Abstract
Nuclear compartments are thought to play a role in three-dimensional genome organization and gene expression. In mammalian brain, the architecture and dynamics of nuclear compartment-associated genome organization is not known. In this study, we developed Genome Organization using CUT and RUN Technology (GO-CaRT) to map genomic interactions with two nuclear compartments—the nuclear lamina and nuclear speckles—from different regions of the developing mouse, macaque and human brain. Lamina-associated domain (LAD) architecture in cells in vivo is distinct from that of cultured cells, including major differences in LADs previously considered to be cell type invariant. In the mouse and human forebrain, dorsal and ventral neural precursor cells have differences in LAD architecture that correspond to their regional identity. LADs in the human and mouse cortex contain transcriptionally highly active sub-domains characterized by broad depletion of histone-3-lysine-9 dimethylation. Evolutionarily conserved LADs in human, macaque and mouse brain are enriched for transcriptionally active neural genes associated with synapse function. By integrating GO-CaRT maps with genome-wide association study data, we found speckle-associated domains to be enriched for schizophrenia risk loci, indicating a physical relationship between these disease-associated genetic variants and a specific nuclear structure. Our work provides a framework for understanding the relationship between distinct nuclear compartments and genome function in brain development and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The mouse and macaque datasets generated in this study are publicly available in the GEO repository under the acession number GSE175679. The human source data described in this paper are available via the PsychENCODE Knowledge Portal (https://psychencode.synapse.org/). The PsychENCODE Knowledge Portal is a platform for accessing data, analyses and tools generated through grants funded by the National Institute of Mental Health (NIMH) PsychENCODE program. Data are available for general research use according to the requirements for data access and data attribution listed at https://psychencode.synapse.org/DataAccess. For access to content described in this paper see https://doi.org/10.7303/syn25931622.
Code availability
All codes for processing and analyzing the data presented in this work are available upon reasonable request.
References
Fraser, P. & Bickmore, W. Nuclear organization of the genome and the potential for gene regulation. Nature 447, 413–417 (2007).
Zhao, R., Bodnar, M. S. & Spector, D. L. Nuclear neighborhoods and gene expression. Curr. Opin. Genet. Dev. 19, 172–179 (2009).
van Steensel, B. & Belmont, A. S. Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 169, 780–791 (2017).
Chen, Y. et al. Mapping 3D genome organization relative to nuclear compartments using TSA-seq as a cytological ruler. J. Cell Biol. 217, 4025–4048 (2018).
Lamond, A. I. & Spector, D. L. Nuclear speckles: a model for nuclear organelles. Nat. Rev. Mol. Cell Biol. 4, 605–612 (2003).
Peric-Hupkes, D. et al. Molecular maps of the reorganization of genome–nuclear lamina interactions during differentiation. Mol. Cell 38, 603–613 (2010).
Poleshko, A. et al. Genome–nuclear lamina interactions regulate cardiac stem cell lineage restriction. Cell 171, 573–587 (2017).
Robson, M. I. et al. Tissue-specific gene repositioning by muscle nuclear membrane proteins enhances repression of critical developmental genes during myogenesis. Mol. Cell 62, 834–847 (2016).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
Vogel, M. J., Peric-Hupkes, D. & van Steensel, B. Detection of in vivo protein–DNA interactions using DamID in mammalian cells. Nat. Protoc. 2, 1467–1478 (2007).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 6, e21856 (2017).
Harr, J. C. et al. Directed targeting of chromatin to the nuclear lamina is mediated by chromatin state and A-type lamins. J. Cell Biol. 208, 33–52 (2015).
Wu, F. & Yao, J. Identifying novel transcriptional and epigenetic features of nuclear lamina-associated genes. Sci. Rep. 7, 100 (2017).
Yu, T. S., Zhang, G., Liebl, D. J. & Kernie, S. G. Traumatic brain injury-induced hippocampal neurogenesis requires activation of early nestin-expressing progenitors. J. Neurosci. 28, 12901–12912 (2008).
