Abstract
Pyramidal cells and GABAergic interneurons fire together in balanced cortical networks. In contrast to this general rule, we describe a distinct neuron type in mice and rats whose spiking activity is anti-correlated with all principal cells and interneurons in all brain states but, most prevalently, during the down state of non-REM (NREM) sleep. We identify these down state-active (DSA) neurons as deep-layer neocortical neurogliaform cells that express ID2 and Nkx2.1 and are weakly immunoreactive to neuronal nitric oxide synthase. DSA neurons are weakly excited by deep-layer pyramidal cells and strongly inhibited by several other GABAergic cell types. Spiking of DSA neurons modified the sequential firing order of other neurons at down–up transitions. Optogenetic activation of ID2+Nkx2.1+ interneurons in the posterior parietal cortex during NREM sleep, but not during waking, interfered with consolidation of cue discrimination memory. Despite their sparsity, DSA neurons perform critical physiological functions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data that support the main findings of this study are publicly available in the Buzsaki Lab Databank: https://buzsakilab.com/wp/public-data/.
Code availability
All custom code is freely available on the Buzsáki Laboratory repository: https://github.com/buzsakilab/buzcode.
References
Buzsaki, G. et al. Nucleus basalis and thalamic control of neocortical activity in the freely moving rat. J. Neurosci. 8, 4007–4026 (1988).
Steriade, M., Nunez, A. & Amzica, F. A novel slow (< 1 Hz) oscillation of neocortical neurons in vivo: depolarizing and hyperpolarizing components. J. Neurosci. 13, 3252–3265 (1993).
Sanchez-Vives, M. V. & McCormick, D. A. Cellular and network mechanisms of rhytmic recurrent activity in neocortex. Nat. Neurosci. 3, 1027–1034 (2000).
Hasenstaub, A. et al. Inhibitory postsynaptic potentials carry synchronized frequency information in active cortical networks. Neuron 47, 423–435 (2005).
Steriade, M. & Timofeev, I. Neuronal plasticity in thalamocortical networks during sleep and waking oscillations. Neuron 37, 563–576 (2003).
Tononi, G. & Cirelli, C. Sleep and the price of plasticity: from synaptic and cellular homeostasis to memory consolidation and integration. Neuron 81, 12–34 (2014).
Takehara-Nishiuchi, K. & McNaughton, B. L. Spontaneous changes of neocortical code for associative memory during consolidation. Science 322, 960–963 (2008).
Todorova, R. & Zugaro, M. Isolated cortical computations during delta waves support memory consolidation. Science 366, 377–381 (2019).
Massimini, M., Huber, R., Ferrarelli, F., Hill, S. & Tononi, G. The sleep slow oscillation as a traveling wave. J. Neurosci. 24, 6862–6870 (2004).
Luczak, A., Barthó, P., Marguet, S. L., Buzsáki, G. & Harris, K. D. Sequential structure of neocortical spontaneous activity in vivo. Proc. Natl Acad. Sci. USA 104, 347–352 (2007).
Gerashchenko, D. et al. Identification of a population of sleep-active cerebral cortex neurons. Proc. Natl Acad. Sci. USA 105, 10227–10232 (2008).
Morairty, S. R. et al. A role for cortical nNOS/NK1 neurons in coupling homeostatic sleep drive to EEG slow wave activity. Proc. Natl Acad. Sci. USA 110, 20272–20277 (2013).
Zielinski, M. R. et al. Somatostatin+/nNOS+ neurons are involved in delta electroencephalogram activity and cortical-dependent recognition memory. Sleep 42, 1–16 (2019).
Compte, A., Sanchez-Vives, M. V., McCormick, D. A. & Wang, X. J. Cellular and network mechanisms of slow oscillatory activity (<1 Hz) and wave propagations in a cortical network model. J. Neurophysiol. 89, 2707–2725 (2003).
Jercog, D. et al. UP-DOWN cortical dynamics reflect state transitions in a bistable network. eLife 6, e22425 (2017).
