RNA-seq expression (row-normalized) for all ribosomal genes in Fig. 2D across all human adult and prenatal samples in this study (left) and in the dorsolateral prefrontal cortex from the BrainSpan Atlas of the Developing Human Brain15. (f) Genomic locus of SUPT5H, a transcript regulated only at the level of translation, and (g) CNTNAP1, a transcript regulated only at the level of transcription. Tracks represent merged and depth-normalized reads across all adult vs. prenatal samples for RNA-seq, Ribo-seq, as well as P-site positions. Box and whisker plots indicate DESeq2-normalized RNA-seq reads and Ribo-seq P-sites in adult vs. prenatal samples. (f) **** = Ribo-seq padj = 2.26*10−4 by DESeq2. (g) **** = RNA-seq padj = 3.74*10−12 by DESeq2; **** = Ribo-seq padj = 1.35*10−6 by DESeq2. (f-g) Data are shown as median ± IQR (whiskers = 1.5*IQR), n = 43 (adult) and 30 (prenatal) biologically independent tissues. (h) Box and whisker plots of DESeq2-normalized RNA-seq reads across human brain samples divided into five age categories (prenatal = 12-18 pcw and 19-24 pcw, adult = 20-39 yrs, 40-59 yrs, >60 yrs) for CNTNAP1 (top), a transcriptionally regulated gene, and SUPT5H (bottom), a translationally regulated gene. (i) Box and whisker plots of DESeq2-normalized Ribo-seq reads across human brain samples divided into five age categories (prenatal = 12-18 pcw (n = 8 biologically independent tissues) and 19-24 pcw (n = 22 biologically independent tissues), adult = 18-39 yrs (n = 18 biologically independent tissues), 40-59 yrs (n = 23 biologically independent tissues), >60 yrs (n = 2 biologically independent tissues)) for CNTNAP1 (left) and SUPT5H (right). (h–i) Data are shown as median ± IQR (whiskers = 1.5*IQR). (j) Line plot of RPKM values from the BrainSpan Atlas of the Developing Human Brain for CNTNAP1 (top) and SUPT5H (bottom) across development in the dorsolateral prefrontal cortex. Data are shown as mean ± SD, n = 1 (8, 9, 17, 19, 21, 24, 26 & 37 pcw; 0.83, 1, 2, 33, 4, 11, 13, 18, 19, 21, 30, 36, 37, 40 yrs), 2 (0.33, 8 yrs), 3 (12, 13, 16 pcw) biologically independent tissues.
Extended Data Fig. 3 Microprotein expression and validation across brain development, Related to Fig. 3.
(a) Violin plot of average ribosome density (RD) by ORF type. Previously described ORFs are shown in red. Average ribosome density is shown in blue. (b) Venn diagram of sORFs detected in human brain (this study), human heart (van Heesch et al.4), or the sORF.org database. In total, 6,071 translated sORFs identified in the human brain perfectly matched the amino acid sequence of a previously reported entry in the sORFs.org database or identified in the human heart, a degree of overlap consistent with prior studies4. (c) Out of 8,590 lincRNA genes expressed across all brain samples, 415 lincRNA genes encode at least one translated ORF. We examined possible differences between ORF-encoding lincRNAs and non-translated lincRNAs. Box and whisker plots of annotated lincRNA features (expression, length, RPKM, and conservation) comparing RNAs that contain at least one ORF in the human brain to lincRNAs that do not contain any ORFs, data are shown as median ± IQR (whiskers = 1.5*IQR), n = 8,175 (no ORF detected) and 415 (ORF detected) lincRNAs. ** p = 0.009079 by two-sided Welch two-sample T-test, n.s. = not significant. (d) Number and type of ORFs identified by size-selection proteomics in the adult and prenatal brain, or by Johnson et al.25. (e) Histograms of number of proteins identified by size-selection proteomics in the adult and prenatal brain, or by Johnson et al.25., binned by protein length. (f) Box and whisker plots of Ribo-seq TPM for all ORFs detected by MS and ORFs not detected, data are shown as median ± IQR (whiskers = 1.5*IQR), n = 352 (adult, Johnson et al.), 3331 (adult, this study), 419 (prenatal, this study), 168,085 (not detected by MS). (g) Box and whisker plots of Ribo-seq TPM for sORFs detected by MS and ORFs not detected. (f-g) data are shown as median ± IQR (whiskers = 1.5*IQR), n = 14 (adult, Johnson et al.), 17 (adult, this study), 31 (prenatal, this study), 16838,125085 (not detected by MS). *** p = 2.92*10−5, **** p < 2.2*10−16, by two-sided Kolmogorov–Smirnov test. While only a fraction of the sORFs identified by ribosome profiling were detected by our mass spectrometry analysis, this is not surprising given that such shotgun proteomic approaches have low sensitivity for the detection of individual proteins, particularly if the proteins are transient or low in abundance. Consistent with this finding, the sORF-encoded proteins that we were able to detect by proteomics exhibited a higher average ribosome density compared to all sORFs detected by ribosome profiling.
