Abstract
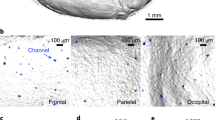
It remains unclear how immune cells from skull bone marrow niches are recruited to the meninges. Here we report that cerebrospinal fluid (CSF) accesses skull bone marrow via dura–skull channels, and CSF proteins signal onto diverse cell types within the niches. After spinal cord injury, CSF-borne cues promote myelopoiesis and egress of myeloid cells into meninges. This reveals a mechanism of CNS-to-bone-marrow communication via CSF that regulates CNS immune responses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
scRNA-seq data are available from the Gene Expression Omnibus under accession number GSE184766. All data are available in the main text or the Supplementary Information files.
Code availability
Custom code used to analyze the RNA sequencing data is freely available at https://doi.org/10.5281/zenodo.5883972.
References
Mrdjen, D. et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity 48, 380–395 (2018).
Hove, H. V. et al. A single-cell atlas of mouse brain macrophages reveals unique transcriptional identities shaped by ontogeny and tissue environment. Nat. Neurosci. 22, 1021–1035 (2019).
Dani, N. et al. A cellular and spatial map of the choroid plexus across brain ventricles and ages. Cell 184, 3056–3074 (2021).
Rustenhoven, J. et al. Functional characterization of the dural sinuses as a neuroimmune interface. Cell 184, 1000–1016 (2021).
Fitzpatrick, Z. et al. Gut-educated IgA plasma cells defend the meningeal venous sinuses. Nature 587, 472–476 (2020).
Croese, T., Castellani, G. & Schwartz, M. Immune cell compartmentalization for brain surveillance and protection. Nat. Immunol. 22, 1083–1092 (2021).
Lima, K. A., de, Rustenhoven, J. & Kipnis, J. Meningeal immunity and its function in maintenance of the central nervous system in health and disease. Annu. Rev. Immunol. 38, 597–620 (2020).
Cugurra, A. et al. Skull and vertebral bone marrow are myeloid cell reservoirs for the meninges and CNS parenchyma. Science 373, eabf7844 (2021).
Sweeney, M. D., Sagare, A. P. & Zlokovic, B. V. Blood–brain barrier breakdown in Alzheimer disease and other neurodegenerative disorders. Nat. Rev. Neurol. 14, 133–150 (2018).
Shibata, M. et al. Clearance of Alzheimer’s amyloid-β1-40 peptide from brain by LDL receptor-related protein-1 at the blood–brain barrier. J. Clin. Invest. 106, 1489–1499 (2000).
Iliff, J. J. et al. A paravascular pathway facilitates CSF flow through the brain parenchyma and the clearance of interstitial solutes, including amyloid β. Sci. Transl. Med. 4, 147ra111 (2012).
Ringstad, G. & Eide, P. K. Cerebrospinal fluid tracer efflux to parasagittal dura in humans. Nat. Commun. 11, 354 (2020).
Herisson, F. et al. Direct vascular channels connect skull bone marrow and the brain surface enabling myeloid cell migration. Nat. Neurosci. 21, 1209–1217 (2018).
Cai, R. et al. Panoptic imaging of transparent mice reveals whole-body neuronal projections and skull–meninges connections. Nat. Neurosci. 22, 317–327 (2018).
Yao, H. et al. Leukaemia hijacks a neural mechanism to invade the central nervous system. Nature 560, 55–60 (2018).
Brioschi, S. et al. Heterogeneity of meningeal B cells reveals a lymphopoietic niche at the CNS borders. Science 373, eabf9277 (2021).
Mesquita, S. D. et al. Functional aspects of meningeal lymphatics in ageing and Alzheimer’s disease. Nature 560, 185–191 (2018).
Munk, A. S. et al. PDGF-B is required for development of the glymphatic system. Cell Rep. 26, 2955–2969 (2019).
Kress, B. T. et al. Impairment of paravascular clearance pathways in the aging brain. Ann. Neurol. 76, 845–861 (2014).
Lehtinen, M. K. et al. The cerebrospinal fluid provides a proliferative niche for neural progenitor cells. Neuron 69, 893–905 (2011).
Baccin, C. et al. Combined single-cell and spatial transcriptomics reveal the molecular, cellular and spatial bone marrow niche organization. Nat. Cell Biol. 22, 38–48 (2020).
Steinman, L. Blocking adhesion molecules as therapy for multiple sclerosis: natalizumab. Nat. Rev. Drug Discov. 4, 510–518 (2005).
Vajkoczy, P., Laschinger, M. & Engelhardt, B. α4-integrin-VCAM-1 binding mediates G protein-independent capture of encephalitogenic T cell blasts to CNS white matter microvessels. J. Clin. Invest. 108, 557–565 (2001).
