Abstract
The organization of the basic tissue and functional properties of the cerebellum across development is unknown. Combining several large datasets, we demonstrate in the human cerebellum a static tissue gradient in adults that mirrors a similar growth-rate gradient across development. Quantitative tissue metrics corroborate unique densities of certain lipids and proteins among lobules, and cerebellar structural development closely follows cerebellar functional properties through childhood.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All data belonging to the HCP are available online at the following persistent weblinks: https://www.humanconnectome.org or https://nda.nih.gov. Users must first be registered for data access. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD034848. Data and code to reproduce developmental qMRI figures can be found at: https://github.com/gomezj/nn_cb_figure_code.
Code availability
The HCP data were preprocessed using the HCP-Pipeline analyses (https://github.com/Washington-University/HCPpipelines). The Statistical Parametric Mapping (SPM12) (https://www.fil.ion.ucl.ac.uk/spm/software/spm12/), the SUIT (https://github.com/jdiedrichsen/suit) and the ACAPULCO (https://gitlab.com/shuohan/acapulco) were further used to process the HCP data regarding the cerebellum. The mrQ toolbox (https://github.com/mezera/mrQ) was used to analyze the qMRI images. Custom code for gradient analysis can be found at https://github.com/BNUCNL/cerebellum_grad_dev. Proteome Discoverer v.2.5.0 suite and its Sequest HT algorithms were used to search the mass spectroscopy raw files. Scaffold v.5.0.1 was used to validate identify proteins and export their density values for plotting. Protein enrichment analysis was conducted within the ToppGene Suite (https://toppgene.cchmc.org).
References
Moore, D. M., D’Mello, A. M., McGrath, L. M. & Stoodley, C. J. The developmental relationship between specific cognitive domains and grey matter in the cerebellum. Dev. Cogn. Neurosci. 24, 1–11 (2017).
King, M., Hernandez-Castillo, C. R., Poldrack, R. A., Ivry, R. B. & Diedrichsen, J. Functional boundaries in the human cerebellum revealed by a multi-domain task battery. Nat. Neurosci. 22, 1371–1378 (2019).
Fatemi, S. H. et al. Consensus paper: pathological role of the cerebellum in autism. Cerebellum 11, 777–807 (2012).
D’Mello, A. M. & Stoodley, C. J. Cerebro-cerebellar circuits in autism spectrum disorder. Front Neurosci. 9, 408 (2015).
Wang, S. S.-H., Kloth, A. D. & Badura, A. The cerebellum, sensitive periods, and autism. Neuron 83, 518–532 (2014).
Burt, J. B. et al. Hierarchy of transcriptomic specialization across human cortex captured by structural neuroimaging topography. Nat. Neurosci. 21, 1251–1259 (2018).
Wagstyl, K., Ronan, L., Goodyer, I. M. & Fletcher, P. C. Cortical thickness gradients in structural hierarchies. Neuroimage 111, 241–250 (2015).
Gomez, J. et al.Microstructural proliferation in human cortex is coupled with the development of face processing. Science 355, 68–71 (2017).
Palay, S. & Chan-Palay, V. Cerebellar cortex: cytology and organization (Springer, 1974).
Van Essen, D. C. et al. The WU-Minn human connectome project: an overview. Neuroimage 80, 62–79 (2013).
Diedrichsen, J., Balsters, J. H., Flavell, J., Cussans, E. & Ramnani, N. A probabilistic MR atlas of the human cerebellum. Neuroimage 46, 39–46 (2009).
Han, S., Carass, A., He, Y. & Prince, J. L. Automatic cerebellum anatomical parcellation using U-Net with locally constrained optimization. Neuroimage 218, 116819 (2020).
Klein, A. P., Ulmer, J. L., Quinet, S. A., Mathews, V. & Mark, L. P. Nonmotor functions of the cerebellum: an introduction. AJNR Am. J. Neuroradiol. 37, 1005–1009 (2016).