Extended Data Fig. 4 Activity-dependent translation in hESC-derived neurons, Related to Fig. 4.
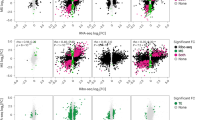
(a) Heatmap of RNA-seq TPM from hESC-derived neuronal cultures for marker genes associated with neuronal and non-neuronal cell types. Rows indicate individual samples and biological replicates. The pattern of gene expression observed in these cultures largely mimics the findings in Nehme et al.29. (b) Heatmap of pairwise Spearman’s r correlation between the top 2,000 expressed genes in RNA-seq samples from NGN2-derived neurons, including cells that were treated with cycloheximide alone (no treatment), harringtonine, or a vehicle control (DMSO). (c) Heatmap of pairwise Spearman’s r correlation between the top 2,000 expressed genes in Ribo-seq samples from NGN2-derived neurons, including cells that were treated with cycloheximide alone (no treatment), harringtonine, or a vehicle control (DMSO). (d) Bar chart of normalized RNA-seq and Ribo-seq reads mapping to FOS, a classic activity-induced gene in neurons, and XIRP1, an activity-induced gene that was previously reported in human GABAergic neuron cultures71. Data are shown as mean ± SD, n = 3 independent cell differentiations. (e) Bar plot of ORF types detected by Ribo-seq in NGN2 neurons, separated based on whether or not the exact ORF was also detected in postmortem adult or prenatal brain tissue. Most canonical ORFs detected in NGN2 neurons are also detected in human postmortem brain tissue. Likewise, we detect substantial, albeit somewhat lower, overlap for a range of non-canonical ORFs across both sample types as well. We are not surprised at the extent of the overlap between these datasets, as NGN2 neuronal cultures largely mimic embryonic cortical excitatory neurons; while cortical excitatory neurons represent a major constituent cell type in the dorsolateral prefrontal cortex, many other cell types exist in this tissue, and a number of the candidate sORFs/noncanonical ORFs (for example PLN, SLN, DWORF) and ncRNAs exhibit cell type specificity in the RNAs from which they are translated. Therefore, it is expected that we would not detect all of the same ORFs between NGN2 neurons and human postmortem tissue. (f) Importantly, the nature of the overlapping ORF population does not simply reflect the likelihood of ORF detection in brain samples. In this regard, we categorized ORFs detected in adult and/or prenatal postmortem tissue based on the number of samples in which the identical ORF was detected, and binned ORFs in each category by whether they were also detected in NGN2 neurons. Stacked bar chart of ORF types detected by Ribo-seq in human postmortem tissue, separated based on the number of tissue samples in which a given ORF was detected (2, 3-5, 6-10, 11-20, 21-50, 50-73) and whether the ORF was also detected in NGN2 neurons. This analysis did not find a notable difference in the proportion of canonical or various non-canonical ORFs based on whether they were detected in NGN2 neurons. (g) Stacked bar plot of start codon usage by ORF type in NGN2 neurons. (h) Metagene plot of Ribo-seq P-sites across the gene body of annotated Refseq ORFs in 3 independent replicates of harringtonine-treated (+ Harr) NGN2 