Abstract
Progressive multiple sclerosis (MS) is characterized by unrelenting neurodegeneration, which causes cumulative disability and is refractory to current treatments. Drug development to prevent disease progression is an urgent clinical need yet is constrained by an incomplete understanding of its complex pathogenesis. Using spatial transcriptomics and proteomics on fresh-frozen human MS brain tissue, we identified multicellular mechanisms of progressive MS pathogenesis and traced their origin in relation to spatially distributed stages of neurodegeneration. By resolving ligand–receptor interactions in local microenvironments, we discovered defunct trophic and anti-inflammatory intercellular communications within areas of early neuronal decline. Proteins associated with neuronal damage in patient samples showed mechanistic concordance with published in vivo knockdown and central nervous system (CNS) disease models, supporting their causal role and value as potential therapeutic targets in progressive MS. Our findings provide a new framework for drug development strategies, rooted in an understanding of the complex cellular and signaling dynamics in human diseased tissue that facilitate this debilitating disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Spatial transcriptomics data have been deposited at Gene Expression Omnibus (GEO) with accession no. GSE174647. Note that a smaller subset of this dataset has been used in one of our previous studies18, where it was employed to test the circumscribed hypothesis that a specific T cell type is present in the brains of patients with progressive MS. As such, no further exploration of the dataset was performed then and all analyses and conclusions of the present study are completely independent. Mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository, accession no. PXD026529. Published scRNA-seq data of peripheral blood mononuclear cells from patients with MS and healthy individuals are available from GEO (no. GSE144744). SnRNA-seq data of human postmortem cortical brain tissue are available from GEO (no. GSE118257); the Sequence Read Archive (no. PRJNA544731); the GTEx portal (http://www.gtexportal.org/home/datasets, see additional data sets, Habib et al.); and the Allen brain atlas (https://portal.brain-map.org/atlases-and-data/rnaseq/human-m1-10x; https://portal.brain-map.org/atlases-and-data/rnaseq/human-multiple-cortical-areas-smart-seq). Literature-curated ligand–receptor pairings are available from https://www.cellphonedb.org/ (v.2.0). Bulk RNA-seq data of human postmortem samples from 54 tissues are available from https://www.gtexportal.org/home/ (GTEX V8).
References
Dendrou, C. A., Fugger, L. & Friese, M. A. Immunopathology of multiple sclerosis. Nat. Rev. Immunol. 15, 545–558 (2015).
Tintore, M., Vidal-Jordana, A. & Sastre-Garriga, J. Treatment of multiple sclerosis—success from bench to bedside. Nat. Rev. Neurol. 15, 53–58 (2019).
Carassiti, D. et al. Neuronal loss, demyelination and volume change in the multiple sclerosis neocortex. Neuropathol. Appl. Neurol. 44, 377–390 (2018).
Scalfari, A. et al. The cortical damage, early relapses, and onset of the progressive phase in multiple sclerosis. Neurology 90, e2107–e2118 (2018).
Magliozzi, R. et al. A gradient of neuronal loss and meningeal inflammation in multiple sclerosis. Ann. Neurol. 68, 477–493 (2010).
Wegner, C., Esiri, M. M., Chance, S. A., Palace, J. & Matthews, P. M. Neocortical neuronal, synaptic, and glial loss in multiple sclerosis. Neurology 67, 960–967 (2006).
Ontaneda, D., Fox, R. J. & Chataway, J. Clinical trials in progressive multiple sclerosis: lessons learned and future perspectives. Lancet Neurol. 14, 208–223 (2015).
Kappos, L. et al. Siponimod versus placebo in secondary progressive multiple sclerosis (EXPAND): a double-blind, randomised, phase 3 study. Lancet 391, 1263–1273 (2018).
Montalban, X. et al. Ocrelizumab versus placebo in primary progressive multiple sclerosis. N. Engl. J. Med. 376, 209–220 (2017).
Longo, S. K., Guo, M. G., Ji, A. L. & Khavari, P. A. Integrating single-cell and spatial transcriptomics to elucidate intercellular tissue dynamics. Nat. Rev. Genet. 22, 627–644 (2021).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Coscia, F. et al. A streamlined mass spectrometry-based proteomics workflow for large‐scale FFPE tissue analysis. J. Pathol. 251, 100–112 (2020).
