Abstract
Understanding the tissue-specific genetic controls of protein levels is essential to uncover mechanisms of post-transcriptional gene regulation. In this study, we generated a genomic atlas of protein levels in three tissues relevant to neurological disorders (brain, cerebrospinal fluid and plasma) by profiling thousands of proteins from participants with and without Alzheimer’s disease. We identified 274, 127 and 32 protein quantitative trait loci (pQTLs) for cerebrospinal fluid, plasma and brain, respectively. cis-pQTLs were more likely to be tissue shared, but trans-pQTLs tended to be tissue specific. Between 48.0% and 76.6% of pQTLs did not co-localize with expression, splicing, DNA methylation or histone acetylation QTLs. Using Mendelian randomization, we nominated proteins implicated in neurological diseases, including Alzheimer’s disease, Parkinson’s disease and stroke. This first multi-tissue study will be instrumental to map signals from genome-wide association studies onto functional genes, to discover pathways and to identify drug targets for neurological diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Both summary statistics and individual-level data have been uploaded to the National Institute on Aging Genetics of Alzheimer’s Disease Data Storage Site repository at https://www.niagads.org/datasets/ng00102 for the three tissues from the Knight ADRC dataset for discovery. Summary statistics (pQTL) data are freely available; as the data exceeds 500 Gb, please email niagads@pennmedicine.upenn.edu to set up an FTP transfer of the data. Summary association results can also be explored through Online Neurodegenerative Trait Integrative Multi-Omics Explorer (ONTIME) (https://ontime.wustl.edu/), a PheWeb (v1.1.14)-based browser.
CSF-Sasayama2017 dataset for replication: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE83711.
Plasma-AddNeuroMed dataset for replication: https://www.synapse.org/#!Synapse:syn4988768.
Drug targets were queried using DrugBank database collected via UniProtKB (as of 3 January 2020) at https://www.uniprot.org/database/DB-0019.
References
Altshuler, D., Daly, M. J. & Lander, E. S. Genetic mapping in human disease. Science 322, 881–888 (2008).
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414 (2019).
Claussnitzer, M. et al. A brief history of human disease genetics. Nature 577, 179–189 (2020).
Aguet, F. et al. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
van der Wijst, M. G. P. et al. The single-cell eQTLGen consortium. eLife 9, e52155 (2020).
Aguet, F. et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Gamazon, E. R., Zwinderman, A. H., Cox, N. J., Denys, D. & Derks, E. M. Multi-tissue transcriptome analyses identify genetic mechanisms underlying neuropsychiatric traits. Nat. Genet. 51, 933–940 (2019).
Võsa, U. et al. Unraveling the polygenic architecture of complex traits using blood eQTL metaanalysis. Preprint at bioRxiv https://doi.org/10.1101/447367 (2018).
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018).
Suhre, K. et al. Connecting genetic risk to disease end points through the human blood plasma proteome. Nat. Commun. 8, 14357 (2017).
Folkersen, L. et al. Mapping of 79 loci for 83 plasma protein biomarkers in cardiovascular disease. PLoS Genet. 13, e1006706 (2017).
Deming, Y. et al. Genetic studies of plasma analytes identify novel potential biomarkers for several complex traits. Sci. Rep. 6, 18092 (2016).
Sasayama, D. et al. Genome-wide quantitative trait loci mapping of the human cerebrospinal fluid proteome. Hum. Mol. Genet. 26, 44–51 (2017).
Kauwe, J. S. K. et al. Genome-wide association study of CSF levels of 59 Alzheimer’s disease candidate proteins: significant associations with proteins involved in amyloid processing and inflammation. PLoS Genet. 10, e1004758 (2014).
Robins, C. et al. Genetic control of the human brain proteome. Preprint at bioRxiv https://doi.org/10.1101/816652 (2019).
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS ONE 5, e15004 (2010).
Haddick, P. C. G. et al. A common variant of IL-6R is associated with elevated IL-6 pathway activity in Alzheimer’s disease brains. J. Alzheimers Dis. 56, 1037–1054 (2017).
Marek, K. et al. The Parkinson Progression Marker Initiative (PPMI). Prog. Neurobiol. 95, 629–635 (2011).