Glasser, M. F. & Van Essen, D. C. Mapping human cortical areas in vivo based on myelin content as revealed by T1- and T2-weighted MRI. J. Neurosci. 31, 11597–11616 (2011).
Flechsig, P. Developmental (myelongenetic) localisation of the cerebral cortex in the human subject. Lancet 158, 1027–1030 (1901).
Elston, G. N. & Fujita, I. Pyramidal cell development: postnatal spinogenesis, dendritic growth, axon growth, and electrophysiology. Front. Neuroanat. 8, 78 (2014).
Buckner, R. L., Krienen, F. M., Castellanos, A., Diaz, J. C. & Yeo, B. T. T. The organization of the human cerebellum estimated by intrinsic functional connectivity.J. Neurophysiol 106, 2322–2345 (2011).
Guell, X., Schmahmann, J. D., Gabrieli, J. & Ghosh, S. S. Functional gradients of the cerebellum. eLife 7, e36652 (2018).
Mezer, A. et al. Quantifying the local tissue volume and composition in individual brains with magnetic resonance imaging. Nat. Med. 19, 1667–1672 (2013).
Liu, J. et al. Impaired adult myelination in the prefrontal cortex of socially isolated mice. Nat. Neurosci. 15, 1621–1623 (2012).
Etxeberria, A. et al. Dynamic modulation of myelination in response to visual stimuli alters optic nerve conduction velocity. J. Neurosci. 36, 6937–6948 (2016).
Zuo, X. N. et al. The oscillating brain: complex and reliable. Neuroimage 49, 1432–1445 (2010).
Zou, Q. H. et al. An improved approach to detection of amplitude of low-frequency fluctuation (ALFF) for resting-state fMRI: fractional ALFF. J. Neurosci. Methods 172, 137–141 (2008).
Barton, R. A. & Venditti, C. Rapid evolution of the cerebellum in humans and other great apes. Curr. Biol. 24, 2440–2444 (2014).
Glasser, M. F. et al. The minimal preprocessing pipelines for the Human Connectome Project. NeuroImage 80, 105–124 (2013).
Somerville, L. H. et al. The Lifespan Human Connectome Project in Development: A large-scale study of brain connectivity development in 5–21 year olds. NeuroImage 183, 456–468 (2018).
Harms, M. P. et al. Extending the Human Connectome Project across ages: Imaging protocols for the Lifespan Development and Aging projects. NeuroImage 183, 972–984 (2018).
Diedrichsen, J. A spatially unbiased atlas template of the human cerebellum. NeuroImage 33, 127–138 (2006).
Diedrichsen, J. & Zotow, E. Surface-based display of volume-averaged cerebellar imaging data. PLoS ONE 10, e0133402 (2015).
Falconer, D. S. & Mackay, T. Introduction to quantitative genetics. (Pearson, Prentice Hall, 2009)
Jacquard, A. Heritability: one word, three concepts. Biometrics 39, 465 (1983).
Eng, J. K., McCormack, A. L. & Yates, J. R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Nesvizhskii, A. I., Keller, A., Kolker, E. & Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem 75, 4646–4658 (2003).
Acknowledgements
HCP-YA data were obtained from ConnectomeDB and HCP-D data were obtained from the National Institute of Mental Health (NIMH) Data Archive (NDA, Collection #2846). This manuscript reflects the views of the authors and may not reflect the opinions or views of the NIMH or of the submitters contributing original data to NDA. We thank S. Kyin and H. Shwe of the Proteomics and Mass Spectroscopy Facility at Princeton University for their technical expertise, and we thank X. Chen and T. Huang for helping with HCP-D data analysis. We thank the Brain Development Lab for several useful discussions. This research was supported by Child Brain-Mind Development Cohort Study in the China Brain Initiative (2021ZD0200534) and the National Natural Science Foundation of China (31771251) to Z.Z. This research was supported by start-up funds from the Princeton Neuroscience Institute to J.G. This research was supported by National Science Foundation Graduate Research Fellowship (2021314805) to F.d'O.U.