Mann, M. The ever expanding scope of electrospray mass spectrometry—a 30 year journey. Nat. Commun. 10, 3744 (2019).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Jäkel, S. et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 566, 543–547 (2019).
Schirmer, L. et al. Neuronal vulnerability and multilineage diversity in multiple sclerosis. Nature 573, 75–82 (2019).
Kaufmann, M. et al. Identifying CNS-colonizing T cells as potential therapeutic targets to prevent progression of multiple sclerosis. Med (N Y) 2, 296–312 (2021).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Han, M. H. et al. Janus-like opposing roles of CD47 in autoimmune brain inflammation in humans and mice. J. Exp. Med. 209, 1325–1334 (2012).
Lisak, R. P. & Benjamins, J. A. Melanocortins, melanocortin receptors and multiple sclerosis. Brain Sci. 7, 104 (2017).
Maimone, D., Guazzi, G. C. & Annunziata, P. IL-6 detection in multiple sclerosis brain. J. Neurol. Sci. 146, 59–65 (1997).
McCandless, E. E. et al. Pathological expression of CXCL12 at the blood-brain barrier correlates with severity of multiple sclerosis. Am. J. Pathol. 172, 799–808 (2008).
Moll, N. M. et al. Imaging correlates of leukocyte accumulation and CXCR4/CXCL12 in multiple sclerosis. Arch. Neurol. 66, 44–53 (2009).
Trebst, C. et al. CCR1+/CCR5+ mononuclear phagocytes accumulate in the central nervous system of patients with multiple sclerosis. Am. J. Pathol. 159, 1701–1710 (2001).
Morrissey, M. A., Kern, N. & Vale, R. D. CD47 ligation repositions the inhibitory receptor SIRPA to suppress integrin activation and phagocytosis. Immunity 53, 290–302 (2020).
Rothlin, C. V., Ghosh, S., Zuniga, E. I., Oldstone, M. B. A. & Lemke, G. TAM receptors are pleiotropic inhibitors of the innate immune response. Cell 131, 1124–1136 (2007).
Cao, W. et al. Regulation of TLR7/9 responses in plasmacytoid dendritic cells by BST2 and ILT7 receptor interaction. J. Exp. Med. 206, 1603–1614 (2009).
Persson, G., Jørgensen, N., Nilsson, L. L., Andersen, L. H. J. & Hviid, T. V. F. A role for both HLA-F and HLA-G in reproduction and during pregnancy? Hum. Immunol. 81, 127–133 (2019).
Cardona, A. E. et al. Control of microglial neurotoxicity by the fractalkine receptor. Nat. Neurosci. 9, 917–924 (2006).
Carrera Silva, E. A. et al. T cell-derived protein S engages Tam receptor signaling in dendritic cells to control the magnitude of the immune response. Immunity 39, 160–170 (2013).
Michael, J., Marschallinger, J. & Aigner, L. The leukotriene signaling pathway: a druggable target in Alzheimer’s disease. Drug Discov. Today 24, 505–516 (2019).
Reis, E. S., Mastellos, D. C., Hajishengallis, G. & Lambris, J. D. New insights into the immune functions of complement. Nat. Rev. Immunol. 19, 503–516 (2019).
Meyer, R. C., Giddens, M. M., Coleman, B. M. & Hall, R. A. The protective role of prosaposin and its receptors in the nervous system. Brain Res. 1585, 1–12 (2014).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2022).
Dong, R. & Yuan, G.-C. SpatialDWLS: accurate deconvolution of spatial transcriptomic data. Genome Biol. 22, 145 (2021).
Elosua-Bayes, M., Nieto, P., Mereu, E., Gut, I. & Heyn, H. SPOTlight: seeded NMF regression to deconvolute spatial transcriptomics spots with single-cell transcriptomes. Nucleic Acids Res. 49, e50 (2021).
Chitu, V., Gokhan, Ş., Nandi, S., Mehler, M. F. & Stanley, E. R. Emerging roles for CSF-1 receptor and its ligands in the nervous system. Trends Neurosci. 39, 378–393 (2016).
