Abstract
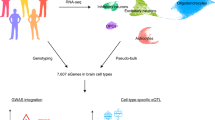
Specific cell populations may have unique contributions to schizophrenia but may be missed in studies of homogenate tissue. Here laser capture microdissection followed by RNA sequencing (LCM-seq) was used to transcriptomically profile the granule cell layer of the dentate gyrus (DG-GCL) in human hippocampus and contrast these data to those obtained from bulk hippocampal homogenate. We identified widespread cell-type-enriched aging and genetic effects in the DG-GCL that were either absent or directionally discordant in bulk hippocampus data. Of the ~9 million expression quantitative trait loci identified in the DG-GCL, 15% were not detected in bulk hippocampus, including 15 schizophrenia risk variants. We created transcriptome-wide association study genetic weights from the DG-GCL, which identified many schizophrenia-associated genetic signals not found in transcriptome-wide association studies from bulk hippocampus, including GRM3 and CACNA1C. These results highlight the improved biological resolution provided by targeted sampling strategies like LCM and complement homogenate and single-nucleus approaches in human brain.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw sequencing reads are available through SRA accession code SRP241159 and BioProject accession code PRJNA600414. Processed data are available at our website: http://research.libd.org/dg_hippo_paper/data.html.
Code availability
Code is available at https://github.com/lieberinstitute/dg_hippo_paper, archived via Zenodo at https://doi.org/10.5281/zenodo.3605718.
References
BrainSeq Consortium. BrainSeq: neurogenomics to drive novel target discovery for neuropsychiatric disorders. Neuron 88, 1078–1083 (2015).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
PsychENCODE Consortiumet al. The PsychENCODE project. Nat. Neurosci. 18, 1707–1712 (2015).
GTEx Consortium et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Jaffe, A. E. et al. Developmental and genetic regulation of the human cortex transcriptome illuminate schizophrenia pathogenesis. Nat. Neurosci. 21, 1117–1125 (2018).
Collado-Torres, L. et al. Regional heterogeneity in gene expression, regulation, and coherence in the frontal cortex and hippocampus across development and schizophrenia. Neuron 103, 203–216 (2019).
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Lake, B. B. et al. Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590 (2016).
Zhu, Y. et al. Spatiotemporal transcriptomic divergence across human and macaque brain development. Science 362, eaat8077 (2018).
Li, M. et al. Integrative functional genomic analysis of human brain development and neuropsychiatric risks. Science 362, eaat7615 (2018).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Bakken, T. E. et al. A comprehensive transcriptional map of primate brain development. Nature 535, 367–375 (2016).
Miller, J. A. et al. Transcriptional landscape of the prenatal human brain. Nature 508, 199–206 (2014).
Arion, D. et al. Transcriptome alterations in prefrontal pyramidal cells distinguish schizophrenia from bipolar and major depressive disorders. Biol. Psychiatry 82, 594–600 (2017).
Kempermann, G., Song, H. & Gage, F. H. Neurogenesis in the adult hippocampus. Cold Spring Harb. Perspect. Biol. 7, a018812 (2015).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Leutgeb, J. K., Leutgeb, S., Moser, M.-B. & Moser, E. I. Pattern separation in the dentate gyrus and CA3 of the hippocampus. Science 315, 961–966 (2007).
Neunuebel, J. P. & Knierim, J. J. CA3 retrieves coherent representations from degraded input: direct evidence for CA3 pattern completion and dentate gyrus pattern separation. Neuron 81, 416–427 (2014).
Hagihara, H., Takao, K., Walton, N. M., Matsumoto, M. & Miyakawa, T. Immature dentate gyrus: an endophenotype of neuropsychiatric disorders. Neural Plast. 2013, 318596 (2013).
Mertens, J. et al. Differential responses to lithium in hyperexcitable neurons from patients with bipolar disorder. Nature 527, 95–99 (2015).
Nakahara, S., Matsumoto, M. & van Erp, T. G. M. Hippocampal subregion abnormalities in schizophrenia: a systematic review of structural and physiological imaging studies. Neuropsychopharmacol. Rep. 38, 156–166 (2018).
