Abstract
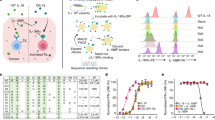
Many immunotherapies act by enhancing the ability of cytotoxic T cells to kill tumor cells. Killing depends on T cell recognition of antigens presented by class I major histocompatibility complex (MHC-I) proteins on tumor cells. In this study, we showed that medulloblastomas lacking the p53 tumor suppressor do not express surface MHC-I and are therefore resistant to immune rejection. Mechanistically, this is because p53 regulates expression of the peptide transporter Tap1 and the aminopeptidase Erap1, which are required for MHC-I trafficking to the cell surface. In vitro, tumor necrosis factor (TNF) or lymphotoxin-β receptor agonist can rescue expression of Erap1, Tap1 and MHC-I on p53-mutant tumor cells. In vivo, low doses of TNF prolong survival and synergize with immune checkpoint inhibitors to promote tumor rejection. These studies identified p53 as a key regulator of immune evasion and suggest that TNF could be used to enhance sensitivity of tumors to immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data are available in the main paper or the supplementary materials. Raw data for all figures are available as Source Data files. The human medulloblastoma sequencing data used in Fig. 3f,g are deposited in the European Genome-phenome Archive as EGAD00001001899 and EGAD00001004958; the data are also available from the authors upon request in the interim.
Change history
14 December 2021
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1038/s41593-021-00994-3
References
Tabori, U. et al. Universal poor survival in children with medulloblastoma harboring somatic TP53 mutations. J. Clin. Oncol. 28, 1345–1350 (2010).
Hill, R. M. et al. Combined MYC and P53 defects emerge at medulloblastoma relapse and define rapidly progressive, therapeutically targetable disease. Cancer Cell 27, 72–84 (2015).
Morrissy, A. S. et al. Divergent clonal selection dominates medulloblastoma at recurrence. Nature 529, 351–357 (2016).
Kantoff, P. W. et al. Sipuleucel-T immunotherapy for castration-resistant prostate cancer. N. Engl. J. Med. 363, 411–422 (2010).
Miller, J. F. & Sadelain, M. The journey from discoveries in fundamental immunology to cancer immunotherapy. Cancer Cell 27, 439–449 (2015).
Rosenberg, S. A. & Restifo, N. P. Adoptive cell transfer as personalized immunotherapy for human cancer. Science 348, 62–68 (2015).
Topalian, S. L., Drake, C. G. & Pardoll, D. M. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell 27, 450–461 (2015).
Braun, D. A., Burke, K. P. & Van Allen, E. M. Genomic Approaches to Understanding Response and Resistance to Immunotherapy. Clin. Cancer Res. 22, 5642–5650 (2016).
ClinicalTrials.gov. Immune checkpoint inhibitor nivolumab in people with select rare CNS cancers. NCT03173950 (2017).
ClinicalTrials.gov. Vaccine immunotherapy for recurrent medulloblastoma and primitive neuroectodermal tumor. NCT01326104 (2020).
Pei, Y. et al. HDAC and PI3K antagonists cooperate to inhibit growth of MYC-driven medulloblastoma. Cancer Cell 29, 311–323 (2016).
Pei, Y. et al. An animal model of MYC-driven medulloblastoma. Cancer Cell 21, 155–167 (2012).
Bowman, T. et al. Tissue-specific inactivation of p53 tumor suppression in the mouse. Genes Dev. 10, 826–835 (1996).
Northcott, P. A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014).
Zurita, E. et al. Genetic polymorphisms among C57BL/6 mouse inbred strains. Transgenic Res. 20, 481–489 (2011).
Cortez, M.A. et al. PDL1 regulation by p53 via miR-34. J. Natl Cancer Inst. 108, dvj303 (2016).
Wang, X. et al. Tumor suppressor miR-34a targets PD-L1 and functions as a potential immunotherapeutic target in acute myeloid leukemia. Cell Signal. 27, 443–452 (2015).
Milde, T. et al. HD-MB03 is a novel Group 3 medulloblastoma model demonstrating sensitivity to histone deacetylase inhibitor treatment. J. Neurooncol. 110, 335–348 (2012).
Yang, Z. J. et al. Medulloblastoma can be initiated by deletion of Patched in lineage-restricted progenitors or stem cells. Cancer Cell 14, 135–145 (2008).