Tang, W. et al. TRAIL receptor mediates inflammatory cytokine release in an NF-κB-dependent manner. Cell Res. 19, 758–767 (2009).
Nibbs, R. J. B. & Graham, G. J. Immune regulation by atypical chemokine receptors. Nat. Rev. Immunol. 13, 815–829 (2013).
Souma, T. et al. Context-dependent functions of angiopoietin 2 are determined by the endothelial phosphatase VEPTP. Proc. Natl Acad. Sci. USA 115, 201714446 (2018).
Zöller, T. et al. Silencing of TGFβ signalling in microglia results in impaired homeostasis. Nat. Commun. 9, 4011 (2018).
Migeotte, I. et al. Identification and characterization of an endogenous chemotactic ligand specific for FPRL2. J. Exp. Med. 201, 83–93 (2005).
Tang, W. et al. The growth factor progranulin binds to TNF receptors and is therapeutic against inflammatory arthritis in mice. Science 332, 478–484 (2011).
Wertz, M. H. et al. Genome-wide in vivo CNS screening identifies genes that modify CNS neuronal survival and mHTT toxicity. Neuron 106, 76–89 (2020).
Lehrman, E. K. et al. CD47 protects synapses from excess microglia-mediated pruning during development. Neuron 100, 120–134 (2018).
Toth, A. B. et al. Synapse maturation by activity-dependent ectodomain shedding of SIRPα. Nat. Neurosci. 16, 1417–1425 (2013).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Spiegel, I. et al. A central role for Necl4 (SynCAM4) in Schwann cell–axon interaction and myelination. Nat. Neurosci. 10, 861–869 (2007).
Golan, N. et al. Genetic deletion of cadm4 results in myelin abnormalities resembling Charcot-Marie-Tooth neuropathy. J. Neurosci. 33, 10950–10961 (2013).
Jolivalt, C. G., Vu, Y., Mizisin, L. M., Mizisin, A. P. & Calcutt, N. A. Impaired prosaposin secretion during nerve regeneration in diabetic rats and protection of nerve regeneration by a prosaposin-derived peptide. J. Neuropathol. Exp. Neurol. 67, 702–710 (2008).
Golde, T. E. The therapeutic importance of understanding mechanisms of neuronal cell death in neurodegenerative disease. Mol. Neurodegener. 4, 8 (2009).
Evans, S. R. et al. A randomized trial evaluating ProsaptideTM for HIV-associated sensory neuropathies: use of an electronic diary to record neuropathic pain. PLoS ONE 2, e551 (2007).
Taylor, E. M., Otero, D. A., Banks, W. A. & O’Brien, J. S. Retro-inverso prosaptide peptides retain bioactivity, are stable In vivo, and are blood-brain barrier permeable. J. Pharmacol. Exp. Ther. 295, 190–194 (2000).
Xie, Y. et al. FGF/FGFR signaling in health and disease. Signal Transduct. Target Ther. 5, 181 (2020).
Jin, K. et al. Cerebral neurogenesis is induced by intranasal administration of growth factors. Ann. Neurol. 53, 405–409 (2003).
Russ, A. et al. Blocking ‘don’t eat me’ signal of CD47-SIRPα in hematological malignancies, an in-depth review. Blood Rev. 32, 480–489 (2018).
Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
Chen, W.-T. et al. Spatial transcriptomics and in situ sequencing to study Alzheimer’s Disease. Cell 182, 976–991 (2020).
Wong, K., Navarro, J. F., Bergenstråhle, L., Ståhl, P. L. & Lundeberg, J. ST Spot Detector: a web-based application for automatic spot and tissue detection for spatial transcriptomics image datasets. Bioinformatics 34, 1966–1968 (2018).
Jemt, A. et al. An automated approach to prepare tissue-derived spatially barcoded RNA-sequencing libraries. Sci. Rep. 6, 37137 (2016).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Zhang, B. & Horvath, S. A general framework for weighted gene Co-Expression network analysis. Stat. Appl Genet. Mol. 4, Article 17 (2005).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Csárdi, G. & Nepusz, T. The igraph software package for complex network research. BibSonomy https://www.bibsonomy.org/bibtex/bb49a4a77b42229a427fec316e9fe515 (2006).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics J. Integr. Biol. 16, 284–287 (2012).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize Implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Colic, M. et al. Identifying chemogenetic interactions from CRISPR screens with drugZ. Genome Med. 11, 52 (2019).