Lovestone, S. et al. AddNeuroMed—the European collaboration for the discovery of novel biomarkers for Alzheimer’s disease. Ann. N. Y. Acad. Sci. 1180, 36–46 (2009).
Kamat, M. A. et al. PhenoScanner V2: an expanded tool for searching human genotype–phenotype associations. Bioinformatics 35, 4851–4853 (2019).
Jayaratnam, S., Khoo, A. K. L. & Basic, D. Rapidly progressive Alzheimer’s disease and elevated 14-3-3 proteins in cerebrospinal fluid. Age Ageing 37, 467–469 (2008).
Foote, M. & Zhou, Y. 14-3-3 proteins in neurological disorders. Int. J. Biochem. Mol. Biol. 3, 152–164 (2012).
Ibanez, L. et al. Overlap in the genetic architecture of stroke risk, early neurological changes, and cardiovascular risk factors. Stroke 50, 1339–1345 (2019).
Lourdusamy, A. et al. Identification of cis-regulatory variation influencing protein abundance levels in human plasma. Hum. Mol. Genet. 21, 3719–3726 (2012).
Walker, R. L. et al. Genetic control of expression and splicing in developing human brain informs disease mechanisms. Cell 179, 750–771 (2019).
Orozco, L. D. et al. Integration of eQTL and a single-cell atlas in the human eye identifies causal genes for age-related macular degeneration. Cell Rep. 30, 1246–1259 (2020).
Urbut, S. M., Wang, G., Carbonetto, P. & Stephens, M. Flexible statistical methods for estimating and testing effects in genomic studies with multiple conditions. Nat. Genet. 51, 187–195 (2019).
Hillary, R. F. et al. Genome and epigenome wide studies of neurological protein biomarkers in the Lothian Birth Cohort 1936. Nat. Commun. 10, 3160–3160 (2019).
Suhre, K., McCarthy, M. I. & Schwenk, J. M. Genetics meets proteomics: perspectives for large population-based studies. Nat. Rev. Genet. 22, 19–37 (2020).
Yao, C. et al. Genome‐wide mapping of plasma protein QTLs identifies putatively causal genes and pathways for cardiovascular disease. Nat. Commun. 9, 3268 (2018).
Paré, G. et al. Novel association of ABO histo-blood group antigen with soluble ICAM-1: results of a genome-wide association study of 6,578 women. PLoS Genet. 4, e1000118 (2008).
Ndungu, A., Payne, A., Torres, J. M., van de Bunt, M. & McCarthy, M. I. A multi-tissue transcriptome analysis of human metabolites guides interpretability of associations based on multi-SNP models for gene expression. Am. J. Hum. Genet. 106, 188–201 (2020).
Cruchaga, C. et al. Cerebrospinal fluid APOE levels: an endophenotype for genetic studies for Alzheimer’s disease. Hum. Mol. Genet. 21, 4558–4571 (2012).
Kibinge, N. K., Relton, C. L., Gaunt, T. R. & Richardson, T. G. Characterizing the causal pathway for genetic variants associated with neurological phenotypes using human brain-derived proteome data. Am. J. Hum. Genet. 106, 885–892 (2020).
Del-Aguila, J. L. et al. A single-nuclei RNA sequencing study of Mendelian and sporadic AD in the human brain. Alzheimer’s Res. Ther. 11, 71 (2019).
Alector Inc. First in human study for safety and tolerability of AL003. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03822208 (2019).
Nalls, M. A. et al. Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol. 18, 1091–1102 (2019).
Bethea, J. W. Clinical Anesthesia, 6th Edition. Anesthesiology 112, 767–768 (2010).
Camerino, G. M. et al. Elucidating the contribution of skeletal muscle ion channels to amyotrophic lateral sclerosis in search of new therapeutic options. Sci. Rep. 9, 3185 (2019).
Savitz, S. I. et al. The novel beta-blocker, carvedilol, provides neuroprotection in transient focal stroke. J. Cereb. Blood Flow. Metab. 20, 1197–1204 (2000).
Nelson, M. R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Gagliano Taliun, S. A. et al. Exploring and visualizing large-scale genetic associations by using PheWeb. Nat. Genet. 52, 550–552 (2020).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. eLife 7, e34408 (2018).