Zhu, K. et al. p53 induces TAP1 and enhances the transport of MHC class I peptides. Oncogene 18, 7740–7747 (1999).
Wang, B., Niu, D., Lai, L. & Ren, E. C. p53 increases MHC class I expression by upregulating the endoplasmic reticulum aminopeptidase ERAP1. Nat. Commun. 4, 2359 (2013).
Leone, P. et al. MHC class I antigen processing and presenting machinery: organization, function, and defects in tumor cells. J. Natl Cancer Inst. 105, 1172–1187 (2013).
Palmer, K. J., Harries, M., Gore, M. E. & Collins, M. K. Interferon-ɑ (IFN-ɑ) stimulates anti-melanoma cytotoxic T lymphocyte (CTL) generation in mixed lymphocyte tumour cultures (MLTC). Clin. Exp. Immunol. 119, 412–418 (2000).
Neumann, H., Schmidt, H., Cavalie, A., Jenne, D. & Wekerle, H. Major histocompatibility complex (MHC) class I gene expression in single neurons of the central nervous system: differential regulation by interferon (IFN)-γ and tumor necrosis factor (TNF)-ɑ. J. Exp. Med. 185, 305–316 (1997).
Slatter, T. L. et al. Antitumor cytotoxicity induced by bone-marrow-derived antigen-presenting cells is facilitated by the tumor suppressor protein p53 via regulation of IL-12. Oncoimmunology 5, e1112941 (2016).
Campbell, I. L., Oxbrow, L., West, J. & Harrison, L. C. Regulation of MHC protein expression in pancreatic β-cells by interferon-γ and tumor necrosis factor-ɑ. Mol. Endocrinol. 2, 101–107 (1988).
Mattsson, R. et al. Placental MHC class I antigen expression is induced in mice following in vivo treatment with recombinant interferon-γ. J. Reprod. Immunol. 19, 115–129 (1991).
Lavi, E. et al. Tumor necrosis factor induces expression of MHC class I antigens on mouse astrocytes. J. Neuroimmunol. 18, 245–253 (1988).
Andrews, J. S., Berger, A. E. & Ware, C. F. Characterization of the receptor for tumor necrosis factor (TNF) and lymphotoxin (LT) on human T lymphocytes. TNF and LT differ in their receptor binding properties and the induction of MHC class I proteins on a human CD4+ T cell hybridoma. J. Immunol. 144, 2582–2591 (1990).
Locksley, R. M., Killeen, N. & Lenardo, M. J. The TNF and TNF receptor superfamilies: integrating mammalian biology. Cell 104, 487–501 (2001).
Chopra, M. et al. Exogenous TNFR2 activation protects from acute GvHD via host T reg cell expansion. J. Exp. Med. 213, 1881–1900 (2016).
Lawrence, T. The nuclear factor NF-κB pathway in inflammation. Cold Spring Harb. Perspect. Biol. 1, a001651 (2009).
Ciancio, M. J., Hunt, J., Jones, S. B. & Filkins, J. P. Comparative and interactive in vivo effects of tumor necrosis factor ɑ and endotoxin. Circ. Shock 33, 108–120 (1991).
Biesmans, S. et al. Peripheral administration of tumor necrosis factor-ɑ induces neuroinflammation and sickness but not depressive-like behavior in mice. Biomed. Res. Int. 2015, 716920 (2015).
Sonabend, A. M. et al. Medulloblasoma: challenges for effective immunotherapy. J. Neurooncol. 108, 1–10 (2012).
Sade-Feldman, M. et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat. Commun. 8, 1136 (2017).
Paulson, K. G. et al. Acquired cancer resistance to combination immunotherapy from transcriptional loss of class I HLA. Nat. Commun. 9, 3868 (2018).
Roberts, N. J., Zhou, S., Diaz, L. A. Jr. & Holdhoff, M. Systemic use of tumor necrosis factor ɑ as an anticancer agent. Oncotarget 2, 739–751 (2011).
Reagan-Shaw, S., Nihal, M. & Ahmad, N. Dose translation from animal to human studies revisited. FASEB J. 22, 659–661 (2008).
Furman, W. L. et al. Phase I clinical trial of recombinant human tumor necrosis factor in children with refractory solid tumors: a Pediatric Oncology Group study. J. Clin. Oncol. 11, 2205–2210 (1993).
Seibel, N. L. et al. Phase I study of tumor necrosis factor-ɑ and actinomycin D in pediatric patients with cancer: a Children’s Cancer Group study. J. Immunother. Emphasis Tumor Immunol. 16, 125–131 (1994).