neurons or vehicle-treated controls (- Harr). The enrichment for P-sites near start codons in + Harr samples is a hallmark of translational stalling induced by harringtonine treatment. (i) Number of ORFs in which the start codon was confirmed in harringtonine-treated NGN2-treated neurons. Two independent computational pipelines were used, ORF-RATER60 and RiboTISH31. While ORF-RATER validates fewer ORFs compared to RiboTISH, this result is expected as ORF-RATER utilizes a different, more stringent filtering process, consistent with our finding that nearly all ORFs validated by ORF-RATER are also validated by RiboTISH. A full list of ORFs validated by ORF-RATER and RiboTISH can be found in Supplementary Table 3. (j&k) Activity-dependent changes in ORF translation were largely driven by transcriptional changes rather than a shift in ribosome density for both canonical ORFs and sORFs. This finding is consistent with observations that activity-dependent translation events coupled to transcription are transient and likely return to basal levels within six hours of membrane depolarization. (j) Scatterplot of fold-changes between stimulated and unstimulated neurons for all canonical ORFs in Ribo-seq data and the corresponding gene in RNA-seq data. Transcriptionally regulated genes (blue), translationally regulated genes (red), buffered genes (light purple), and intensified genes (dark purple) are highlighted. (k) Scatterplot of fold-changes between stimulated and unstimulated neurons for all sORFs in Ribo-seq data and the corresponding gene in RNA-seq data. Transcriptionally regulated genes (blue), translationally regulated genes (red), buffered genes (light purple), and intensified genes (dark purple) are highlighted. (l&m) Genomic loci of two activity-dependent ncRNAs with evidence of translation, MIR22HG (l) and LOC107986102 (m). Tracks represent merged and depth-normalized reads across 3 biological replicates of membrane-depolarized (6 h KCl) and unstimulated neurons for RNA-seq, Ribo-seq, as well as P-site positions for Ribo-seq and harringtonine-treated Ribo-seq. sORFs identified by RibORF are shown in gold.
Extended Data Fig. 5 Evolutionary origins of human brain sORFs, Related to Fig. 5.
(a) Bar plot of the number of sORFs (40-110 AA) grouped by evolutionary age. (b) Bar plot of the number of sORFs (40-100 AA) detected by mass spectrometry (See Fig. 3d) grouped by evolutionary age. We detected many evolutionarily conserved ORFs that were previously detected in other species. For example, uORFs translated from the 5′UTRs of FTH1 and CCNi show conservation within the mammalian lineage by phylostratigraphy and show experimental evidence of translation in rodents4,36. (c) Number and percentage of sORFs ≥40 AA that are translated from brain-enriched transcripts, grouped by evolutionary age. (d) Criteria for filtering TE insertion events at start codons (left) and pie chart of TE type for all ORFs in our dataset with a TE insertion at the start codon. (e) Histogram of ORF length for all ORFs encoded within ncRNAs, pseudogenes, and uORFs. Most ORFs encoded by ncRNAs, pseudogenes, and uORFs are sORFs.
Extended Data Fig. 6 Effects of uORF expression on downstream ORF translation, Related to Fig. 6.