Del-Aguila, J. L. et al. Assessment of the genetic architecture of Alzheimer’s disease risk in rate of memory decline. J. Alzheimers Dis. 62, 745–756 (2018).
Huang, K. et al. A common haplotype lowers PU.1 expression in myeloid cells and delays onset of Alzheimer’s disease. Nat. Neurosci. 20, 1052–1061 (2017).
van Rheenen, W. et al. Genome-wide association analyses identify new risk variants and the genetic architecture of amyotrophic lateral sclerosis. Nat. Genet. 48, 1043–1048 (2016).
Ferrari, R. et al. Frontotemporal dementia and its subtypes: a genome-wide association study. Lancet Neurol. 13, 686–699 (2014).
Malik, R. et al. Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat. Genet. 50, 524 (2018).
Demenais, F. et al. Multiancestry association study identifies new asthma risk loci that colocalize with immune-cell enhancer marks. Nat. Genet. 50, 42–53 (2018).
Chen, H. VennDiagram: generate high-resolution Venn and Euler plots. https://rdrr.io/cran/VennDiagram/ (2018).
Morris, J. C. The Clinical Dementia Rating (CDR): current version and scoring rules. Neurology 43, 2412–2414 (1993).
Mirra, S. S. et al. The Consortium to Establish a Registry for Alzheimer’s Disease (CERAD). Part II. Standardization of the neuropathologic assessment of Alzheimer’s disease. Neurology 41, 479–486 (1991).
Khachaturian, Z. S. Diagnosis of Alzheimer’s disease. Arch. Neurol. 42, 1097–1105 (1985).
Sattlecker, M. et al. Alzheimer’s disease biomarker discovery using SOMAscan multiplexed protein technology. Alzheimers Dement. 10, 724–734 (2014).
Williams, S. A. et al. Plasma protein patterns as comprehensive indicators of health. Nat. Med. 25, 1851–1857 (2019).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
UniProt Consortium. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 47, D506–D515 (2019).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Pruim, R. J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, 2009).
Iotchkova, V. et al. GARFIELD classifies disease-relevant genomic features through integration of functional annotations with association signals. Nat. Genet. 51, 343 (2019).
Mancuso, N. et al. Integrating gene expression with summary association statistics to identify genes associated with 30 complex traits. Am. J. Hum. Genet. 100, 473–487 (2017).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. circlize implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Wallace, C. Statistical testing of shared genetic control for potentially related traits. Genet. Epidemiol. 37, 802–813 (2013).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Ng, B. et al. An xQTL map integrates the genetic architecture of the human brain’s transcriptome and epigenome. Nat. Neurosci. 20, 1418–1426 (2017).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Ongen, H., Buil, A., Brown, A. A., Dermitzakis, E. T. & Delaneau, O. Fast and efficient QTL mapper for thousands of molecular phenotypes. Bioinformatics 32, 1479–1485 (2016).
Wishart, D. S. et al. DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res. 34, D668–D672 (2006).
Acknowledgements
We thank all the participants and their families as well as the many involved institutions and their staff. Funding: This work was supported by grants from the National Institutes of Health (NIH) (R01AG044546 (C.C.), P01AG003991 (C.C. and J.C.M.), RF1AG053303 (C.C.), RF1AG058501 (C.C.), U01AG058922 (C.C.), R01NS118146 (B.A.B.) and R01AG057777 (O.H.)) and the Alzheimer Association (NIRG-11-200110 (C.C.), BAND-14-338165 (C.C.), AARG-16-441560 (C.C.) and BFG-15-362540 (C.C.)). This work was supported by access to equipment made possible by the Hope Center for Neurological Disorders and the Departments of Neurology and Psychiatry at Washington University School of Medicine. The recruitment and clinical characterization of research participants at Washington University were supported by NIH P50AG05681 (J.C.M.), P01AG03991 (J.C.M.) and P01AG026276 (J.C.M.).