Suzuki, H., et al. Recurrent non-coding U1-snRNA mutations drive cryptic splicing in Shh medulloblastoma. Nature https://dx.doi.org/10.1038/s41586-019-1650-0 (2019).
Futterer, A., Mink, K., Luz, A., Kosco-Vilbois, M. H. & Pfeffer, K. The lymphotoxin β receptor controls organogenesis and affinity maturation in peripheral lymphoid tissues. Immunity 9, 59–70 (1998).
Banks, T. A. et al. A lymphotoxin-IFN-β axis essential for lymphocyte survival revealed during cytomegalovirus infection. J. Immunol. 174, 7217–7225 (2005).
Hong, H. et al. Suppression of induced pluripotent stem cell generation by the p53–p21 pathway. Nature 460, 1132–1135 (2009).
Kojima, K. et al. p53 activation of mesenchymal stromal cells partially abrogates microenvironment-mediated resistance to FLT3 inhibition in AML through HIF-1ɑ-mediated down-regulation of CXCL12. Blood 118, 4431–4439 (2011).
Tsuchiya, S. et al. Establishment and characterization of a human acute monocytic leukemia cell line (THP-1). Int. J. Cancer 26, 171–176 (1980).
Sundstrom, C. & Nilsson, K. Establishment and characterization of a human histiocytic lymphoma cell line (U-937). Int. J. Cancer 17, 565–577 (1976).
Smardova, J., Pavlova, S., Svitakova, M., Grochova, D. & Ravcukova, B. Analysis of p53 status in human cell lines using a functional assay in yeast: detection of new non-sense p53 mutation in codon 124. Oncol. Rep. 14, 901–907 (2005).
Weisberg, E. et al. Inhibition of wild-type p53-expressing AML by the novel small molecule HDM2 inhibitor CGM097. Mol. Cancer Ther. 14, 2249–2259 (2015).
Misuraca, K. L. et al. Pax3 expression enhances PDGF-B-induced brainstem gliomagenesis and characterizes a subset of brainstem glioma. Acta Neuropathol. Commun. 2, 134 (2014).
Lewis, P. W. et al. Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340, 857–861 (2013).
Hashizume, R. et al. Characterization of a diffuse intrinsic pontine glioma cell line: implications for future investigations and treatment. J. Neurooncol. 110, 305–313 (2012).
Caretti, V. et al. Human pontine glioma cells can induce murine tumors. Acta Neuropathol. 127, 897–909 (2014).
Kreft, L. et al. ConTra v3: a tool to identify transcription factor binding sites across species, update 2017. Nucleic Acids Res. 45, W490–W494 (2017).
Acknowledgements
The authors gratefully acknowledge the Animal Facilities at UCSD and SBP for help with animal care and husbandry; S. Tacheva Grigorova for assistance generating conditional Ptch1 knockout mice; Y. Altman of the Flow Cytometry Shared Resource at SBP; and J. Olvera and C. Fine of the Stem Cell Core at UCSD for assistance with FACS sorting and analysis. We are also indebted to T. Milde at the Hopp Children’s Cancer Center, Heidelberg, for providing HD-MB03 and BT-084 cells; to M. Monje at Stanford University for sharing human DIPG cell lines; and to J. Scott-Browne and A. Rao at the La Jolla Institute for Immunology for providing p53 shRNA constructs. We thank S. Baker and L. Hover at St. Jude Children’s Research Hospital, L. Attardi at Stanford University, O. Ayrault and A. Forget at Institut Curie, and J. Rich, B. Prager and L. Kim from UCSD for help with analysis of gene and protein expression in p53-mutant and wild-type tumors. M.D.T. is supported by the National Institutes of Health (NIH) (R01CA148699 and R01CA159859), the Pediatric Brain Tumour Foundation, the Terry Fox Research Institute, the Canadian Institutes of Health Research, the Cure Search Foundation, b.r.a.i.n. child, Meagan’s Walk, the SWIFTY Foundation, the Brain Tumour Charity, Genome Canada, Genome BC, Genome Quebec, the Ontario Research Fund, Worldwide Cancer Research, the V Foundation for Cancer Research and the Ontario Institute for Cancer Research through funding provided by the Government of Ontario. M.D.T. is also supported by a Canadian Cancer Society Research Institute Impact grant, a Cancer Research UK Brain Tumour Award and a Stand Up To Cancer (SU2C) St. Baldrick’s Pediatric Dream Team Translational Research Grant (SU2C-AACR-DT1113) and SU2C Canada Cancer Stem Cell Dream Team Research Funding (SU2C-AACR-DT-19-15) provided by the Government of Canada through Genome Canada and the Canadian Institutes of Health Research, with supplementary support from the Ontario Institute for Cancer Research through funding provided by the Government of Ontario. M.D.T. is also supported by the Garron Family Chair in Childhood Cancer Research at the Hospital for Sick Children and the University of Toronto. This work was supported by funding from the National Cancer Institute (2R01 CA159859-06 and P30 CA30199 (to R.J.W.