Senzai, Y., Fernandez-Ruiz, A. & Buzsáki, G. Layer-specific physiological features and interlaminar interactions in the primary visual cortex of the mouse. Neuron 101, 500–513(2019).
Watson, B. O., Levenstein, D., Greene, J. P., Gelinas, J. N. & Buzsáki, G. Network homeostasis and state dynamics of neocortical sleep. Neuron 90, 839–852 (2016).
Okun, M. et al. Diverse coupling of neurons to populations in sensory cortex. Nature 521, 511–515 (2015).
English, D. F. et al. Pyramidal cell–interneuron circuit architecture and dynamics in hippocampal networks. Neuron 96, 505–520 (2017).
Tremblay, R., Lee, S. & Rudy, B. GABAergic interneurons in the neocortex: from cellular properties to circuits. Neuron 91, 260–292 (2016).
Tasic, B. et al. Shared and distinct transcriptomic cell types across neocortical areas. Nature 563, 72–78 (2018).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Krienen, F. M. et al. Innovations present in the primate interneuron repertoire. Nature 586, 262–269 (2020).
Oláh, S. et al. Regulation of cortical microcircuits by unitary GABA-mediated volume transmission. Nature 461, 1278–1281 (2009).
Overstreet-Wadiche, L. & McBain, C. J. Neurogliaform cells in cortical circuits. Nat. Rev. Neurosci. 16, 458–468 (2015).
Schuman, B. et al. Four unique interneuron populations reside in neocortical layer 1. J. Neurosci. 39, 125–139 (2019).
Jiang, X. et al. Principles of connectivity among morphologically defined cell types in adult neocortex. Science 350, aac9462 (2015).
Cadwell, C. R. et al. Electrophysiological, transcriptomic and morphologic profiling of single neurons using Patch-seq. Nat. Biotechnol. 34, 199–203 (2016).
Szabadics, J., Tamás, G. & Soltesz, I. Different transmitter transients underlie presynaptic cell type specificity of GABAA,slow and GABAA,fast. Proc. Natl Acad. Sci. USA 104, 14831–14836 (2007).
Goldberg, E. M. et al. K+ channels at the axon initial segment dampen near-threshold excitability of neocortical fast-spiking GABAergic interneurons. Neuron 58, 387–400 (2008).
Karube, F., Kubota, Y. & Kawaguchi, Y. Axon branching and synaptic bouton phenotypes in GABAergic nonpyramidal cell subtypes. J. Neurosci. 24, 2853–2865 (2004).
Peyrache, A., Battaglia, F. P. & Destexhe, A. Inhibition recruitment in prefrontal cortex during sleep spindles and gating of hippocampal inputs. Proc. Natl Acad. Sci. USA 108, 17207–17212 (2011).
Levenstein, D., Buzsáki, G. & Rinzel, J. NREM sleep in the rodent neocortex and hippocampus reflects excitable dynamics. Nat. Commun. 10, 1–12 (2019).
Harvey, C. D., Coen, P. & Tank, D. W. Choice-specific sequences in parietal cortex during a virtual-navigation decision task. Nature 484, 62–68 (2012).
Hoffman, K. L. & McNaughton, B. L. Coordinated reactivation of distributed memory traces in primate neocortex. Science 297, 2070–2073 (2002).
Kubota, Y., Hattori, R. & Yui, Y. Three distinct subpopulations of GABAergic neurons in rat frontal agranular cortex. Brain Res. 649, 159–173 (1994).
Perrenoud, Q., Rossier, J., Geoffroy, H., Vitalis, T. & Gallopin, T. Diversity of GABAergic interneurons in layer VIa and VIb of mouse barrel cortex. Cereb. Cortex 23, 423–441 (2013).
Tamás, G., Lörincz, A., Simon, A. & Szabadics, J. Identified sources and targets of slow inhibition in the neocortex. Science 299, 1902–1905 (2003).
Craig, M. T. & McBain, C. J. The emerging role of GABAB receptors as regulators of network dynamics: fast actions from a ‘slow’ receptor? Curr. Opin. Neurobiol. 26, 15–21 (2014).