Borsos, M. et al. Genome–lamina interactions are established de novo in the early mouse embryo. Nature 569, 729–733 (2019).
Hebert, J. M. & Fishell, G. The genetics of early telencephalon patterning: some assembly required. Nat. Rev. Neurosci. 9, 678–685 (2008).
Puelles, L. & Rubenstein, J. L. Forebrain gene expression domains and the evolving prosomeric model. Trends Neurosci. 26, 469–476 (2003).
Delgado, R. N. & Lim, D. A. Maintenance of positional identity of neural progenitors in the embryonic and postnatal telencephalon. Front. Mol. Neurosci. 10, 373 (2017).
van Schaik, T., Vos, M., Peric-Hupkes, D., Hn Celie, P. & van Steensel, B. Cell cycle dynamics of lamina-associated DNA. EMBO Rep. 21, e50636 (2020).
Leemans, C. et al. Promoter-intrinsic and local chromatin features determine gene repression in LADs. Cell 177, 852–864 (2019).
Workman, A. D., Charvet, C. J., Clancy, B., Darlington, R. B. & Finlay, B. L. Modeling transformations of neurodevelopmental sequences across mammalian species. J. Neurosci. 33, 7368–7383 (2013).
Meuleman, W. et al. Constitutive nuclear lamina–genome interactions are highly conserved and associated with A/T-rich sequence. Genome Res. 23, 270–280 (2013).
Sun, Z. et al. EGR1 recruits TET1 to shape the brain methylome during development and upon neuronal activity. Nat. Commun. 10, 3892 (2019).
Preciados, M., Yoo, C. & Roy, D. Estrogenic endocrine disrupting chemicals influencing NRF1 regulated gene networks in the development of complex human brain diseases. Int. J. Mol. Sci. 17, 2086 (2016).
Han, H. et al. Multilayered control of alternative splicing regulatory networks by transcription factors. Mol. Cell 65, 539–553 (2017).
Iotchkova, V. et al. GARFIELD classifies disease-relevant genomic features through integration of functional annotations with association signals. Nat. Genet. 51, 343–353 (2019).
Pardinas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
Grove, J. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 51, 431–444 (2019).
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Gandal, M. J. et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science 362, eaat8127 (2018).
Markenscoff-Papadimitriou, E. et al. A chromatin accessibility atlas of the developing human telencephalon. Cell 182, 754–769 (2020).
Won, H. et al. Chromosome conformation elucidates regulatory relationships in developing human brain. Nature 538, 523–527 (2016).
Yokochi, T. et al. G9a selectively represses a class of late-replicating genes at the nuclear periphery. Proc. Natl Acad. Sci. USA 106, 19363–19368 (2009).
Kind, J. et al. Single-cell dynamics of genome–nuclear lamina interactions. Cell 153, 178–192 (2013).
Jaffe, A. E. et al. Developmental and genetic regulation of the human cortex transcriptome illuminate schizophrenia pathogenesis. Nat. Neurosci. 21, 1117–1125 (2018).
Bannister, A. J. & Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 21, 381–395 (2011).
Jiang, C. & Pugh, B. F. Nucleosome positioning and gene regulation: advances through genomics. Nat. Rev. Genet. 10, 161–172 (2009).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380(2012).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Lim, D. A. et al. Chromatin remodelling factor Mll1 is essential for neurogenesis from postnatal neural stem cells. Nature 458, 529–533 (2009).
Skene, P. J., Henikoff, J. G. & Henikoff, S. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat. Protoc. 13, 1006–1019 (2018).
Lund, E., Oldenburg, A. R. & Collas, P. Enriched domain detector: a program for detection of wide genomic enrichment domains robust against local variations. Nucleic Acids Res. 42, e92 (2014).
Stovner, E. B. & Saetrom, P. epic2 efficiently finds diffuse domains in ChIP-seq data. Bioinformatics 35, 4392–4393 (2019).