(a) Scatterplot of fold-changes in translation between adult and prenatal brain for sORFs and canonical ORFs expressed from the same transcript. Positive values indicate enrichment in the adult brain, whereas negative values indicate enrichment in the prenatal brain. Red points indicate genes where sORF translation is significantly (DESeq2 padj < 0.05) enriched in prenatal brain while canonical ORF translation is significantly (DESeq2 padj < 0.05) enriched in adult brain. Blue points indicate genes where sORF translation is significantly (DESeq2 padj < 0.05) enriched in adult brain while canonical ORF translation is significantly (DESeq2 padj < 0.05) enriched in prenatal brain. While most sORFs exhibited concordant translation with their associated canonical ORFs across development, we identified 50 sORFs that were discordant with nearby canonical ORF translation, and these discordant sORFs were strongly enriched for uORFs translated from 5′UTRs of annotated protein-coding genes. (b) Stacked bar plot of numbers and percentages of sORFs detected in human brain (all sORFs), or sORFs exhibiting oppositely regulated expression across development compared to a canonical ORF translated from the same gene, separated by sORF type. (c) Scatterplot of fold-changes in translation between adult and prenatal brain for sORFs and canonical ORFs expressed from the same transcript, separated by type of ORF. Positive values indicate enrichment in the adult brain, whereas negative values indicate enrichment in the prenatal brain. Red points indicate genes where sORF translation is significantly (DESeq2 padj < 0.05) enriched in prenatal brain whereas canonical ORF translation is significantly (DESeq2 padj < 0.05) enriched in adult brain. Blue points indicate genes where sORF translation is significantly (DESeq2 padj < 0.05) enriched in adult brain whereas canonical ORF translation is significantly (DESeq2 padj < 0.05) enriched in prenatal brain. (d) Genomic locus of DLGAP1. Tracks represent merged and depth-normalized reads across all adult vs. prenatal samples for RNA-seq, Ribo-seq, as well as P-site positions. The sORF identified by RibORF is shown in gold.
Extended Data Fig. 7
Microprotein functional characterization, Related to Fig. 7. (a) Protein functions of known genes that share homology with sORFs. (b) Characterization of sORFs that overlap with a domain in the PFam database. (c) Line plot of sequence complexity in the known human proteome (Known Uniprot), the disordered proteome (Known IDRs), and sORFs detected in human postmortem tissue that do not share homology with known protein domains. (d) Line plot of the log2 ratio of arginine to lysine in the categories of ORFs described in c. (e) Line plot of the proportion of aromatic amino acids (phenylalanine, tyrosine, and tryptophan) in the categories of ORFs described in c. (f) Line plot of the isoelectric point for the categories of ORFs described in c. (g) Heatmap and hierarchical clustering of z-scores for 109 sequence and physicochemical features associated with the known disordered proteome as well as all sORFs that do not contain a BlastP hit and do not overlap with annotated ORFs. Boxes to the right of the heatmap indicate clusters of IDRs with similar properties. Yellow = clusters significantly enriched for sORFs.
Supplementary information
Supplementary Table 1
Full demographic and Ribo-seq quality metrics. Legend included in file as Tab 1. Columns describe the following metrics: Sample ID (adult or prenatal sample ID); Age (adult age in years, prenatal age in post-conception weeks (pcw)); Postmortem Interval (adult postmortem interval); Sex; % Alignment RNA-seq (percentage of RNA-seq reads that aligned to hg38 genome after filtering rRNA, tRNA and mtRNA); % rRNA Ribo-seq (percentage of Ribo-seq reads that aligned to hg38 rRNA); % Alignment Ribo-seq (percentage of Ribo-seq reads that aligned to hg38 genome after filtering rRNA, tRNA and mtRNA); 28-nt % reads in frame 1 (percentage of Ribo-seq reads (28-nt length) that mapped to frame 1 after offset correction); 29-nt % reads in frame 1 (percentage of Ribo-seq reads (29-nt length) that mapped to frame 1 after offset correction); 30-nt % reads in frame 1 (percentage of Ribo-seq reads (30-nt length) that mapped to frame 1 after offset correction); 31-nt % reads in frame 1 (percentage of Ribo-seq reads (31-nt length) that mapped to frame 1 after offset correction); % reads mapping to annotated CDS; percentage of Ribo-seq reads mapping to annotated 5′ UTR; percentage of Ribo-seq reads mapping to annotated 3′ UTR; Total number of ORFs identified by RibORF.