Tricoire, L. et al. Common origins of hippocampal ivy and nitric oxide synthase expressing neurogliaform cells. J. Neurosci. 30, 2165–2176 (2010).
Niquille, M. et al. Neurogliaform cortical interneurons derive from cells in the preoptic area. eLife 7, e32017 (2018).
Taniguchi, H., Lu, J. & Huang, Z. J. The spatial and temporal origin of chandelier cells in mouse neocortex. Science 339, 70–74 (2013).
Van Der Werf, Y. D., Witter, M. P. & Groenewegen, H. J. The intralaminar and midline nuclei of the thalamus. Anatomical and functional evidence for participation in processes of arousal and awareness. Brain Res. Rev. 39, 107–140 (2002).
Brombas, A., Fletcher, L. N. & Williams, S. R. Activity-dependent modulation of layer 1 inhibitory neocortical circuits by acetylcholine. J. Neurosci. 34, 1932–1941 (2014).
Olsen, S. R., Bortone, D. S., Adesnik, H. & Scanziani, M. Gain control by layer six in cortical circuits of vision. Nature 483, 47–54 (2012).
Stark, E., Koos, T. & Buzsáki, G. Diode probes for spatiotemporal optical control of multiple neurons in freely moving animals. J. Neurophysiol. 108, 349–363 (2012).
Pachitariu, M., Steinmetz., N. A., Kadir, S. N., Carandini, M. & Harris, K. D. Fast and accurate spike sorting of high-channel count probes with KiloSort. Adv. Neural Inform. Process. Sys. 29, 4448–4456 (2016).
Valero, M. et al. Mechanisms for selective single-cell reactivation during offline sharp-wave ripples and their distortion by fast ripples. Neuron 94, 1234–1247.e7 (2017).
Navas-Olive, A. et al. Multimodal determinants of phase-locked dynamics across deep-superficial hippocampal sublayers during theta oscillations. Nat. Commun. 11, 2217 (2020).
Stark, E. & Abeles, M. Unbiased estimation of precise temporal correlations between spike trains. J. Neurosci. Methods 179, 90–100 (2009).
Barrio-Alonso, E., Fontana, B., Valero, M. & Frade, J. M. Pathological aspects of neuronal hyperploidization in Alzheimer’s disease evidenced by computer simulation. Front. Genet. 11, 287 (2020).
Stimberg, M., Brette, R. & Goodman, D. F. M. Brian 2, an intuitive and efficient neural simulator. eLife 8, e47314 (2019).
Valero, M. et al. Determinants of different deep and superficial CA1 pyramidal cell dynamics during sharp-wave ripples. Nat. Neurosci. 18, 1281–1290 (2015).
Viney, T. J. et al. Network state-dependent inhibition of identified hippocampal CA3 axo-axonic cells in vivo. Nat. Neurosci. 16, 1802–1811 (2013).
Salib, M. et al. GABAergic medial septal neurons with low-rhythmic firing innervating the dentate gyrus and hippocampal area CA3. J. Neurosci. 39, 4527–4549 (2019).
Hovde, K., Gianatti, M., Witter, M. P. & Whitlock, J. R. Architecture and organization of mouse posterior parietal cortex relative to extrastriate areas. Eur. J. Neurosci. 49, 1313–1329 (2019).
Feng, L., Zhao, T. & Kim, J. Neutube 1.0: a new design for efficient neuron reconstruction software based on the SWC format. eNeuro 2, ENEURO.0049-14.2014 (2015).
Acknowledgements
We would like to thank L. Menéndez de la Prida, K. McClain, D. Levenstein and A. Fernández-Ruiz and the rest of the members of the Buzsáki Laboratory for helpful comments on the manuscript. This work was supported by the European Molecular Biology Organization (EMBO) postdoctoral fellowship (EMBO ALTF 1161-2017), a Human Frontiers Science Program postdoctoral fellowship (LT0000717/2018) to M.V., the UK Medical Research Council (MR/R011567/1) to T.J.V., a Leon Levy Neuroscience Fellowship to I.Z., P01NS074972, R01NS110079, R01NS107257 and F31NS106793 to B.R. and NIH MH54671, NIH MH107396, NS 090583, NSF PIRE (grant no. 1545858), U19 NS107616 and U19 NS104590.