Elvsåshagen, T. et al. Dentate gyrus–cornu ammonis (CA) 4 volume is decreased and associated with depressive episodes and lipid peroxidation in bipolar II disorder: longitudinal and cross-sectional analyses. Bipolar Disord 18, 657–668 (2016).
Hibar, D. P. et al. Novel genetic loci associated with hippocampal volume. Nat. Commun. 8, 13624 (2017).
Rasetti, R. et al. Altered hippocampal–parahippocampal function during stimulus encoding: a potential indicator of genetic liability for schizophrenia. JAMA Psychiatry 71, 236–247 (2014).
Weinberger, D. R., Berman, K. F., Suddath, R. & Torrey, E. F. Evidence of dysfunction of a prefrontal–limbic network in schizophrenia: a magnetic resonance imaging and regional cerebral blood flow study of discordant monozygotic twins. Am. J. Psychiatry 149, 890–897 (1992).
Pardiñas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–252 (2016).
Gandal, M. J. et al. Shared molecular neuropathology across major psychiatric disorders parallels polygenic overlap. Science 359, 693–697 (2018).
Eisch, A. J. & Petrik, D. Depression and hippocampal neurogenesis: a road to remission? Science 338, 72–75 (2012).
Overall, R. W., Paszkowski-Rogacz, M. & Kempermann, G. The mammalian adult neurogenesis gene ontology (MANGO) provides a structural framework for published information on genes regulating adult hippocampal neurogenesis. PLoS One 7, e48527 (2012).
Pollen, A. A. et al. Molecular identity of human outer radial glia during cortical development. Cell 163, 55–67 (2015).
Shapiro, E., Biezuner, T. & Linnarsson, S. Single-cell sequencing-based technologies will revolutionize whole-organism science. Nat. Rev. Genet. 14, 618–630 (2013).
Darmanis, S. et al. A survey of human brain transcriptome diversity at the single cell level. Proc. Natl Acad. Sci. USA 112, 7285–7290 (2015).
Xu, X. et al. Species and cell-type properties of classically defined human and rodent neurons and glia. eLife 7, e37551 (2018).
Gandal, M. J. et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science 362, eaat8127 (2018).
Sekar, A. et al. Schizophrenia risk from complex variation of complement component 4. Nature 530, 177–183 (2016).
Li, M. et al. A human-specific AS3MT isoform and BORCS7 are molecular risk factors in the 10q24.32 schizophrenia-associated locus. Nat. Med. 22, 649–656 (2016).
Ma, L. et al. Schizophrenia risk variants influence multiple classes of transcripts of sorting nexin 19 (SNX19). Mol. Psychiatry https://doi.org/10.1038/s41380-018-0293-0 (2019).
Wang, D. et al. Comprehensive functional genomic resource and integrative model for the human brain. Science 362, eaat8464 (2018).
Dong, X. et al. Enhancers active in dopamine neurons are a primary link between genetic variation and neuropsychiatric disease. Nat. Neurosci. 21, 1482–1492 (2018).
Pietersen, C. Y. et al. Molecular profiles of parvalbumin-immunoreactive neurons in the superior temporal cortex in schizophrenia. J. Neurogenet. 28, 70–85 (2014).
Datta, D., Arion, D., Corradi, J. P. & Lewis, D. A. Altered expression of CDC42 signaling pathway components in cortical layer 3 pyramidal cells in schizophrenia. Biol. Psychiatry 78, 775–785 (2015).
Pietersen, C. Y. et al. Molecular profiles of pyramidal neurons in the superior temporal cortex in schizophrenia. J. Neurogenet. 28, 53–69 (2014).
Egan, M. F. et al. Variation in GRM3 affects cognition, prefrontal glutamate, and risk for schizophrenia. Proc. Natl Acad. Sci. USA 101, 12604–12609 (2004).
Green, E. K. et al. The bipolar disorder risk allele at CACNA1C also confers risk of recurrent major depression and of schizophrenia. Mol. Psychiatry 15, 1016–1022 (2010).
Wagner, M. J., Kim, T. H., Savall, J., Schnitzer, M. J. & Luo, L. Cerebellar granule cells encode the expectation of reward. Nature 544, 96–100 (2017).