Supplementary Table 2
ORFs validated by proteomics. Columns describe the following metrics: orfID (ORF ID assigned by RibORF); chrom (chromosome); codon5 (genomic position of ORF start codon); codon3 (genomic position of ORF stop codon); readNum (number of P-sites within ORF); f1 (number of P-sites in frame 1); f2 (number of P-sites in frame 2); f3 (number of P-sites in frame 3); entropy (entropy value of read distribution); MAXentropy (maximum entropy value of randomized distribution); PME (percentage of maximum entropy value); codonNum (number of codons with sequencing reads); f1max (fraction of codons with 1st nucleotides containing more reads than 2nd and 3rd); DNA_Sequence (DNA sequence within ORF exons); Peptide Sequence (ORF protein sequence); type (ORF type); Peptide_Length (ORF length (aa)); DNA_Length (ORF DNA length across exons); SpecificTypes (annotated RNA type in lncRNAKB annotation); ENST_IDs (Ensembl ID of gene encoding ORF); maxp (maximum translation probability across all adult and prenatal samples); numsamples (number of adult/prenatal samples in the ORF was detected).
Supplementary Table 3
ORFs independently validated in NGN2 neurons by harringtonine treatment. Columns describe the following metrics: orfID ORF identifier assigned by RibORF; type (ORF type); start.codon (ORF start codon); chrom (chromosome); strand (positive or negative DNA strand); codon5 (genomic position of ORF start codon); codon3 (genomic position of ORF stop codon); RiboTISH (was ORF start codon validated by the RiboTISH pipeline?); ORFrater (was ORF start codon validated by ORF-RATER pipeline?).
Supplementary Table 4
Paralogs of human brain sORFs. Columns describe the following metrics: orfID (ORF identifier assigned by RibORF); orf_gene_name (ORF identifier assigned by RibORF); orf_gene_description (full gene description of ORF); paralog.protID (Ensembl protein identifier for paralog); paralog.transcriptID (Ensembl transcript identifier for paralog); paralog.geneID (gene identifier for paralog); paralog.gene.name (gene name for paralog); paralog_gene_description (full gene description of paralog).
Supplementary Table 5
Physicochemical analysis of human brain sORFs. The ‘Feature Comparisons’ table describes the mean and standard deviation (s.d.) of molecular characteristics in sORFs >20 aa and all proteins in UniProt. The molecular features are fully described in Zarin et al.51. Table 1 (https://cdn.elifesciences.org/articles/46883/elife-46883-supp1-v4.docx). The composition of the feature clusters in Fig. 7c are described in ‘Cluster Lists’. The molecular features are listed in row 1 (K-rich, acidic, N-rich, Q-rich, S-rich, aliphatic, hydrophobic, aromatic, P-rich, G-rich, RGG, R-rich and RG-repeats). The paired ‘id’ and ‘name’ columns list the proteins comprising each cluster. The ‘id’ columns list the UniProt accession ID (or lncRNAKB ID for sORFs). The ‘name’ columns list the full protein name or, in the case of sORFs, the source of tissue where the sORF was identified (fetal, adult or both).
Supplementary Table 6
Sequence information for sORF expression constructs. Legend included in file as Tab 1. Columns describe the following metrics: ORF ID (open reading frame identifier); Gene Name (gene name corresponding to the ORF); Length (aa) (length of the ORF in amino acids); Category (rationale for inclusion in the experiment); Protein sequence (protein sequence of the ORF); DNA sequence (DNA sequence of the ORF).
Source data
Source Data Fig. 7
Unprocessed western blots
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Duffy, E.E., Finander, B., Choi, G. et al. Developmental dynamics of RNA translation in the human brain. Nat Neurosci 25, 1353–1365 (2022). https://doi.org/10.1038/s41593-022-01164-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-022-01164-9
This article is cited by
-
FMRP-mediated spatial regulation of physiologic NMD targets in neuronal cells
Genome Biology (2024)
-
Small open reading frames: a comparative genetics approach to validation
BMC Genomics (2023)
-
Evolution and implications of de novo genes in humans
Nature Ecology & Evolution (2023)
-
The status of the human gene catalogue
Nature (2023)
-
Celf4 controls mRNA translation underlying synaptic development in the prenatal mammalian neocortex
Nature Communications (2023)