Author information
Authors and Affiliations
Contributions
M.V. and G.B. designed the experiments. M.V., S.M., I.Z. and Y.S. performed the in vivo physiological experiments. B.S. performed the in vitro physiological experiments. T.J.V. and M.V. did the anatomical characterization of DSA neurons. S.M. and I.Z. helped with behavioral testing. R.M. and B.R. designed and performed the breeding of the ID2/Nkx.21 line. M.V. generated in silico experiments and analyzed results. G.B. and M.V. wrote the paper with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Neuroscience thanks Giulio Tononi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Unit classification and physiological properties of DOWN-State Active (DSA) neurons.
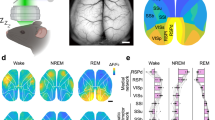
a, Units (n = 2,279 from 11 mice) were first classified based on trough-to-peak latency and firing rate. Each dot corresponds to one unit. Units with trough-to-peak latency < 0.55 ms were tentatively classified as narrow-waveform putative interneurons (INT; blue). Wide waveform units were grouped into putative excitatory cells (PYR cells; red) and inhibitory interneurons (INT; blue), on the basis of the bimodality of the marginal distribution of trough-to-peak latencies (top histogram), firing rate (right histogram) and burstiness (dot size) (Methods). Optogenetically identified neurons (Sst, PV, nNOS and ID2/Nkx2.1) are superimposed on the clouds. PV cells were recorded from V1. b, DOWN-state rate (P < 10−101, KW test), DOWN-state participation (P < 10−69, KW test) and UP-state rate (P < 10−136, KW test) distributions for putative pyramidal cells (PYR, red), interneurons (INT, blue) and DSA neurons (gray). c, Similar layout as in C but for REM sleep (left) and walking behavior (right). d, Peri-DOWN-state Z scored firing raster plot for all putative principal cells (PYR, left) and putative interneurons (INT, middle) for events detected during quiet waking state, as ranked according to Fig. 1D. Dashed lines delimit the ±50 ms window used to estimate unit responses. e, DSA neurons in the rat PFC (left) and in mouse primary visual cortex, V1 (right). Left (top to bottom): spike auto-correlogram and spike-waveform, peri-DOWN-state raster plot and average spike cross-correlation (CCG) between a reference neuron and all other simultaneously recorded units in a single session (each row corresponds to spikes of a target neuron referenced to the spike of DSA neuron at time 0. Black line is average). Right, autocorrelogram of 6 V1 DSA neurons. Middle panel: cross-correlograms between DOWN-state (time 0 is the trough of DOWN state) and 6 DSA neurons. Bottom: average spike cross-correlation (CCG) between a reference DSA neuron (time zero) and all other simultaneously recorded units in that session. ***P < 0.001.
Extended Data Fig. 2 Network property statistics of DSA interneurons.