Author information
Authors and Affiliations
Contributions
C.Y. performed the analyses, interpreted the results and wrote the manuscript. F.H.G.F., L.I., M.V.F., F.W., J.L.B., Z.L., U.D., Y.S., K.M. and J.P.B. contributed to data collection, data processing, quality control and cleaning. J.C.M., A.M.F. and R.J.P. contributed samples and/or data. B.S. wrote the manuscript. J.A.B., B.E. and O.H. developed the PheWeb browser. B.A.B. interpreted the results. H.R., O.H. and C.C. designed the study, collected the data, supervised the analyses, interpreted the results and wrote the manuscript. C.Y., A.S. and C.C. addressed the comments from peer review and updated the manuscript. All authors read and contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
C.C. receives research support from Biogen, EISAI, Alector and Parabon. C.C. is a member of the advisory board of Vivid Genomics, Halia Therapeutics and ADx Healthcare. The remaining authors declare no competing financial interests.
Additional information
Peer review information Nature Neuroscience thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 QC pipeline.
QC on both proteins (a to c) and samples (d) were described as follows: a, Flowchart of CSF protein level QC, starting from 1305; after step-1, Limit Of Detection VS 2-StDeviation, 807 proteins were kept with a pass-rate > = 85%; after step-2, given Max Difference of Scale Factor < 0.5, 749 proteins were kept; after step-3, given Coefficient of Variation (of calibrator) < 0.15 & step-4, given IQR, sum(outliers) < 15%, 746 proteins were kept. After step-5, 713 proteins that shared by < 30 samples (shared by ~80% of the subject outliers) were kept. b, Flowchart of plasma protein level QC, starting from 1305; after step-1, 1301 proteins were kept with a pass-rate > = 85%; after step-2, 956 proteins were kept; after step-3 & step-4, 955 proteins were kept. After step-5, 931 proteins that shared by < 10 samples were kept. c, Flowchart of brain protein level QC, starting from 1305; after step-1, 1109 proteins were kept with a pass-rate > = 85%; after step-2, 1107 proteins were kept; after step-3 & step-4, given IQR, sum(outliers) < 15%, 1106 proteins were kept. After step-5, 1079 proteins that shared by < 21 samples were kept. d, Table of sample size after each step of QC in genotype and proteomics. Within each tissue (1st column), we profiled proteomics from 1300 CSF, 648 plasma and 459 samples (2nd column). From unique donors in proteomics data (3rd column), we first kept donors with genotyping array data (4th column). We next kept only the donors with a European ancestry after checking principal components (5th column). Moreover, we kept donors that were not close with each other (PI_HAT < 0.05) after checking identity by descent (6th column). Finally, the samples remained only passing both the genotype and protein data QC (7th column).
Extended Data Fig. 2 Reproducibility of proteomic data.
a, Table of total sample size for each tissue before and after QC, including the biological and technical replicates. b, Venn diagram on the designed donor overlap across tissues. c, Scatterplot of 321 subjects with both longitudinal and baseline samples from CSF indicates a Pearson correlation coefficient of 0.995 (95% confidence interval from 0.995 to 0.995). d, Scatterplot of 11 subjects with both fasted and nonfasted samples from plasma indicates a Pearson correlation coefficient of 0.907 (95% confidence interval from 0.904 to 0.911). e, Scatterplot of one subject with both longitudinal and baseline samples from plasma indicates a Pearson correlation coefficient of 0.938 (95% confidence interval from 0.930 to 0.945). f, Scatterplot of one subject with two technical replicates from brain indicates a Pearson correlation coefficient of 0.976 (95% confidence interval from 0.976 to 0.981). All statistical tests used were two-sided from (c) to (f).
Extended Data Fig. 3 Overview of the sample size and number of pQTLs from pQTL studies mentioned in this paper and the summary statistics from the meta-analyses.
a, Scatter plot of sample size (log10-scaled) and number of total pQTLs after clumping or unique proteins when no clumping was performed (log10-scaled). Dot color represents the tissue type; dot size represents total number of proteins profiled. b, Table of these nine datasets listed the exact numbers for drawing the scatter plot. c, Table of three different combinations of meta-analyses: 2) meta2_WUcsf_PPMI19_JP17: meta-analysis on all three CSF studies by Sasayama and colleagues published in 2017, by PPMI released in 2019, and by Washington University cohort (this study); 3) meta3_WUcsf_WUplasma_WUbrain: meta-analysis on all three-tissue findings from CSF, plasma and brain respectively by Washington University cohort (this study); 4) meta4_ WUcsf_WUplasma_WUbrain_ PPMI19_JP17: meta-analysis on both the CSF studies by Sasayama and colleagues published in 2017 and by PPMI released in 2019 plus all three-tissue findings from CSF, plasma and brain respectively by Washington University cohort (this study). The columns include number of proteins in common, number of protein-level GWAS hits after meta-analysis, number of protein-level GWAS hits before meta-analysis using only the common proteins within each tissue for each combination. d, Stacked Manhattan plots for all three different combinations of meta-analyses. The darkred line represents P = 5 × 10-8.