Acknowledgements
We thank M. Esiri (University of Oxford) for advice and assistance in characterization of postmortem brain tissue, and G. McVean (Big Data Institute, University of Oxford) for analytical support. M.K. is supported by an MS Clinician-Scientist Fellowship from Stifterverband für die Deutsche Wissenschaft and by a Walter-Benjamin-Fellowship from Deutsche Forschungsgemeinschaft (nos. KA5554/1-1 and KA5554/1-2). F.C. is supported by the European Union’s Horizon 2020 research and innovation program under grant agreement no. 846795 (Marie Skłodowska-Curie Fellowship). P.M.M. is supported by the Edmond J. Safra Foundation and Lily Safra, an NIHR Senior Investigator Award and the UK Dementia Research Institute, which receives its funding from UK DRI Ltd, funded by the UK Medical Research Council, Alzheimer’s Society and Alzheimer’s Research UK. M.A.F. is supported by intramural funds of the University Medical Center Hamburg-Eppendorf, Deutsche Forschungsgemeinschaft (nos. SFB1328, SPP1738, KFO296, FOR2289, FOR2879, FOR5068 and FR1720/11-2), Gemeinnützige Hertie-Stiftung, Deutsche Multiple Sklerose Gesellschaft (no. V6.2) and Bundesministerium für Bildung und Forschung (Target validation, no. 16GW0308K). M.M. is supported by the Max Planck Society for the Advancement of Science, the Novo Nordisk Foundation (grant agreement nos. NNF14CC0001 and NNF15CC0001) and the European Union’s Horizon 2020 research and innovation program under grant agreement no. 686547 (MSmed Project). L.F. is supported by Wellcome (no. 100308/Z/12/Z), Medical Research Council UK (no. MC_UU_12010/3), the Oak Foundation (no. OCAY-15-520) and NIHR Oxford BRC.
Author information
Authors and Affiliations
Contributions
Conceptualization was carried out by M.K., A.-L.S., K.E.A., M.A.F. and L.F. Methodology was the responsibility of M.K., A.-L.S., R.S., F.C., C.A.D., A.M., G.C.D. and J.L. Validation was undertaken by M.K., F.C. and J.F.N. Formal analysis, data curation and visualization were performed by M.K. Investigation was carried out by M.K., A.-L.S., R.S., F.C., C.A.D., A.C., G.K., H.G.E., J.F.N., J.K.S., C.M., G.C.D. and K.E.A. Resources were provided by A.M., J.L., P.M.M. and M.A.F. Writing of the original draft was done by M.K., A.-L.S., K.E.A., M.A.F. and L.F. Writing, review and editing were performed by M.K., A.-L.S., G.C.D., P.M.M., K.E.A., M.A.F. and L.F. M.A.F., M.M. and L.F. supervised the project. Funding acquisition was done by M.M. and L.F. R.S. and F.C. contributed equally.
Corresponding author
Ethics declarations
Competing interests
G.C.D. has received research funding from Merck Serono, travel expenses from Bayer Schering, Biogen Idec, Genzyme, Merck Serono, Novartis and honoraria as an invited speaker/faculty/advisor for Novartis and Roche. J.L. and A.M. are scientific consultants for 10X Genomics. P.M.M. has received consultancy fees from Roche, Adelphi Communications, Celgene, Neurodiem and Medscape, honoraria or speakers’ fees from Novartis and Biogen and has received research or educational funds from Biogen, Novartis and GlaxoSmithKline. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Lawrence Steinman and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cell type signatures.
Enrichment of intact neuronal signature and cell type marker signatures in scRNA-seq and snRNA-seq data sets14,15,16,17,18 aggregated per sample for n = 47 (intact neurons), n = 20 (B cells), n = 29 (endothelial cells), n = 40 (microglia), n = 20 (monocytes), n = 47 (oligodendrocytes), n = 41 (OPC) and n = 20 (T cells) biological replicates. Box plots centered on median, bounds defined between the 25th and 75th percentile with minimum and maximum defined as median ± 1.5 x interquartile range and whiskers extending to the lowest / highest value within this range.