a, Statistical contrast matrices (two-sided Tukey’s test) for firing rates of pyramidal cells (PYR), interneurons (INT) and DSA interneurons during waking quiescence (QWake), NREM sleep, REM sleep and active behavior (Walk). b, Same layout and statistical comparisons as in A but for the average unit CCGs as a function of brain state. c, Pearson correlations (tested using a Student’s t distribution) between average CCG responses of NREM sleep, QWake, REM sleep and walking behavior for all groups. Note that spike vs population relationships are preserved across brain states. d, Top: Average joint Z-score rate density between 10% of the interneurons (100 shufflings), 90% remaining interneurons and pyramidal cells neurons (left) and between 10% of the pyramidal cells (100 shufflings), interneurons and remaining pyramidal neurons. Bottom: Spearman correlation and statistical contrast matrices (two-sided Tukey’s test) for interneuron and pyramidal neurons rate and Z-scored population for all shown joint histograms. e, Average (mean ± IC95) partial correlation values of the Z-scored rate for all groups (blue for ρDSA,INT controlling for PYR; red for ρDSA,PYR controlling for INT; magenta for ρPYR,INT controlling for DSA), after truncating high firing rate units to match median the spikes number of pyramidal cells (average from n = 32 sessions, P < 10−25, F(2, 2787) = 57.42, repeated measures ANOVA). f, Left: distribution of excitatory divergence in all groups. Only putative pyramidal units excited their postsynaptic target cells (incidence probability for all groups in top-inset; P < 10−110, χ2(2) = 504.24, χ2 test). Right: distribution of excitatory convergence for all cell groups (P < 10−67, χ2(2) = 306.15, χ2 test). DSA neurons have fewer excitatory connections than the interneurons group (P < 10−4, χ2(1) = 11.36, χ2 test). ***P < 0.001.
Extended Data Fig. 3 Mechanisms of DSA neuron firing during DOWN states – model results.
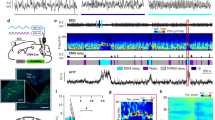
a, Spiking neural model containing 100 leaky DSA neurons receiving an asymmetric inhibitory/excitatory drive (top-left scheme). Bottom, UP/DOWN transitions with DOWN-selective firing DSA neurons (gray dots in the rastergram at the top). Top right, log firing rate distributions in the model corresponded to the those of the recorded neurons. b, Peri-DOWN-state Z scored firing raster plot for all simulated principal cells (PYR, left) and interneurons (INT, right), including (incl) 10% of DSAn, ranked according to their event response. Bottom, Z-scored average CCG for each model neuron group (compare with Fig. 1e). The peri-DOWN state raster plots and unit cross-correlograms (Extended Data Fig. 3b) were similar to experimental results (compare with Fig. 1d, e) for both pyramidal and interneuron populations (Supp. Fig. 3C-D; PYR rate mean ± SD: 0.2 ± 0.09 Hz during DOWN, 1.31 ± 0.41 Hz during UP; P ~ 0; INT: 0 ± 0.1 Hz during DOWN, 8.38 ± 2.52 Hz during UP, P ~ 0) as well as for the DSA group (7.43 ± 0.90 Hz during DOWN, 1.29 ± 0.34 Hz during UP, P < 10−83; Kruskal-Wallis test). c, Magnitude of DOWN-state response as a function of the CCG response. Marginal CCG (top, n = 20.000, 5000 and 500 PYR, INT and DSAn; P ~ 0, KW test) and DOWN-state response (right, P ~ 0, KW test) distributions for simulated pyramidal cells, interneurons and DSA neurons in each group. d, DOWN-state/UP-state firing rate ratio (P ~ 0, KW test), DOWN-state rate (P ~ 0, KW test), DOWN-state participation (P ~ 0, KW test) and UP-state rate (P ~ 0, KW test) distributions for all simulated pyramidal cells (PYR, red), interneurons (INT, blue) and DSA neurons (gray). e, Average (mean ± IC95) partial correlation values of the Z-scored rate for DSA and INT (blue), DSA and PYR (red), and PYR and INT (magenta) for different values of inhibitory weight simulated in the INT-DSA connection (0 to −3 mV, n = 2 simulations of 5100 cells per condition; P ~ 0, F(2, 44.094) = 278.36, repeated measures ANOVA). Model neurons showed that partial correlation between pyramidal cells and interneurons was positive at all time bins but negative for DSA neurons versus pyramidal cells and interneurons (compare with Fig. 2i), and it was strongly dependent on the inhibitory drive