Author information
Authors and Affiliations
Contributions
J.G. and Z.Z. conceived of the idea and design of the study. X.L. compiled the data, performed the analyses, and prepared visualizations. F.d'O.U. performed the analyses. A.N.V. provided expertise and postmortem tissue samples. J.G., X.L. and Z.Z. drafted and revised the manuscript with input from all authors. J.G. and Z.Z. supervised the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Ye Tian and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Analyzing T1w/T2w ratio from medial to lateral bins of the cerebellum.
Within each individual, a cerebellar hemisphere was split into six bins from medial to lateral based on the percentage of slices to normalize inter-individual differences in cerebellar volume. Data are averaged across hemispheres. Within each bin, median values are extracted from each lobule and colored according to the inset cartoonized cerebellum. While the S-shaped gradient reported in the main finding can be observed within each bin (colored box plots are well differentiated in each bin), there is little variation along the medial-lateral axis. For each box subplot, center: median value; box: interquartile range; whiskers: 1.5 times of interquartile range.
Extended Data Fig. 2 The structural gradient and its development replicated with a cerebellar segmentation built specifically for children.
(a) ACAPULCO neural network trained with pediatric data to segment lobules I-V to X shows similar results to the full ACAPULCO network: An S-shaped gradient with lower T1w/T2w ratio values in dorsal lobules like Crus I, replicating the developmental gradient with dorsal lobules showing the most pronounced development. Bold line denotes linear fit with shaded 95% confidence intervals. Reported F-test and two-sided P value result from an ANOVA with factors of age and lobule. (b) Slopes (% annual change in T1w/T2w ratio) from data shown in part a. Asterisks denotes slope significantly (two-sided P < 0.05 with FDR correction, n = 519 participants) different from zero in linear regression of predicting ratio value with age. (c) The T1w/T2w ratio across the lobules of the developmental HCP dataset binned by ages shown in the inset legend. The gradient is more pronounced in younger children (blue) compared to adults (red). (d) The interlobular heterogeneity (standard deviation across T1w/T2w ratio across an individual’s lobules) decreases with development. Bold line denotes linear fit with shaded 95% confidence intervals. The resulting P value from the linear model is two-sided.
Extended Data Fig. 3 A myelin gradient still emerges along the sensory-association gradient even when defined functionally from cortical connectivity.
(a) T1w/T2w values from the young adult HCP participants within a functional parcellation defined from the Bucker 7-network resting-state functional connectivity atlas (each network is assigned an arbitrary color). A given voxel within the cerebellum is assigned to one of the seven cortical networks with which it is most correlated. All but the limbic network are shown, to which a disproportionately small number of cerebellar voxels were assigned. The median T1w/T2w ratio value was extracted across these functionally-defined regions of the cerebellum from each participant. T1w/T2w values are significantly different from one another. In this analysis, a functionally-defined region of interest (ROI) can span more than one lobule. Sensory networks, such as Vision and Somatomotor, are the most myelinated compared to association-related regions such as the Ventral Attention and Frontoparietal networks. The inner box indicates interquartile range and the central dot indicates median value. Reported F-test and two-sided P value result from an ANOVA with main factor of functional network. (b) Similar to Fig. 1g, the slope (annual rate of change) was extracted from linear regressions (n = 1014 participants) across all developmental HCP dataset participants within a given parcel (for example, one of the 6 networks). Asterisks denote that the rate-of-change is significantly different from zero. Qualitatively, functionally-defined networks show differential development of T1w/T2w ratio, largely replicating the main results that association-related ROIs (Dorsal and Ventral Attention, green and pink respectively) show the most protracted development compared to sensory regions (like Vision and Somatomotor networks, in purple and blue respectively). Although it is worth noting that an association network such as the Default Network in red shows less development than potentially expected, but given that this network spans both anterior and posterior lobules, the averaged structural development is less extreme given the opposing developmental trajectories of these different lobules. *two-sided P < 0.05 FDR corrected.