-R.)), P01 CA177322 (to C.F.W.), K22 CDA CA229613 (to S.H), NIH (R01 AI048073) (to C.F.W.), Forschungsgemeinschaft (DFG, German Research Foundation) (324392634–TRR221 and WA 1025/31-1) (to H.W.), the V Foundation, the Pediatric Brain Tumor Foundation, Accelerate Brain Cancer Cure, Alex’s Lemonade Stand Foundation (to R.J.W.-R. and O.J.B.), William’s Superhero Fund (to R.J.W.-R.), the McDowell Charity Trust (to R.J.W.-R.), Madox’s Warriors (to O.J.B.), the Cristian Rivera Foundation (to O.J.B.), the Fly A Kite Foundation (to O.J.B.) and the California Institute for Regenerative Medicine (to R.J.W.-R.). This work is dedicated to the memory of William Schultz and Errol McDowell.
Author information
Authors and Affiliations
Contributions
A.G. and R.J.W.-R. conceived and designed experiments. A.G., L.Q.C., M.B.M. and J.M.R. performed experiments. A.G. H.S., V.R., S.H., S.A.M., P.S., F.M.G.C., H.F., S.J.M.J., R.A.M., A.J.M., Y.M., N.T., Y.L. and M.M. analyzed data. L.Q., M.K., Y.D., P.B., J.J.H., J.R.C., M.M.R, O.J.B., M.L.L., J.M.O., Y.-J.C., A.M., L.C., A.J.D. and X.-N.L. provided patient samples, cell lines or xenografts and associated data. P.S.N., C.F.W. and H.W. provided reagents and protocols. P.S.N., F.C., L.M.B., C.F.W. and M.D.T. discussed results and strategy. R.J.W.-R. supervised the study. A.G. and R.J.W.-R. wrote the manuscript, which was revised and approved by all authors.
Corresponding author
Ethics declarations
Competing interests
The Sanford Burnham Prebys Medical Discovery Institute has filed a patent application with R.J.W.-R., A.G. and C.F.W. listed as inventors. PCT/US2018/048916 was filed on August 30, 2018, and is currently pending. This application covers the potential of p53 as a biomarker for responsiveness to T cell-based immunotherapy and the use of TNF receptor superfamily ligands to increase MHC expression and enhance immunotherapy.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The RNA sequencing data from human medulloblastoma samples are deposited in the European Genome-phenome Archive as EGAD00001001899 and EGAD00001004958.
This article has been retracted. Please see the retraction notice for more detail: https://doi.org/10.1038/s41593-021-00994-3
Extended data
Extended Data Fig. 1 Immune microenvironment in p53-mutant and wild-type murine medulloblastoma.
a–d, Intracranially transplanted MP tumors (black) and MG tumors (red) labelled with GFP were harvested 2, 5 and 7 days after transplant (n = 3 mice per tumor type per day). Percentages of B-cells (a, B220 positive), Macrophages (b, CD11b positive), Natural killer (NK) cells (c, NK1.1 positive) and cytotoxic T-cells (d, CD8 positive) were determined by FACS analysis. The average curve and single data points are shown for each condition. p-values were determined by two-sided unpaired t-test. In d, *P= 0.0014. e–g, Depletion of T-cells (αCD8, red line), or no T-cell depletion (Control, black line), was performed in aB6 MG tumor-bearing mice. (e) Bioluminescence imaging of representative mice. f, Quantification of the average bioluminescence signal (dark lines) and the signals for individual mice (light lines) for each group. i, Survival curves for control mice (n = 6); and αCD8-treated mice (n = 6). p-value for survival curve was determined by a two-sided log-rank (Mantel-Cox) test. All error bars represent the mean ± SD. h-i, MP tumor cells or MP tumor cells overexpressing GFI1 (MP+G) were transplanted into NSG or aB6 mice. Bioluminescence imaging of representative mice h, and survival curves (n = 10) i, are shown. p-values for the difference in survival between MP and MP+G transplanted into NSG or aB6 were determined by two-sided log-rank (Mantel-Cox) test. In i, NS (not significant), P= 0.608 (NSG) and P= 0.597 (aB6). j,k. Expression of T-cell suppression molecules j, and Dendritic cell and T-cell activation molecules k, in MP tumors (n = 3) and MG tumors (n = 3) were measured by qRT-PCR. Quantification of the relative expression of the genes for three independent tumors is shown; data points represent values for individual tumors. Bars represent means ± SD. p-values were determined by two-sided unpaired t-test. None of the differences between MP and MG were significant.