imposed on the DSA neurons by other inhibitory neurons. f, DSA neurons Z-scored DOWN-state responses (mean ± IC95) for different values of inhibition simulated on the DSA neurons (n = 200 DSAn per condition). DSA neurons rate during UP-states, but not during DOWN-states, depends on the inhibitory strength between INT and DSA neuron pairs. g, The peak delay of DSA neurons during DOWN-states correlates with the inhibitory strength of the INT-DSA connection (mean ± IC95, n = 2 simulations of 5100 cells per conditions; Pearson Correlations correlations tested using a Student’s t distribution). h, Partial correlation spectra (mean ± IC95) of the Z-scored simulated rate for each group as in Fig. 2d (n = 5 simulations of 5100 cells using a DSA Inh Drive = 1.5 mV; P ~ 0, F(2, 31.495) = 921.33, repeated measures ANOVA). i, Temporal dynamics of the Z-scored DOWN-state responses across all model neurons from H (top-left, mean ± IC95; P < 10−104, KW test). Simulated DSA neurons reach peak firing rate ~ 40 ms after maximum rate decrease of pyramidal cells and interneurons during DOWN-states (top-right; medians, interquartile ranges and error bars). Bottom, the average CCG delay between simulated DSA neuron spikes and spikes of other cells (bottom, dashed gray line; P < 10−315, KW test) matches DOWNs states responses (~40 ms, left avCCG average curves, right, medians, interquartile ranges, maxima and minima). j, Similar displays as in I but generated from recorded neurons (mean ± IC95; n = 1457, 775 and 47 PYR, INT and DSAn, respectively, from 11 mice; P < 10−22 for the temporal dynamic of the Z-scored DOWN-state responses and P < 10−7 for the average CCG delay, KW test). The trough of the cross-correlogram of DSA neurons versus other neurons occurred earlier than the peak of the cross-correlograms of other neurons as in the model. *P < 0.05; **P < 0.01; ***P < 0.001. For model details, see Methods.
Extended Data Fig. 4 Mechanisms of DSA neurons firing during the DOWN-states.
a, Top: Z-scored peristimulus histogram (PSTH) for cells recorded in the Sst::ChR animals (n = 2 mice), grouped as pyramidal cells (PYR, red), interneurons (INT, blue) and DSA neurons (gray). Horizontal bar: duration of optogenetic stimulation. Middle: firing rate trajectory for the DSA – INT response (P<10−29 between baseline and rebound epochs, KW test). Bottom: Firing trajectory for the DSA – PYR response (P < 10−11, KW test). b, Same than A but for PV::ChR animals (n = 4 mice; P=0.009 and P = 0.39 for ‘INT resp vs DSA resp’ and ‘PYR resp vs DSA resp’, respectively, KW test). Non-significant, n.s. c, Spike auto-correlogram (top, mean ± IC95) and spike waveform (bottom, mean ± IC95) of ID2/Nkx2.1-expressing DSA neurons (gray, n = 6) in optogenetic experiments are similar to non-responding DSA neurons (dark yellow, n = 3, ID2-/Nkx2.1- or non-recombined neurons) as well as to DSA neurons observed in other mice (black, n = 41; KW test). d, Depth distribution of all recorded units (as in Fig. 1i), highlighting the position of the nNOS+ neurons (yellow diamonds; n = 25 cells optogenetically responding cells from 4 Nos1::ChR mice). Note the deep position of the two DSA/nNOS+ neurons (encircled). e, Same layout as in D but highlighting the position of the ID2 + /Nkx2.1+ cells in black circles (n = 8 from 6 ID2/Nkx2.1::Ai80 mice). Note that the two ID2 + /Nkx2.1+ cells, residing above layer 5 (encircled), were not DSA neurons. **P < 0.01; ***P < 0.001, after Tukey’s honesty post hoc multiple comparisons.
Extended Data Fig. 5 Intrinsic electrophysiological features of L5/6 ID2/Nkx2.1 neurons.
a, Group differences (mean ± SD) for three electrophysiological parameters between L1 neurogliaform cells (green, n = 25 L1 NGFC), L5/6 ID2/Nkx2.1 (gray, n = 18 cells) and delayed fast spiking cells (in blue, from ref. 30; n = 23 dFS). Note that Goldberg et al used 600 ms second square pulses on the delayed respond tests. b, Example of rebound spiking test protocol. Average traces from a L5/6 ID2/Nkx2.1 cell held at −70 mV, then hyperpolarized to −90 mV for 3 s, then depolarized to either −70 mV (black) or −50 mV (red). If present, rebound spikes should be apparent immediately following the depolarizing step (n = 11 L5/6 ID2/Nkx2.1 cells).