Extended Data Fig. 2 Relation of spatial transcriptomics to neuropathological assessment.
a, HE-staining of n = 1 cortical brain tissue section from patient MS1 used for spatial transcriptomics. b, Structural annotations. c, Spatial transcriptomics-based mapping of intact neuronal signature enrichment values onto gray matter. d, Spatial transcriptomics-based mapping of microglial signature enrichment values onto gray matter. e, Immunohistochemistry for myelin oligodendrocyte glycoprotein (MOG) of n = 1 tissue section adjacent to a. f, Oil Red O staining of n = 1 tissue section adjacent to a.
Extended Data Fig. 3 GO term enrichment maps.
GO term enrichment maps for gene co-expression modules. Nodes represent GO terms; edges connect terms that share genes. Groups of similar GO terms are jointly labelled for clarity. Only significantly (FDR-adjusted P value ≤ 0.1, methods) enriched GO terms for each module are shown. GO term networks with less than three nodes are not depicted unless no other terms were enriched in a module. All individual GO terms ranked by adjusted P values are listed in Supplementary Table 3.
Extended Data Fig. 4 Exclusion of potential age and sex bias and proteomic validation of ligand–receptor analysis results.
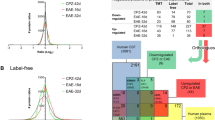
A potential bias resulting from comparing MS patients, who were on average younger and female, with older male controls (Supplementary Table 1) could impact the results of the ligand–receptor co-expression analysis (Figs. 4,5). To test for this, genome-wide RNA-expression data from post mortem cortical brain tissue of n = 336 donors with known gender and age from the GTEx consortium53 was analyzed. A linear model was fitted for each gene to assess the degree to which age and sex predicted cortical gene expression (methods). a, Genome-wide quantification of age-/sex-bias of gene expression in cortical brain tissue. The y-axis displays the variance explained by age and sex (R2), with the cut-off for a significant influence of age and/or sex on cortical expression denoted with a dotted line (FDR-adj. P value≤ 0.1, linear model, two-sided). The set of differentially co-expressed ligand/receptor genes contained less age-/sex-biased genes (13.93%) than would be expected based on the overall frequency of age-/sex-biased genes in the whole genome (19.96%). b, Mean cortical expression of exemplary genes. ZFY shows a strong sex bias, GPR26 expression is significantly influenced by age and GPR37L1 (the top candidate from our study, compare Fig. 7b) appears to be neither influenced by age nor sex. The linear model explained 63.5% of the variance (R2) for ZFY (FDR adj. P value = 8.1×10−63), 32.8% for GPR26 (FDR adj. P value = 1.09×10−21) and 0% for GPR37L1 (FDR adj. P value = 0.56). c, Further exploration of the 13.93% age-/sex-biased ligand–receptor genes to test whether they follow a pattern that could be explained by age/sex imbalance between our study cohorts. The only gene found to match an expected pattern was PDGFRA. d, Validation of ligand–receptor analysis results at the protein level. The log2 fold change between intact MS gray matter and control areas is shown for ligand–receptor genes that were detected both at the RNA and protein level. n = 18 control regions (RNA), n = 111 intact MS gray matter regions (RNA), n = 8 control regions (protein), n = 31 intact MS gray matter regions (protein). Two-sided Pearson correlation, mean centered 95% confidence band shown in gray. The correlation between ligand/receptor gene expression was positive (r = 0.41) and significant (P value ≤ 0.044).
Extended Data Fig. 5 Validation of snRNA-seq based cell type deconvolution.