Ma, Q., Ineichen, B. V., Detmar, M. & Proulx, S. T. Outflow of cerebrospinal fluid is predominantly through lymphatic vessels and is reduced in aged mice. Nat. Commun. 8, 1434 (2017).
Chen, Z. W. et al. Deep amino acid sequencing of native brain GABAA receptors using high-resolution mass spectrometry. Mol. Cell. Proteomics 11, M111.011445 (2012).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Acknowledgements
We thank all the members of the Kipnis Laboratory for their valuable comments during many discussions of this work. We thank P. Bayguinov for assistance with two-photon microscopy. We thank R. R. Townsend, Q. Zhang and P. Erdmann-Gilmore for assistance with LC–MS experiments and analysis. We also thank McDonnell Genome Institute for processing scRNA-seq and the Flow Cytometry Core of the Department of Pathology and Immunology, School of Medicine, Washington University in St. Louis, for assistance with cell sorting. We also acknowledge D. Bender and the Bursky Center for Human Immunology & Immunotherapy Programs for assistance with the Luminex analysis. This work was funded by National Institutes of Health grants AT010416 and NS096967, a Cure Alzheimer’s Fund (Berg Brain Entry and Exit Consortium) grant and the BJC HealthCare Investigators Program, all to J.K, as well as by National Institutes of Health grant T32NS121881, to J.A.M. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization: J.A.M., L.C.D.S., K.A.C., J.S., T.M., J.R. and J.K. Methodology: J.A.M., L.C.D.S., K.A.C., T.D., J.S., S.D., T.M., I.S., J.R. and J.K. Investigation: J.A.M., L.C.D.S., K.A.C., T.D., J.S., S.D., T.M., I.S. and J.R. Visualization: J.A.M., L.C.D.S., K.A.C., T.D., J.R. and J.K. Funding acquisition: J.K. Project administration: J.K. Supervision: J.R. and J.K. Writing—original draft and revision: J.A.M., L.C.D.S., K.A.C., J.R. and J.K. These authors contributed equally and are listed alphabetically: K.A.C., T.D. and J.S.
Corresponding authors
Ethics declarations
Competing interests
J.K. is a scientific advisor for, holds shares of and has a licensing agreement with PureTech. The other authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of stem and immune cell populations in the basal skull marrow.
a, Flow cytometry gating strategy for major immune populations in the skull bone marrow. b, Absolute numbers of CD45+ cells in the dorsal and basal skull marrow. n = 3 mice. Mean ± SEM. c, Relative frequencies of immune populations in the dorsal skull, basal skull, and tibial bone marrow. n = 3 mice. Mean ± SEM.
Extended Data Fig. 2 Characterization of differences between the skull and tibial marrow populations.
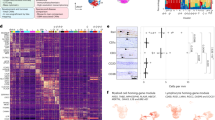
a, Dot plot demonstrating scaled gene expression and percentage of cells expressing genes for cluster phenotyping markers for bone marrow cell types from scRNA-seq analysis. b, Analysis of cluster proportions in skull and tibial bone marrow. c-f, Volcano plots of differentially expressed genes in neutrophils, monocytes, macrophages, and HSCs. Magenta dots represent upregulated transcripts, while cyan dots represent downregulated transcripts in skull populations compared to the tibia. y-axes represent adjusted log2 p value for cluster changes between skull and tibia. Dotted line represents an adjusted p value of 0.05 (general linear mixed model with Benjamini-Hochberg correction). g-j, Top 10 downregulated gene ontology pathways in skull vs. tibia for differentially expressed genes in neutrophils, monocytes, macrophages, and HSCs. k, Dot plot of receptor expression in skull bone marrow cells, scaled by gene expression and percentage of cells expressing the gene, showing expression of receptors for which there is a cognate CSF ligand.
Extended Data Fig. 3 Effects of AMD3100 on immune cell composition of the dura and bone marrow.
a, Experimental design for injections for skull bone marrow egress experiments. AMD3100 (10 μg) or artificial cerebrospinal fluid (aCSF) was injected intra-cisterna magna (i.c.m.), and mice were left for 24 hours. The following day, tissues were processed for immunolabeling or flow cytometry. b, Representative images of Ly6b+ cells and CD3+ cells in non-sinus regions of the dura. Scale bar: 200 μm. c, d, Regional analysis of Ly6b+ myeloid and CD3+ cells in the dura following AMD3100 administration. n = 3 mice per group. Data are means ± SEM, p values represent two-way ANOVA with Sidak’s post hoc test. e, Flow cytometry gating strategy for neutrophils, Ly6Chi monocytes, macrophages, and T cells in the bone marrow following AMD3100 administration. f, g, Relative numbers of neutrophils, Ly6Chi monocytes, macrophages, and T cells in the skull and tibial bone marrow 24 hours following i.c.m. AMD3100 administration. n = 5 mice per group. Data are means ± SEM, p values represent a two-sided Student’s t test.