Huckins, L. M. et al. Gene expression imputation across multiple brain regions provides insights into schizophrenia risk. Nat. Genet. 51, 659–674 (2019).
Bigos, K. L. et al. Genetic variation in CACNA1C affects brain circuitries related to mental illness. Arch. Gen. Psychiatry 67, 939–945 (2010).
Eckart, N. et al. Functional characterization of schizophrenia-associated variation in CACNA1C. PLoS One 11, e0157086 (2016).
Deep-Soboslay, A. et al. Reliability of psychiatric diagnosis in postmortem research. Biol. Psychiatry 57, 96–101 (2005).
Lipska, B. K. et al. Critical factors in gene expression in postmortem human brain: focus on studies in schizophrenia. Biol. Psychiatry 60, 650–658 (2006).
Ziegenhain, C. et al. Comparative analysis of single-cell RNA sequencing methods. Mol. Cell 65, 631–643 (2017).
Babraham Bioinformatics. FastQC https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Feng, Y.-Y., et al. RegTools: integrated analysis of genomic and transcriptomic data for discovery of splicing variants in cancer. Preprint at bioRxiv https://doi.org/10.1101/436634 (2018).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Wang, L., Wang, S. & Li, W. RSeQC: quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185 (2012).
Kent, W. J., Zweig, A. S., Barber, G., Hinrichs, A. S. & Karolchik, D. BigWig and BigBed: enabling browsing of large distributed datasets. Bioinformatics 26, 2204–2207 (2010).
Delaneau, O., Coulonges, C. & Zagury, J.-F. Shape-IT: new rapid and accurate algorithm for haplotype inference. BMC Bioinformatics 9, 540 (2008).
Howie, B., Marchini, J. & Stephens, M. Genotype imputation with thousands of genomes. G3 1, 457–470 (2011).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Hu, P. et al. Dissecting cell-type composition and activity-dependent transcriptional state in mammalian brains by massively parallel single-nucleus RNA-seq. Mol. Cell 68, 1006–1015 (2017).
Habib, N. et al. Div-Seq: single-nucleus RNA-Seq reveals dynamics of rare adult newborn neurons. Science 353, 925–928 (2016).
Lacar, B. et al. Nuclear RNA-seq of single neurons reveals molecular signatures of activation. Nat. Commun. 7, 11022 (2016).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Houseman, E. A. et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinformatics 13, 86 (2012).
Burke, E. E. et al. Dissecting transcriptomic signatures of neuronal differentiation and maturation using iPSCs. Nat. Commun. 11, 462 (2019).
Jaffe, A. E. et al. qSVA framework for RNA quality correction in differential expression analysis. Proc. Natl Acad. Sci. USA 114, 7130–7135 (2017).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Shabalin, A. A. Matrix eQTL: ultra fast eQTL analysis via large matrix operations. Bioinformatics 28, 1353–1358 (2012).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Edlund, C. K., Conti, D. V. & Van Den Berg, D. J. raggr http://raggr.usc.edu/ (USC, 2017).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Acknowledgements
We would like to express our gratitude to our colleagues whose tireless efforts have led to the donation of postmortem tissue to advance these studies: the Office of the Chief Medical Examiner of the District of Columbia, the Office of the Chief Medical Examiner for Northern Virginia, Fairfax, Virginia, and the Office of the Chief Medical Examiner of the State of Maryland, Baltimore, Maryland. We also would like to acknowledge the contributions of L.B. Bigelow for his diagnostic expertise. Finally, we are indebted to the generosity of the families of the decedents, who donated the brain tissue used in these studies.
Author information
Authors and Affiliations
Contributions
A.E.J. performed data processing and analysis and wrote the manuscript. D.J.H., R.T., K.T., M.N.T., K.R.M., K.M., J.H.S., A.D.S. and J.E.K. performed data collection and generation. T.S., E.E.B. and L.C.T. performed data analysis. L.B. and J.U. performed data generation and analysis. D.R.W. performed data interpretation and contributed to the writing of the manuscript. M.M. and T.M.H. supervised data collection and generation and contributed to writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.J.H., T.S., K.T. and M.M. are employees and stockholders of Astellas Pharma. No other authors have competing interests.