Extended Data Fig. 2 Knockdown of p53 decreases MHC-I expression in murine and human medulloblastoma cells.
a, FACS histogram of MHC-I expression in normal neural stem cells (NSCs). Isotype control is shown in gray and MHC-I staining in black. Quantification of the mean fluorescence intensity (MFI) values for three independent NSC samples (isolated from three separate litters of mice) is shown below the histogram; data points represent MFIs for individual NSC samples. p-value, determined by two-sided unpaired t-test, is indicated on bar graph. b,d, FACS histograms of MHC-I or HLA-I expression in murine MG tumor cells (b) and human HD-MB03 tumor cells (d) transduced with control shRNA (black histograms) or shRNAs that target sequences conserved in mouse and human p53 (shp53 #109, blue histograms; shp53 #110, pink histograms). Quantification of the MFI values for three independent tumors are shown below each histogram; data points represent MFIs for individual tumor samples. p-values were determined by two-sided unpaired t-test. p-values are indicated on each bar graph. c,e, Protein levels of p53 and β-Actin were determined by western blotting in MG tumor cells (c) and HD-MB03 cells (e). Western blots are cropped at the molecular weights for p53 (53 kDa) and β-actin (42 kDa); original blots are available in Source Data. f, Murine Ptch-KO MB tumor cells overexpressing dominant negative p53 (Ptch-KO MB-DNP53) were transplanted into NSG or aB6 mice. Survival curves (n = 8) are shown. p-values for the difference in survival between Ptch-KO MB and Ptch-KO MB-DNP53 were determined by two-sided log-rank (Mantel-Cox) test. NS (not significant), P= 0.6939 (NSG) and **P= 0.0021 (aB6). g, FACS histograms of MHC-I expression in Ptch-KO MB (black) and Ptch-KO MB-DNP53 (blue) tumor cells. Quantification of MFI values for three independent tumor samples is shown below the histogram; data points represent MFIs for each tumor sample. p-value, determined by two-sided unpaired t-test, is indicated on bar graph.
Extended Data Fig. 3 p53 status determines MHC-I expression in medulloblastoma patient-derived xenografts and DIPG cells.
(a–j) FACS histograms of HLA-I expression in human medulloblastoma PDXs expressing mutant p53 (upper panel, a-e) or wild-type p53 (lower panel, f-j. Isotype controls are shown in black and HLA-I staining in purple. e,j, Quantification of average mean fluorescence intensity (MFI) for triplicate samples of each human medulloblastoma PDX. Two-sided t-tests were used to evaluate the significance of the difference between HLA-I and isotype control staining for p53-mutant tumors (e, NS, not significant, P= 0.3793) and for p53-WT tumors (j, *P= 0.0112). k–n, FACS histograms of MHC-I expression in p53 wild-type (k, m) or p53 deficient (l and n) murine DIPG lines (k-l) and human DIPG lines (m,n). Isotype controls are shown in black and murine MHC-I and human HLA-I staining in purple. Quantification of the MFI values for three independent tumor samples are shown below each histogram; data points represent MFIs for individual tumor samples. p-values were determined by two-sided unpaired t-test and are indicated on each bar graph.