Extended Data Fig. 4 Laminectomy does not affect CSF efflux to skull bone marrow.
a, Laminectomy, or sham surgery, was performed on mice and 3 hours later OVA-488 was injected into the cisterna magna. Tissues were collected 1 hour later for flow cytometry. Representative flow plots of macrophages in skull and tibia bone marrow with either sham surgery or laminectomy. b, Quantification of i.c.m. injected OVA uptake in macrophages following sham surgery or laminectomy. n = 5 mice per group. Data are means ± SEM, p values represent a two-way ANOVA. c, Representative flow plots of i.c.m. anti-c-Kit-PE staining in LSKs in skull and tibia bone marrow with either sham surgery or laminectomy. d, Quantification of i.c.m. injected cKit-PE uptake in LSKs following sham surgery or laminectomy. n = 5 mice per group. Data are means ± SEM, p values represent a two-way ANOVA with Sidak’s post hoc test.
Extended Data Fig. 5 Effects of spinal cord injury on vertebral bone marrow.
a, Experimental paradigm for spinal cord injury experiments. Spinal cord injury (SCI) or laminectomy (sham) was performed, and at 3 hours post-injury vertebra adjacent to the site of injury were processed for flow cytometry. b-e, Relative numbers of monocyte dendritic precursors (MDPs), common monocyte progenitors (cMoPs), Ly6Chi monocytes, and actively proliferating (Ki-67+, EdU+) monocytes in vertebral bone marrow. n = 5 mice per group. p values represent a two-sided Student’s t test. f, Multiplexed measurement of cytokines and chemokines in the CSF of sham and SCI mice using Luminex. n = 5. p values represent two-sided t tests with Holm-Sidak’s multiplicity adjustment. Data are means ± SEM.
Extended Data Fig. 6 Intracisternal injection of LPS enhances hematopoiesis in skull bone marrow and triggers myeloid egress to the dura.
a, LPS (1.25 µg, 4 µL) was injected into the skull bone marrow. After 24 hours, skullcaps and dura were processed for flow cytometry. Representative flow plots of neutrophils in the dura in aCSF and LPS-treated mice. b, Representative flow plots of Ly6Chi monocytes in the dura in aCSF and LPS-treated mice. c, Representative flow plots of LSKs in the skull BM of aCSF and LPS-treated mice. d, Quantification of the proportion of CD45+ immune cells and absolute number of neutrophils and Ly6Chi monocytes in the dura of aCSF and LPS-treated mice. n = 5 mice per group. Mean ± SEM. p values represent a two-sided Student’s t test. e, Quantification of the proportion of live cells and the proportion of actively proliferating Ki67+ stem/progenitor (LSK, MDP, cMoP, GMP, GP) and myeloid (neutrophils, Ly6Chi monocytes) cells in the skull bone marrow of aCSF and LPS-treated mice. n = 5 mice per group. Mean ± SEM. p values represent a two-sided Student’s t test.
Extended Data Fig. 7 Summary schematic for proposed mechanism.
Brain interstitial fluid and cerebrospinal fluid can efflux to skull bone marrow during healthy conditions. During CNS insults—for example pathogenic infections or spinal cord injury—CSF-derived cues can promote skull bone marrow hematopoiesis and egress of myeloid cells to underlying dura. HSC; hematopoietic stem cell, CSF; cerebrospinal fluid, ISF; interstitial fluid, SAS; subarachnoid space, BM; bone marrow, CNS, central nervous system.
Supplementary information
Supplementary Tables
Supplementary Table 1. Full list of CSF–bone marrow ligand–receptor pairs used for the chord plot in Fig. 2c and the dot plot in Extended Data Fig. 2k. Supplementary Table 2. Antibodies used in flow cytometry and immunostaining experiments.
41593_2022_1029_MOESM3_ESM.mov
Supplementary Video 1. Perivascular and bone marrow localization of tracer after ICM OVA injection. Z-stack movie of a region of interest in a dura–skull whole mount stained with CD31 (green), OsteoSense (blue) and ICM OVA (magenta). Video direction is from dura to skull bone marrow. Frames shown in Fig. 1b. Scale bar, 100 µm.
Rights and permissions
About this article
Cite this article
Mazzitelli, J.A., Smyth, L.C.D., Cross, K.A. et al. Cerebrospinal fluid regulates skull bone marrow niches via direct access through dural channels. Nat Neurosci 25, 555–560 (2022). https://doi.org/10.1038/s41593-022-01029-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-022-01029-1