Additional information
Peer review information Nature Neuroscience thanks Nenad Sestan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 LCM of granule cell layer from human postmortem dentate gyrus.
a, Cresyl violet stain before and (B) after -LCM. The difference indicates the captured granule cell layer.
Extended Data Fig. 2 Quality control metrics across the DG-GCL and bulk hippocampus.
a, mitochondrial chromosome mapping rate, b, overall mapping rate, c, exonic mapping (also known as gene assignment rate) and d, RNA Integrity number (RIN) compared between Hippocampus and DG-GCL. Colors indicate two different RiboZero library preparation kits: RiboZero Gold and RiboZero HMR (Human-Mouse-Rat).
Extended Data Fig. 3 Principal component analyses (PCA) of various subsets of data.
a, PCA on 224 subjects with data on both HIPPO (red) and DG-GCL (blue). Grey lines connect data from the same donors. b, PCA on 596 subjects used in age, eQTL and diagnosis analyses. c, PCA on 263 DG-GCL samples (blue circles) with HIPPO samples (red squares) projected into component space. d, PCA on the 333 HIPPO samples (red circles) with 263 DG-GCL samples (blue squares) projected into this component space.
Extended Data Fig. 4 Transcript-level overlap of eQTL signal.
Venn diagram displays how eFeatures are mapped to Gencode-annotated transcripts.
Extended Data Fig. 5 Additional DG-GCL eQTL characterizations.
eQTL directional consistency collections between eQTL statistics calculated in HIPPO and DG-GCL, stratified by a, feature type, and then categories related to the b, number of eSNPs and c, statistical significance. d, Local cis heritability differences comparing DG-GCL and DLPFC (which complements Fig. 3b, c).
Extended Data Fig. 6 DG-GCL eQTLs among schizophrenia risk SNPs.
a, Feature-level support for PGC-associated index SNPs in DG-GCL dataset and (B) the number of significant index SNPs as eQTLs (at FDR < 5%) in three brain datasets.
Extended Data Fig. 7 Cross-dataset TWAS analyses.
a, TWAS Z scores across three datasets are highly correlated. Correlations were highest between DLPFC and HIPPO (ρ = 0.64), followed by DG-GCL and HIPPO (ρ = 0.53) then DG-GCL and DLPFC (ρ = 0.50). Overlap of TWAS gene-level associations across three datasets at b, FDR < 5% significance and c, Bonferroni < 10% significance.
Extended Data Fig. 8 Case-control differential expression analyses.
a, Comparisons of log2 fold change effect sizes across diagnoses in the DG-GCL dataset. Comparison of the schizophrenia b, log2 fold changes and c, T-statistics for DG-GCL versus bulk Hippocampus. Color indicates FDR < 0.05 significance indicated in the legend. d, TWAS versus differential expression for schizophrenia effects among 12256 expressed genes with positive heritability (N = 12256) shows no association.
Extended Data Fig. 9 tSNE plot of hippocampus snRNA-seq data.
Shown is a tSNE plot of snRNA-seq data from 4,127 nuclei from bulk hippocampus, colored by data-driven cell types.
Supplementary information
Supplementary Tables
Supplementary Tables 1–10.
Rights and permissions
About this article
Cite this article
Jaffe, A.E., Hoeppner, D.J., Saito, T. et al. Profiling gene expression in the human dentate gyrus granule cell layer reveals insights into schizophrenia and its genetic risk. Nat Neurosci 23, 510–519 (2020). https://doi.org/10.1038/s41593-020-0604-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-0604-z
This article is cited by
-
Central neurocytoma exhibits radial glial cell signatures with FGFR3 hypomethylation and overexpression
Experimental & Molecular Medicine (2024)
-
Prioritization of potential causative genes for schizophrenia in placenta
Nature Communications (2023)
-
Isoform-level transcriptome-wide association uncovers genetic risk mechanisms for neuropsychiatric disorders in the human brain
Nature Genetics (2023)
-
Single-cell and spatial transcriptomics: deciphering brain complexity in health and disease
Nature Reviews Neurology (2023)
-
Integrating genetics and transcriptomics to study major depressive disorder: a conceptual framework, bioinformatic approaches, and recent findings
Translational Psychiatry (2023)