Extended Data Fig. 4 Disease stratified analysis on comparing pQTLs effect size.
To investigate of disease status effect on pQTLs, we performed linear regression on the same protein-loci pairs (before conditioning on top variants) identified from above default model using three additional models: a, joint analysis but with disease status as another covariate (CO vs non-CO). Pearson correlation coefficient was 0.999 (p-value < 2.2 × 10-16, 95%CI = 0.999 to 0.999), 0.999 (p-value = 4.3 × 10-202, 95%CI = 0.999 to 0.999), 0.999 (p-value = 9.5 × 10-52, 95%CI = 0.999 to 0.999) for CSF, plasma, and brain respectively. Sample size for this joint analysis was 835, 529, and 380 for CSF, plasma, and brain respectively. b, AD case (CA) only using the same covariates as default model. Pearson correlation coefficient of 0.991 (p-value = 3.9 × 10-160, 95%CI = 0.988 to 0.993), 0.989 (p-value = 1.8 × 10-83, 95%CI = 0.983 to 0.992), 0.998 (p-value = 2.4 × 10-29, 95%CI = 0.995 to 0.999) for CSF, plasma, and brain respectively. Sample size for this AD case (CA) only analysis was 217, 168, and 248 for CSF, plasma, and brain respectively. c, Cognitive unimpaired (CO) only using the same covariates as default model. Pearson correlation coefficient of 0.999 (p-value = 5.2 × 10-234, 95%CI = 0.998 to 0.999), 0.998 (p-value = 1.17 × 10-122, 95%CI = 0.997 to 0.999), 0.602 (p-value = 0.002, 95%CI = 0.262 to 0.809) for CSF, plasma, and brain respectively. Sample size for this cognitive unimpaired (CO) only analysis was 614, 357, and 24 for CSF, plasma, and brain respectively. The relatively low correlation in default model comparison with control only in brain samples was due to much smaller sample size as a control for brain samples. All statistical tests used were two-sided from (a) to (c).
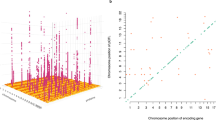
Extended Data Fig. 5 Global view of pleiotropic regions in CSF.
In total, 59 Pleiotropic regions passing genome-wide significance threshold (5 × 10-8) in CSF (sample size = 835). Unique non-overlapping regions associated with a given SOMAmer were first defined as 1-Mb region upstream and downstream of each significant variant for that SOMAmer. Within the region (2 Mb) containing the variant with the smallest P value, any overlapping regions were then merged into the same locus. Next, an LD-based clumping approach was adapted to identify whether a region was associated with multiple SOMAmers. Variants were combined into a single region per LD (EUR) defined loci. Any loci associated with more than one protein were identified as pleiotropic regions. Genomic locations of pQTLs were visualized by a squared-Manhattan plot. Dark-green represents cis-pQTLs; gold represents trans-pQTLs. X-axis indicates the positions of the top variant; and Y-axes indicates the gene encoding the protein. All pleiotropic genomic regions are annotated at the top of each plot along the X-axis.
Extended Data Fig. 6 Global view of pleiotropic regions in plasma.
In total, 34 pleiotropic regions passing genome-wide significance threshold (5 × 10-8) in plasma (sample size = 529). Genomic locations of pQTLs were visualized by a squared-Manhattan plot, same as Extended Data Fig. 5.
Extended Data Fig. 7 Global view of pleiotropic regions in brain.
In total, 10 pleiotropic regions passing genome-wide significance threshold (5 × 10-8) in brain (sample size = 380). Genomic locations of pQTLs were visualized by a squared-Manhattan plot, same as Extended Data Fig. 5.