Brahma, S. & Henikoff, S. RSC-associated subnucleosomes define MNase-sensitive promoters in yeast. Mol. Cell 73, 238–249 (2019).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Park, D. H. et al. Activation of neuronal gene expression by the JMJD3 demethylase is required for postnatal and adult brain neurogenesis. Cell Rep. 8, 1290–1299 (2014).
Mirzadeh, Z., Merkle, F. T., Soriano-Navarro, M., Garcia-Verdugo, J. M. & Alvarez-Buylla, A. Neural stem cells confer unique pinwheel architecture to the ventricular surface in neurogenic regions of the adult brain. Cell Stem Cell 3, 265–278 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
McLeay, R. C. & Bailey, T. L. Motif Enrichment Analysis: a unified framework and an evaluation on ChIP data. BMC Bioinformatics 11, 165 (2010).
Acknowledgements
The authors thank members of the S. Henikoff laboratory (Fred Hutchinson Cancer Research Center) for providing aliquots of pA-MNase; A. Bhaduri and T. Mukhtar (UCSF) for help with primary human tissue samples; N. K. Matharu (UCSF) for suggesting the name GO-CaRT and helpful discussions throughout the writing of this manuscript and UCSF Parnassus Flow Core, RRID:SCR_018206. Funding: This study was supported by National Institutes of Health grants 1R01-NS112357, 1R01NS091544 and Veterans Affairs grant 5I01 BX000252; the Chad Tough Foundation; the Childhood Brain Tumor Foundation and Sandler Program for Breakthrough Biomedical Research (to D.A.L.); National Institute of Mental Health (NIMH) grant 1U01MH116438 (to T.J.N. and A.A.P.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
S.H.A. and D.A.L. conceived the study and designed the experiments. S.H.A. performed the experiments, interpreted data and wrote the manuscript. R.N.D. performed tissue dissections and data analyses. E.G. assisted in computational analyses and quantification of DNA-FISH. M.A.C. performed the GWAS variant analysis in nuclear compartments. J.Z. performed conservation analyses of LADs. S.J.H. performed immunohistochemistry on mouse brain sections. A.R.K., T.J.N. and A.A.P. supervised human and macaque experiments. D.A.L. supervised research and helped write the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Peer review information Nature Neuroscience thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Gene expression and chromatin features of LADs identified by GO-CaRT in 3T3 MEFs.
a, Venn diagram showing percent overlap of LAD coverage between GO-CaRT and DamID in MEFs. b, Box and whiskers plot depicting average gene expression (RNA-seq- SRX803943, SRX803944) in LADs and inter-LADs defined by GO-CaRT and DamID in MEFs. P = Two-tailed student’s t-test. GO-CaRT LADs, n = 4390, inter-LADs, n = 30499; DamID LADs, n = 5313, inter-LADs, n = 29576, n = number of genes. Boxes show the range from lower (25th percentile) to upper quartiles (75th percentile), with the median line (50th percentile); whiskers extend 1.5 times the inter-quartile range from bounds of box. c-e, Average H3K9me2 (c), H3K9me3 (d) and H3K27me3 (e) signal over LADs and flanking regions (400 Kb). Average LaminB signal (red) is also shown in each plot.
Extended Data Fig. 2 Gene density and expression in LADs identified in mouse brain NPCs in vivo.
a, Circos plot showing LaminB enrichment and LADs (horizontal bars) across the mouse chromosomes (1-19) in dorsal and ventral NPCs. b, LAD and inter-LADs gene density in brain NPCs (GO-CaRT) and indicated cultured cell lines (DamID). ASCs, astrocytes., ESCs, embryonic stem cells. c, Box plot depicting average gene expression in LADs (dorsal, n = 3683; ventral, n = 3520) and inter-LADs (dorsal, n = 18849; ventral, n = 19014) as determined by RNA-seq in dorsal and ventral NPCs, n = number of genes. P = Two-tailed students t-test. Boxes show the range from lower (25th percentile) to upper quartiles (75th percentile), with the median line (50th percentile); whiskers extend 1.5 times the inter-quartile range from bounds of box.