Extended Data Fig. 6 Anatomical features of deep and superficial cortical ID2/Nkx2.1 interneurons.
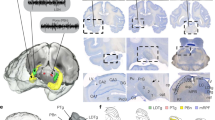
a, Depth distribution of ID2 + /Nkx2.1+ neurons (tdTomato+ or GFP+) in the medial prefrontal cortex (mPFC), PPC and anterior and posterior regions of the primary visual cortex (V1). b, Photographs showing distribution of GFP+ (or tdTomato+) neurons in the mPFC of an ID2/Nkx2.1::Ai80 mouse (Ai65) (top and middle images, reverse contrast epifluorescence; n = 47 neurons in 9 samples from 3 mice). Most neurons were found in layer 6. c, A layer 6 tdTomato+ ID2/Nkx2.1 neuron (red) at two different depth levels. Note the putative PV+ (cyan) puncta in close apposition to the tdTomato+ cell (arrows). Top, 0.38 μm-thick single optical section. Bottom, 0.76 μm-thick maximum intensity z-projection (representative micrograph, tested in 3 mice). d, Top: Z projections (confocal maximum-intensity projection) of two intracellularly-filled tdTomato+ lD2/Nkx2.1 neuron from two ID2/NKx2.1::Ai65 mouse. Bottom: reconstruction of an intracellularly-filled tdTomato+ lD2/Nkx2.1 neuron (from a 223.2 µm thick confocal stack, maximum-intensity projection as inset) of an ID2/NKx2.1::Ai65 mouse. The two reconstructions of the ID2/Nkx2.1 neurons showed similarities to neurogliaform cells in L2/324,29 and L126. Total dendritic length: 7968.5 µm and 11641 µm for the cell in Fig. 5d and the cell in Extended Data Fig. 6, respectively; total axonal length: 77873 µm and 53669 µm; membrane surface in dendrites and soma: 1.02 × 105 µm2 and 2.81 × 105 µm2; membrane surface in axons: 9.21 × 105 µm2 and 1.14 × 106 µm2; number of dendritic branches: 117 and 105; axonal branches: 747 and 314. e, ID2+/Nkx2.1+ neuron (red) in L6b weakly immunoreactive for both NPY and nNos (arrow; n = 64 tdTomato+ ID2/Nkx2.1 neuron from 3 mice were tested against NPY). Note also strongly immunoreactive NPY+ nNOS+ neuron lacking tdTomato expression (arrowhead on top). f, A tdTomato+ ID2/Nkx2.1 neuron in L5 of PPC immunoreactive for calretinin (white arrowheads) and lacking expression of Calbindin (widefield epifluorescence; n = 54 tdTomato+ ID2/Nkx2.1 neuron from 3 mice were tested, 2 of which were positive for calretinin). g, PV+ tdTomato+ neuron in L2 of V1 (representative micrograph, tested in 3 mice). h, Left: a layer 2 tdTomato+ ID2/Nkx2.1 neuron in layer 2 (widefield epifluorescence, reverse contrast). Right, high magnification of the marked rectangle area shows the tdTomato+ ID2/Nkx2.1 neuron axon distribution (‘cartridges’, some of them marked by arrows) typical of axo-axonic neurons (widefield epifluorescence, reverse contrast; representative micrograph, tested in 3 mice). The ID2/Nkx2.1 cells labeled here are identical to the Lamp5/Lhx6 type described in ref. 21,22. ID2 and Lamp5 are both broadly expressed in putative neurogliaform interneurons21, and Lhx6 is expressed downstream of Nkx2.1 in cortical interneurons. The proportion of the Lamp5/Lhx6 subtype has been reported to be 1.4 ± 0.2% of total interneurons22, with the majority located in L5/6.