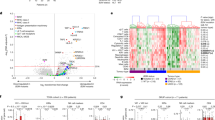
Extended Data Fig. 4 p53 mutations are associated with decreased ERAP1 expression in other cancers.
a,b, Expression of ERAP1 and TAP1 in p53-mutant relative to p53 wild type tumors in TCGA datasets. Cancer sites are indicated along the X-axis. Each column represents the fold change in expression in p53 mutant relative to p53 WT tumors. Positive values indicate increased expression of ERAP1 a, or TAP1 b, mRNA levels in p53 mutant tumors compared to p53 wild-type (WT) tumors; negative values indicate decreased expression of ERAP1 or TAP1 in p53 mutant tumors compared to WT tumors; 0 indicates no change in expression. Error bars indicate standard error of the mean. *P<0.05; **P<0.01; ***P<0.001; ns, not significant, based on two-sided Wilcoxon Rank Sum test. Precise p-/q-values and sample sizes are provided in Supplemental Table 1. (c,d) Box plots describing mRNA expression of ERAP1 c, and TAP1 d, in patient tumors from TCGA datasets (AML, Head and Neck, lung and stomach cancer) with p53-mutant samples shown in red and p53-wild type (WT) samples shown in blue. Box plot center lines show median, box limits indicate the 25th and 75th percentiles, lower and upper whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles, respectively. p-values represent significance, based on two-sided Wilcoxon Rank Sum test. (e, f) Erap1 (e) and Tap1 (f) expression in p53-WT (blue) and mutant (red) human AML cell lines. (g, h) Erap1 (g) and Tap1 (h) expression in p53-WT (blue) and mutant (red) murine and human DIPG cell lines. Erap1 and Tap1 expression were measured by qRT-PCR. Quantification of 3 independent experiments is shown. Error bars represent means ± SD. p-values were determined by two-sided unpaired t-test comparing p53 WT and mutant lines. In (e) * P= 0.0457 ; in (f) NS, not significant, P= 0.7724 ; in (g) **P= 0.007 (murine) and *P= 0.033 (human) ; in (h) **P= 0.0009 (murine) and *P= 0.028 (human). (I, j) Erap2 (i) and Tap2 (j) expression in MP tumors (n = 3), MG tumors (n = 3), MG tumors overexpressing DNp53 (MG+P) (n = 3) were measured by qRT-PCR. Error bars represent means ± SD. p-values were determined by two-sided unpaired t-test. In (j) NS, not significant, P= 0.21 (MG vs MP), P = 0.394 (MG vs MG+P) ; In (j) NS, P= 0.112 (MG vs MP) and P= 0.149 (MG vs MG+P).
Extended Data Fig. 5 Perturbation of Tap1 alters expression of cell surface MHC-I.
a–d, MG tumor cells were transduced with control shRNA (shCtl) or shRNAs targeting Tap1 (shTap1#1, shTap1#2). Knockdown efficiency was determined by western blotting (a). Western blots are cropped at the molecular weights for Tap1 (68 kDa) and β-actin (42 kDa); original blots are available in Source Data. MHC-I expression was determined by FACS in control cells (shCtl, black) and Tap1 knockdown cells (shTap1, red) (b). Quantification of the mean fluorescence intensity (MFI) values for three independent tumors is shown below each histogram; data points represent MFIs for individual tumor samples. p-values, determined by two-sided unpaired t-test, are indicated on bar graph. c-d, Tap1 knockdown cells were transplanted into aB6 mice. Bioluminescence imaging of representative mice (c) and survival curves (n = 6) (d) are shown. p-values for the difference in survival between shTap1 and shCtl were determined using the two-sided log-rank (Mantel-Cox) test. *P = 0.0198 (shTap1#1) and *P= 0.0246 (shTap1#2). (e, f) MP tumor cells were transduced with empty vector (vect) or vectors encoding Erap1 or Tap1. Analysis of MHC-I expression by FACS in control cells (vect, black), Erap1-overexpressing cells (e) or Tap1-overexpressing cells (f) (blue histograms in e and f). Quantification of the MFI for three independent experiments is shown below the histogram; data points represent MFIs for individual tumor samples. The p-values were determined by two-sided unpaired t-test and are indicated on each bar graph.
Extended Data Fig. 6 TNF increases MHC-I expression in human medulloblastoma and DIPG.