Extended Data Fig. 8 Tissue specificity exploration with permissive thresholds.
To determine whether our tissue-specificity results were biased by statistical power, we performed similar analyses with two more permissive p-values on the 411 proteins. a, Venn diagrams of all pQTLs across all three tissues by fixing genome-wide significance threshold (5 × 10-8) for all three tissues. b, Venn diagrams of all pQTLs across all three tissues by fixing genome-wide significance threshold for one tissue and 0.001 for the other two tissues. For example, when checking CSF pQTLs shared in plasma or brain, we chose 5 × 10-8 as threshold for CSF and 0.001 for plasma or brain. c, Venn diagrams of all pQTLs across all three tissues by fixing genome-wide significance threshold for one tissue and 0.05 for the other two tissues. For example, when checking CSF pQTLs shared in plasma or brain, we chose 5 × 10-8 as threshold for CSF and 0.05 for plasma or brain.
Extended Data Fig. 9 Tissue specificity exploration with plasma result from INTERVAL study.
To further demonstrate that tissue-specificity findings are not a product of different sample size, we performed similar comparisons by analyzing the plasma pQTLs from the INTERVAL study on 616 proteins that passed QC in our CSF, brain and plasma INTERVAL. a, Venn diagrams of proteins passing QC across all three tissues: CSF and brain results are from WashU cohort, plasma result is from INTERVAL study. b, Venn diagrams of all pQTLs across all three tissues by fixing genome-wide significance threshold (5 × 10-8) for all three tissues. c, Venn diagrams of all pQTLs across all three tissues by fixing genome-wide significance threshold for one tissue and 0.001 for the other two tissues. For example, when checking CSF pQTLs shared in plasma or brain, we chose 5 × 10-8 as threshold for CSF and 0.001 for plasma or brain. d, Venn diagrams of all pQTLs across all three tissues by fixing genome-wide significance threshold for one tissue and 0.05 for the other two tissues. For example, when checking CSF pQTLs shared in plasma or brain, we chose 5 × 10-8 as threshold for CSF and 0.05 for plasma or brain.
Extended Data Fig. 10 Properties of pQTLs.
a, Dot plots of -log10(P) from all significant associations (via linear regression) against the distance of sentinel SNPs from TSS within each tissue. b, Dot plots of absolute effect size associated with MAF within each tissue. c, Forest plot of enrichment on the predicted functional annotation classes of pQTLs versus null sets of variants from permutation within each tissue (Data are presented as mean values of Odds Ratio + /- 95% confidence interval from Fisher’s Exact Test) and Bar plots of the proportion of variants annotate in each class. (Note: Features on exonic_splicing/ncRNA_splicing/splicing/UTR5_UTR3 are not shown due to not all tissues have these features). d, Histograms of variance explained by conditionally independent variants within each tissue. For CSF, the mean = 0.141, standard deviation = 0.144, mode = 0.061; For plasma, the mean = 0.157, standard deviation = 0.125, mode = 0.188; For brain, the mean = 0.208, standard deviation = 0.151, mode = 0.092.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9 and Supplementary Results.
Supplementary Table 1
Supplementary Tables 1–35.
Rights and permissions
About this article
Cite this article
Yang, C., Farias, F.H.G., Ibanez, L. et al. Genomic atlas of the proteome from brain, CSF and plasma prioritizes proteins implicated in neurological disorders. Nat Neurosci 24, 1302–1312 (2021). https://doi.org/10.1038/s41593-021-00886-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-021-00886-6
This article is cited by
-
Associations between genetically predicted plasma protein levels and Alzheimer’s disease risk: a study using genetic prediction models
Alzheimer's Research & Therapy (2024)
-
Investigation of the acute pathogenesis of spondyloarthritis/HLA-B27-associated anterior uveitis based on genome-wide association analysis and single-cell transcriptomics
Journal of Translational Medicine (2024)
-
A multi-ancestry genetic study of pain intensity in 598,339 veterans
Nature Medicine (2024)
-
Identifying novel proteins for suicide attempt by integrating proteomes from brain and blood with genome-wide association data
Neuropsychopharmacology (2024)
-
TOPMed imputed genomics enhances genomic atlas of the human proteome in brain, cerebrospinal fluid, and plasma
Scientific Data (2024)