Extended Data Fig. 7 Effector role of the DSA neurons.
a, Cross-correlation (CCG) between Non-Mod (reference, n = 648 units) and –Mod (yellow, n = 118 units) compared to the CCG within only Non-Mod cells (green) (top and bottom-left; mean ± IC95; P = 0.004, KW test), and CCG asymmetry [before-after]/[before+after] (bottom-right; medians, interquartile ranges, maxima and minima; P < 10−5, KW test). b, Amplitude response differences (medians, interquartile ranges, maxima and minima) for –Mod (n = 51 units from 9 session with at least 1 DSAn from 6 ID2/Nkx2.1::CatCh mice) and Non-Mod cells (n = 195 units) in LOW and HIGH DSA neuron rate DOWN-UP events. c, Temporal dynamics of the Z-scored DOWN-UP events responses for LOW and HIGH events (n = 2.954 and 5.317 events, respectively; medians, interquartile ranges, maxima and minima). d, Averages (mean ± IC95) of the Fourier-transform spectra for non-stimulated (control, black) and stimulated (green) epochs (top) during NREM (n = 12 sessions from 6 mice). Peak frequency, peak amplitude and delta power (0.5–4 Hz) comparisons for non-stimulated and stimulated epochs (bottom). Note decreased peak frequency of in the delta band during stimulated epochs, likely reflecting the prolongation of DOWN state upon activation of ID2/Nkx2.1 neurons (see Fig. 6). e, Bottom, Z-scored sequential activity of a simultaneously recorded population during DOWN-UP events sorted for ‘High DSA rate’ events (session mean template), vertically arranged by latency. Top, session mean template for ‘Low DSA rate’ events but arranged according to ‘High DSA rate’ classification (see Fig. 6f for ‘Low DSA rate’). Note that the two types of event sequences are different when DSA spiking activity in the slow oscillation event is high or low. *p < 0.05; **p < 0.01; ***p < 0.001.
Extended Data Fig. 8 Stimulation of DSA neurons during NREM sleep.
a, Multitaper spectrogram from 0–40 Hz of one entire recording day with NREM optogenetic stimulation (including baseline, behavioral session 1, rest period and behavioral session 2) utilized for state scoring. Middle, electromyogram (EMG) over the same session (au, arbitrary units). Bottom, state scoring of the same session (as in ref. 17). States are coded as label to the left. b, Three-dimensional plot showing state segregations. Each point corresponds to 1 s of recording time, with colors indicating the identified state during that second as labeled. c, d, Same than A-B but for a no-simulation (control) day. e, NREM sleep durations (mean ± SD) between test sessions 1 and 2 are similar in the four different conditions. f, Normalized fractions of time spent in Wake, NREM and REM in the home cage between behavioral sessions 1 and 2. g, CCG trace (mean ± IC95) between light pulses and UP and DOWN states (left) for all NREM stimulated sessions (n = 15 sessions). Right, fraction of stimulated events over all detected events (medians, interquartile ranges, maxima and minima; P < 10−5, KW test). ***p < 0.001.
Supplementary information
Rights and permissions
About this article
Cite this article
Valero, M., Viney, T.J., Machold, R. et al. Sleep down state-active ID2/Nkx2.1 interneurons in the neocortex. Nat Neurosci 24, 401–411 (2021). https://doi.org/10.1038/s41593-021-00797-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-021-00797-6
This article is cited by
-
Wake slow waves in focal human epilepsy impact network activity and cognition
Nature Communications (2023)
-
The role of superficial and deep layers in the generation of high frequency oscillations and interictal epileptiform discharges in the human cortex
Scientific Reports (2023)
-
Somatostatin neurons in prefrontal cortex initiate sleep-preparatory behavior and sleep via the preoptic and lateral hypothalamus
Nature Neuroscience (2023)
-
Preconfigured dynamics in the hippocampus are guided by embryonic birthdate and rate of neurogenesis
Nature Neuroscience (2022)
-
Cortical regulation of two-stage rapid eye movement sleep
Nature Neuroscience (2022)