Spatial transcriptomics data were integrated with single cell RNA-seq data in this study through enrichment score-based deconvolution10 implemented with DEseq265 and AUCell66. To validate the applied integration approach three additional published methods for integration between single cell RNA-seq data and spatial transcriptomics, RCTD35, spatialDWLS36 and SPOTlight37, were performed and the results compared. a, Correlation matrix for all four integration methods and all six CNS-resident cell types. All four methods are strongly correlated with each other. b, Representative projection of spot deconvolution results for all four integration methods at the same tissue location as in Fig. 5c. No major differences in cell type distribution are observed. c, Direct correlation of all alternative integration methods with enrichment score-based deconvolution for all CNS-resident cell types. n = 83,256 gray matter spot transcriptomes, r and P values derived using two-sided Pearson correlation, P values shown are FDR-adjusted. Highly significant positive correlation is observed for enrichment score-based deconvolution in comparison to each alternative method.
Extended Data Fig. 6 Correlation between ligand–receptor co-expression and module expression.
Two-sided Pearson correlations between each model and each ligand–receptor interaction are shown.
Extended Data Fig. 7 Organ immunohistochemistry.
a,b, Immunohistochemistry of indicated proteins in representative tissues. Image credit: Human Protein Atlas49. Only stainings based on antibodies whose reliability was classified as ‘supported’ or ‘enhanced’ in the Human Protein Atlas are shown. For image URLs see Supplementary Table 6. Matching proteins for transcripts found to be a, CNS-enriched or b, not enriched in the CNS at the RNA level.
Supplementary information
Supplementary Table 1
Study cohort information. Metadata associated with each donor (sheet ‘donor characteristics’) and with each sample (sheet ‘sample characteristics’) are provided.
Supplementary Table 2
Gene signatures. Cell type marker gene signatures extracted from single-cell/single-nuclei RNA-seq datasets (Methods) are provided for intact cortical neurons (sheet ‘Intact neuronal consensus sign.) and other cell types supporting cell type deconvolution analysis (sheet ‘cell type marker signatures’).
Supplementary Table 3
GO terms. Unique GO terms, significantly enriched (FDR-adjusted P ≤ 0.1, enrichGO algorithm; Methods) in gene coexpression modules.
Supplementary Table 4
Annotations of ligand–receptor interactions differentially coexpressed between intact MS gray matter and control tissue. The table provides additional information for each significant (FDR-adjusted P ≤ 0.1, unpaired two-sided t-tests) interaction, including a reference for experimental validation of the ligand–receptor relationship (ref_interaction_pubmed_id) and supporting literature for the classification as trophic factor, anti-inflammatory factor or proinflammatory factor (annotation_ref_pubmed_id). In addition, numeric values for mean absolute module membership (mean_abs_module_membership) and CNS specificity score (mean_ratio_CNS_nonCNS) are listed.
Supplementary Table 5
Published in vivo perturbations. The table provides additional details for the analysis presented in Fig. 6. The sheets ‘Candidate genes queried’ and ‘Disease models queried’ summarize search terms that were combined as queries (sheet ‘Queries’) for Europe PMC searches. Raw search results are listed in the sheet ‘Search results’. The extracted perturbations of candidate ligand–receptor interactions (Methods) are listed in the sheet ‘Perturbations_identified’.
Supplementary Table 6
Human Protein Atlas URLs. Web addresses for each image used from the Human Protein Atlas are provided.
Supplementary Table 7
Oligonucleotide sequences. Sequences are provided for oligonucleotides referred to in Methods describing spatial transcriptomics.
Supplementary Table 8
Software versions. The table lists all software used for data processing, analysis and data visualization, including version, reference and web addresses.
Rights and permissions
About this article
Cite this article
Kaufmann, M., Schaupp, AL., Sun, R. et al. Identification of early neurodegenerative pathways in progressive multiple sclerosis. Nat Neurosci 25, 944–955 (2022). https://doi.org/10.1038/s41593-022-01097-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-022-01097-3
This article is cited by
-
Arachidonic acid-derived lipid mediators in multiple sclerosis pathogenesis: fueling or dampening disease progression?
Journal of Neuroinflammation (2024)
-
A repair pathway lost in multiple sclerosis provides a new drug opportunity
Nature Immunology (2024)
-
SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
Nature Communications (2024)
-
The astrocyte-produced growth factor HB-EGF limits autoimmune CNS pathology
Nature Immunology (2024)
-
Spatial transcriptomics in development and disease
Molecular Biomedicine (2023)