Extended Data Fig. 3 Comparison of in vitro and in vivo identified LADs.
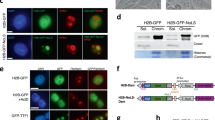
a, Venn diagram showing percent overlap of LAD coverage between brain NPCs and in vitro NPCs (DamID). b, Representative micrographs of DNA-FISH (red) with LaminB ICC (green) in brain NPCs and SVZ-cNSCs for four loci previously annotated as cLADs. Scale bar 2 𝜇m. Multiple images were taken to quantitate sub-nuclear localization of the DNA-FISH loci. c, Quantifications of DNA-FISH for loci shown in b. Percentages indicate loci within 0.5 μm (grey area) from the nuclear lamina. 50-60 nuclei were quantified for each locus. d, LaminB GO-CaRT tracks in brain NPCs, SVZ-cNSCs and in vitro NPCs (DamID) over cLAD DNA-FISH loci (dashed box) shown in b. e, LaminB GO-CaRT tracks of brain NPCs, E13 lung, E13 liver, SVZ-cNSCs and 3T3 MEFs. In vitro NPC and cLAD track (shown on top) are DamID derived6. Dashed box indicates a region where some previously annotated “cLADs” are not observed in tissues in vivo but detected by GO-CaRT in SVZ-NSCs and 3T3 MEFs. f, Venn diagram showing percent overlap of LAD coverage between cLADs and LADs common to brain, lung and liver.
Extended Data Fig. 4 Regional differences in LAD architecture of mouse brain NPCs.
a, Heatmap showing gene expression differences (row normalized) of marker genes in dorsal and ventral regions of the E13 mouse forebrain. R1 and R2 indicate two biological replicates. b, Venn diagram showing percent overlap of LAD coverage between dorsal and ventral NPCs. c, LaminB GO-CaRT tracks showing additional examples of brain region-specific LADs (dashed boxes). d, DNA-FISH micrographs of indicated loci in dorsal and ventral NPCs. Nuclear lamina is marked by LaminB1 staining (green). Scale bar 2 μm. Multiple images were taken to quantitate sub-nuclear localization of the DNA-FISH loci. e, Quantifications of DNA-FISH for the loci shown in c. Percentages indicate loci within 0.5 μm (grey area) from the nuclear lamina. 50-60 nuclei were quantified for each locus. f, LaminB GO-CaRT tracks in brain NPCs and in vitro NPCs (DamID) over DNA-FISH loci (dashed boxes) shown in d. g, Percent coverage of SINE and LINE repeat elements in brain region-specific LADs, common LADs and inter-LADs. h, Box plot depicting differential expression of genes that have quantitative differences in LaminB enrichment (top 20 percentile) between ventral vs. dorsal NPCs. LaminB decreased, n = 150, LaminB increased, n = 150. P = Two-tailed students t-test. Boxes show the range from lower (25th percentile) to upper quartiles (75th percentile), with the median line (50th percentile); whiskers extend 1.5 times the inter-quartile range from bounds of box. i, Top enriched GO-terms (biological process) of differentially expressed genes in dorsal- and ventral-specific LADs. GO terms were sorted based on their significance (-log10(q value)), the size of the bubble represents the gene ratio for each term.