Extended Data Fig. 4 Gradients of T1w/T2w ratio and fALFF map differentially onto, but are largely unique from, previously observed adult functional gradients of the cerebellum.
(a) Left column, flatmaps of the two orthogonal gradients previously observed from Guell and colleagues. Right Column, flatmaps of the fALFF and T1w/T2w ratio from the current data. (b) Left column, R-squared values from regressions performed in individuals relating either fALFF (top) or T1w/T2w (bottom) maps to either Gradient 1 (blue) or Gradient 2 (pink). The fALFF map has more variance explained by Gradient 1, while the T1w/T2w map is explained more by Gradient 2 and shows a decreasing relationship with development. Center: median value; box: interquartile range; whiskers: 1.5 times of interquartile range; fliers: outliers. Right column, coefficients from a regression using Gradient 1 and Gradient 2 to explain either fALFF (top) or T1w/T2w (bottom) values within individuals. Coefficients are shown as a ratio compared to their values in the oldest age group (19 years old). For T1w/T2w, Gradient 2 shows higher betas compared to Gradient 1, and these betas decrease with development. Shaded regions indicate 95% confidence intervals.
Extended Data Fig. 5 Data-driven analysis replicates the T1w/T2w ratio lobular gradient across the cerebellum.
(a) A spatial PCA that uses participants as features and voxels as samples to discover a set of orthogonal PC axes by applying a linear transformation on the original set of features and then projecting each sample on these axes to form unrelated spatial maps. That is, PCA decomposes spatial maps into unrelated spatial PCs, which are sorted in decreasing order according to its contribution in explaining the total variance. To simultaneously discover the principal gradient and characterize its development, we performed the PCA on the developmental HCP dataset. (b) Weights from the first PC show to what extent participants in a given age-bin contribute to the PC. PC1 shows decreasing weights with development, suggesting the spatial gradient identified by PC1 becomes less pronounced with development, similar to the main finding of decreasing interlobular heterogeneity in T1w/T2w ratio. Error bars indicate 95% confidence intervals. Individual participants plotted in gray circles. (c) PCA allows for the identification of gradients (for example, spatial PCs) across space (lobules). On the left, the lobular values of PC1 replicate the adult S-shaped structural gradient reported in Fig. 1c which is shown here on the right for ease of comparison. The inner box of the violin plot indicates interquartile range and the center dot indicates median value.
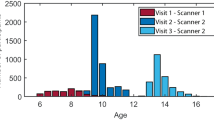
Extended Data Fig. 6 Sample distribution of age and sex.
Participants with ages 9 to 19 years old from HCP-D dataset were included in the study. Stacked bar plots show how many participants in a given age-bin are female (light blue) or male (dark blue).
Supplementary information
Supplementary Information
Supplementary Table 1.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, X., d’Oleire Uquillas, F., Viaene, A.N. et al. A multifaceted gradient in human cerebellum of structural and functional development. Nat Neurosci 25, 1129–1133 (2022). https://doi.org/10.1038/s41593-022-01136-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-022-01136-z
This article is cited by
-
Population-wide cerebellar growth models of children and adolescents
Nature Communications (2024)
-
Population imaging cerebellar growth for personalized neuroscience
Nature Communications (2024)
-
A multimodal submillimeter MRI atlas of the human cerebellum
Scientific Reports (2024)
-
Evaluation of histological and ultrastructural changes provoked by prenatal tramadol on postnatal cortical cerebellar neuronal development in rats: possible implication of Ki67, GFAP and MicroRNA-7/P53 signalling trajectories
Journal of Molecular Histology (2024)
-
Phylogenetic comparative analysis of the cerebello-cerebral system in 34 species highlights primate-general expansion of cerebellar crura I-II
Communications Biology (2023)