MG (a) and MP (b) tumor cells were treated in vitro for 48h with interleukin-12 (IL-12, 1 μg mL-1). MHC-I expression in untreated cells (Ctl, black histograms) and cells treated with IL-12 (pink) were analyzed by FACS. (c–h) Fresh patient samples (c,d) and PDXs (e-h) from p53-mutant MB patients were treated in vitro for 48h with TNF (50 pg mL-1). HLA-I expression in untreated cells (Ctl, black histograms) and cells treated with TNF (green histograms) were analyzed by FACS. In (c) and (d), no error bars are shown because no replicates were possible from primary patient samples. (i, j) cells from p53-mutant murine (i) and human (j) DIPG cell lines were treated with TNF (50 pg mL-1) for 48h. Murine MHC-I and human HLA-I expression was analyzed by FACS in untreated cells (Ctl, black histograms) and cells treated with TNF (green histograms). k-l, MP tumor cells generated from p53 knockout (p53 KO) mice were treated for 48h with TNF (50pg mL-1) or LtβRag (1.6ug mL-1). MHC-I expression was analyzed by FACS in untreated cells (Ctl, black histograms) and cells treated with TNF (green histograms) or LtβRag (blue histograms). For each FACS analysis (except for fresh MB patient samples in c and d), quantification of the mean fluorescence intensity (MFI) for three independent experiments is shown below each histogram; data points represent MFIs for individual tumor samples. p-values were determined by two-sided unpaired t-test. p-values are indicated on each bar graphs.
Extended Data Fig. 7 TNFR2 and LTβR are necessary for induction of MHC-I by TNF and LTβRag.
a–c, TNFR1 (a, green), TNFR2 (b, magenta) and LTβR (c, blue) expression on MP tumor cells were analyzed by FACS. Isotype control antibody is shown in gray. d–f, MP tumor cells generated from TNFR1 knockout mice (TNFR1 KO, d), TNFR1/2 double knockout mice (TNFR1/2 KO, e) or LTβR knockout mice (LTβR KO, f) were treated for 48h with TNF (50 pg mL-1). MHC-I expression in untreated cells (Ctl, black histograms) and cells treated with TNF (green histograms) were analyzed by FACS. g–i, MP tumor cells generated from TNFR1 knockout mice (TNFR1 KO, g), TNFR2 knockout mice (TNFR2 KO, h) or LTβR knockout mice (LTβR KO, i) were treated for 48h with LTβRag (1.6 μg mL-1). MHC-I expression in untreated cells (Ctl, black histograms) and cells treated with LTβRag (blue histograms) were analyzed by FACS. For each FACS analysis, quantification of the mean fluorescence intensity (MFI) for three independent tumor samples is shown below each histogram; data points represent MFIs for individual tumor samples. p-values were determined by two-sided unpaired t-test, and are indicated on the corresponding graphs. j–l, MP tumor cells were transduced with control shRNA (shCtl) or shRNAs targeting RelA (shRelA#832, shRelA#833), p50 (shp50#484, shp50#485) or RelB (shRelB#494, shRelB#495). Knockdown efficiency of RelA (j), p50 (k) and RelB (l) was determined by western blot. Western blots are cropped at the molecular weights for RelA (65 kDa), p50 (50 kDa), RelB (68 kDa) and GAPDH (37 kDa); original blots are available in Source Data.
Extended Data Fig. 8 TNF increases sensitivity to immune checkpoint inhibitors in a dose-dependent manner.
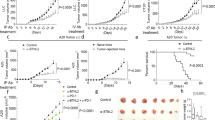
a, TNF toxicity was evaluated in aB6 mice after daily dosing with TNF at 0.5 µg kg-1 (red line), 1 µg kg-1, 10 µg kg-1 and 20 µg kg-1 (n = 3 per dose of TNF). Body weight was measured every 5 days. b,c, MP tumor cells were transplanted into aB6 mice, and after 10 days, animals were treated with vehicle (Ctl) or TNF (0.5 µg kg-1) for 9 days, and tumor tissue was harvested and dissociated. Percentages of isolated cells representing tumor cells (GFP+) and CD3+ T-cells (anti-CD3-APC) were determined by FACS analysis. (b) Representative dot plots (isotype control antibody, black dots; anti-CD3 antibody, green dots) for control mice (left panel) and TNF-treated mice (right panel) are shown, with percentages of cells in each quadrant indicated on the plot. c, Bar graph indicating the average percentage of CD3+ T-cells in tumors from control mice (n = 3, black) and TNF-treated mice (n = 3, green). p-value was determined by two-sided unpaired t-test. ** P = 0.0051). (d–k) MP tumor cells (d–g), MHC-I knock-out (KO) MP tumor cells (h, i) or DNp53-expressing Ptch-KO j,k, tumor cells were transplanted