Extended Data Fig. 5 Regional differences in LAD architecture of human brain NPCs.
a, Schematic of a GW20 human brain (coronal view) showing regions sampled for GO-CaRT and RNA-seq. b, Circos plot showing LaminB enrichment and LADs (horizontal bars) across the human chromosomes (1-22) in GW20 cortex and MGE. c, Heatmap showing gene expression differences (row normalized) of marker genes in GW20 cortex and MGE. R1 and R2 indicate two biological replicates. d, Jaccard similarity matrix of GO-CaRT LADs identified in human cortex and MGE and DamID LADs in indicated cultured cell lines19. e, Venn diagram showing percent overlap of LAD coverage between GW20 cortex and MGE. f, Representative LaminB GO-CaRT profiles of GW20 cortex and MGE. Dashed boxes show brain region-specific LADs. g, Genomic features of human brain region-specific LADs, common LADs and inter-LADs. h, Volcano plot showing differentially expressed genes in GW20 cortex- and MGE- specific LADs. Each dot represents a single gene with its differential gene expression level plotted on the x-axis and statistical significance (q < 0.05) on y-axis. P = Chi-square goodness of fit test. i, Top enriched GO-terms (biological process) of differentially expressed genes in cortex- and MGE- specific LADs. GO terms were sorted based on their significance (-log10(q value)), the size of the bubble represents the gene ratio for each term.
Extended Data Fig. 6 Chromatin context of active LAD genes in human cortex and H3K9me2-depleted LAD sub-domains in mouse cortex.
a, H3K9me2, LaminB, ATAC-seq and DNA methylation signal over TSS and surrounding regions for active genes in LADs (TPM > 1, red), repressed genes in LADs (TPM < 1, turquoise) and genes in inter-LADs (grey) in GW20 human cortex. b, Representative LaminB GO-CaRT tracks showing H3K9me2-depleted LAD sub-domains (marked by dashed boxes) in the mouse cortical/dorsal NPCs. c, Average H3K9me2 and LaminB signal over LADs (blue), H3K9me2-depleted LAD sub-domains (purple) and inter-LADs (grey) in mouse cortical NPCs scaled to the same relative size depicted by a solid horizontal bar. Traces in the 50Kb flanking proximal and distal regions are unscaled. d, Violin plot depicting the size in base pairs (log10) of LADs (n = 743) and H3K9me2-depleted LAD sub-domains (n = 664) identified in mouse cortical NPCs. e, Average gene expression in LADs (n = 3546), H3K9me2-depleted LAD sub-domains (n = 799) and inter-LADs (n = 18442) in mouse cortical NPCs, n = number of genes. P = Wilcoxon rank sum test, two-sided, non-adjusted. Boxes show the range from lower (25th percentile) to upper quartiles (75th percentile), with the median line (50th percentile); whiskers extend 1.5 times the inter-quartile range from bounds of box.
Extended Data Fig. 7 LAD conservation in cortex and MGE across macaque, human and mouse.
a, Circos plot showing LaminB enrichment and LADs (horizontal bars) across the macaque chromosomes (1-20) in PCD80 cortex and MGE. b, Percent LAD coverage in cortex and MGE across human, macaque and mouse. c, Alluvial plot depicting conserved LADs (blue) across human, macaque and mouse cortex using human as anchor species for liftover. Inter-LADs are shown in grey. d, A donut plot depicting conservation of mouse cortex/MGE shared LADs (brown) in macaque and human. Brain region-specific LADs are shown in white. e, Representative LaminB GO-CaRT profiles over LADs that are shared between cortex and MGE across human, macaque and mouse.
Extended Data Fig. 8 Features of neural and non-neural genes in conserved LADs.
a, Gene density in conserved LADs, all LADs, non-conserved LADs and inter-LADs identified based on mouse and human liftover. b, Top enriched GO-terms of genes in conserved LADs based on mouse liftover. GO terms were sorted based on their significance (-log10(q value)), the size of the bubble represents the gene ratio for each term. BP- Biological Process; MF- Molecular Function. c, Violin plot showing average gene expression in GW20 human cortex for all genes (red, n = 1365), neural genes (pink, n = 271) and non-neural genes (green, n = 1094) in conserved LADs. For comparison expression is also shown for genes in inter-LADs (grey, n = 17291). P = Wilcoxon rank sum test, two-sided, non-adjusted. d, LaminB signal (human GW20 cortex) over TSS and surrounding regions over the same gene sets as in c. e, Enriched motifs in neural and non-neural gene promoters in human. The size of the circle represents enrichment scores based on the P value from HOMER and color indicates the gene expression of the corresponding TFs in human GW20 cortex.
Extended Data Fig. 9 Gene expression and chromatin features of SPADs, inter-LAD, nonSPADs and LADs in human cortex.
a, Genome-wide scatter plot showing Pearson correlation between SON GO-CaRT in HEK293T cells and SON TSA-seq in K562 cells4. b, Circos plot showing SON GO-CaRT (green) and LaminB GO-CaRT profiles (blue) across the human chromosomes (1-22) in GW20 cortex. SPADs and LADs are shown by green and blue horizontal bars, respectively. c, Box plot depicting average gene expression in LADs (blue, n = 3105), non-speckle inter-LADs (grey, n = 9452) and SPADs (green, n = 8272) as determined by RNA-seq in GW20 human cortex, n = number of genes. P = Wilcoxon rank sum test, two-sided non-adjusted. Boxes show the range from lower (25th percentile) to upper quartiles (75th percentile), with the median line (50th percentile); whiskers extend 1.5 times the inter-quartile range from bounds of box. d, Average enrichment of various chromatin marks48 in LADs (blue), non-speckle inter-LADs (grey) and SPADs (green) in GW20 cortex. Mean enrichment of chromatin marks over each region scaled to the relative size of each region, which is depicted by the black horizontal. Traces in the 25 Kb flanking proximal and distal regions are unscaled.
Extended Data Fig. 10 DNA-FISH validation of SPADs and TF motif analyses of SPADs and LADs.
a, DNA-FISH micrographs of four SPAD loci in human GW20 cortical cells. Nuclear speckles are marked by SON ICC (green). Scale bar 2 μm, higher magnification 0.5 μm. b, SON and LaminB enrichment for SPAD loci shown in a. c, Quantifications of DNA-FISH for SPAD loci shown in a. Percentages indicate loci within 0.5 μm (grey area) from the nuclear speckles. 50-60 nuclei were quantified for each locus. d, Enriched motif in SPADs and LADs of GW20 cortex. The size of the circle represents enrichment scores based on the P value from HOMER and color indicates the gene expression of the corresponding TFs in GW20 cortex. e, Log odds ratio of enrichment of significant variants overlapping a given genomic compartment with 95% confidence intervals for T2D GWAS, where significant variants are defined as having a GWAS summary statistic of P < 1e-6 (left) or P < 1e-5 (right). Size of circle represents the fraction of significant variants overlapping a genomic compartment. Wald Test, two-sided non-adjusted. Bars represent standard error. LAD, n = 704, SPAD, n = 1047, inter-LAD, non-SPAD, n = 1810.
Supplementary information
Supplementary Tables 1–3
Supplementary Table 1: Sample information used in the study. Supplementary Table 2: Reproducibility of the biological replicates as calculated by Spearman correlation. Supplementary Table 3: Probes used for DNA-FISH and their genomic coordinates.
Rights and permissions
About this article
Cite this article
Ahanger, S.H., Delgado, R.N., Gil, E. et al. Distinct nuclear compartment-associated genome architecture in the developing mammalian brain. Nat Neurosci 24, 1235–1242 (2021). https://doi.org/10.1038/s41593-021-00879-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-021-00879-5
This article is cited by
-
Identifying quantitatively differential chromosomal compartmentalization changes and their biological significance from Hi-C data using DARIC
BMC Genomics (2023)
-
3D chromatin architecture and transcription regulation in cancer
Journal of Hematology & Oncology (2022)
-
The wide and growing range of lamin B-related diseases: from laminopathies to cancer
Cellular and Molecular Life Sciences (2022)
-
Chromatin domain alterations linked to 3D genome organization in a large cohort of schizophrenia and bipolar disorder brains
Nature Neuroscience (2022)