into aB6 mice. 15 days later, mice were treated with vehicle (Ctl, black), anti-PD-1 (10 mg kg-1, blue), TNF (0.1–1 µg kg-1, as indicated in panels d, f, h and j, green) or anti-PD-1 and TNF (orange). For combinations of TNF and anti-PD-1, each cycle of treatment consisted of a dose of TNF on day 1, a dose of anti-PD-1 on day 2, and no treatment on day 3. Mice received up to 20 cycles of treatment. d,f,h,j, Bioluminescence imaging of representative mice. e,g,i,k, Survival curves (in (e) : n = 10 ; in (g) : n = 25 ; in (i) : n = 16; in (k): n = 10 per group). p-values for the difference in survival between Ctl mice and treated mice were determined by a two-sided log-rank (Mantel-Cox) test. (e) *P = 0.019 (anti-PD-1), **P= 0.0007 (TNF), ***P = 9.3 × 10-6 (TNF + anti-PD-1); (g) *** P= 8.1 × 10-6 (anti-PD-1), P= 2.6 × 10-10 (TNF), P= 5.3 × 10-12 (TNF + anti-PD-1); (i) NS, Not Significant P= 0.29 (anti-PD-1), P= 0.151 (TNF), P= 0.094 (TNF + anti-PD-1) ; (k) *** P= 8.1 × 10-5 (anti-PD-1), P = 3.5 × 10-6 (TNF), P= 3.5 × 10-6 (TNF + anti-PD-1).
Extended Data Fig. 9 Representative gating strategy used for analysis of HLA A/B/C in human cells or MHC-I expression in murine cells.
a–d, Gating strategy for unlabeled human tumor cells (data for RCMB18 are shown). (a) Dot plot representing FSC-A and SSC-A profile of the entire cell population, with box indicating intact cells (as opposed to debris and clumps) selected for analysis. (b) Dot plot showing the FSC-H and FSC-A profile of the population selected in (a), with a box indicating the single cell (as opposed to doublet) population selected for analysis. (c) Dot plot showing 7-AAD staining vs. FSC-A of the population in (b), with a box indicating the viable (7-AAD-negative) cell population selected for analysis. (d) Analysis of HLA-I staining of the cell population in (c). (e–i) Gating strategy for GFP-labeled murine (MG) tumor cells. (e) Dot plot representing FSC-A and SSC-A profile of the entire cell population, with box indicating intact cells (as opposed to debris and clumps) selected for analysis. (f) Dot plot showing the FSC-H and FSC-A profile of the population selected in (e), with a box indicating the single cell (as opposed to doublet) population selected for analysis. (g) Dot plot showing 7-AAD staining vs. FSC-A of the population in (f), with a box indicating the viable (7-AAD-negative) cell population selected for analysis. (h) Dot plot showing GFP expression vs. FSC-A in the population selected in (g), with a box indicating the tumor cells (GFP+) selected for analysis. (i) Analysis of MHC-I staining of the population in (h).
Supplementary information
Supplemental Information
Supplementary Tables 1–4.
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 2
Uncropped Western Blots
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 3
Uncropped Western Blots
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 4
Uncropped Western Blots
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 5
Uncropped Western Blots
Source Data Fig. 6
Statistical Source Data
Source Data Fig. 6
Uncropped Western Blots
Source Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 2
Uncropped Western Blots
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 5
Uncropped Western Blots
Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 7
Uncropped Western Blots
Source Data Extended Data Fig. 8
Statistical Source Data
About this article
Cite this article
Garancher, A., Suzuki, H., Haricharan, S. et al. RETRACTED ARTICLE: Tumor necrosis factor overcomes immune evasion in p53-mutant medulloblastoma. Nat Neurosci 23, 842–853 (2020). https://doi.org/10.1038/s41593-020-0628-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-0628-4
This article is cited by
-
Heterogeneity and tumoral origin of medulloblastoma in the single-cell era
Oncogene (2024)
-
Distinct immune escape and microenvironment between RG-like and pri-OPC-like glioma revealed by single-cell RNA-seq analysis
Frontiers of Medicine (2023)
-
FABP4 activates the JAK2/STAT2 pathway via Rap1a in the homocysteine-induced macrophage inflammatory response in ApoE mice atherosclerosis
Laboratory Investigation (2022)
-
Mutant p53 in cancer: from molecular mechanism to therapeutic modulation
Cell Death & Disease (2022)
-
The proprotein convertase furin in cancer: more than an oncogene
Oncogene (2022)