Abstract
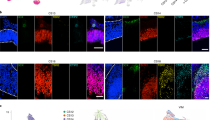
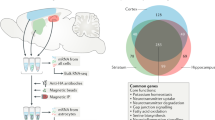
Although the cerebral cortex is organized into six excitatory neuronal layers, it is unclear whether glial cells show distinct layering. In the present study, we developed a high-content pipeline, the large-area spatial transcriptomic (LaST) map, which can quantify single-cell gene expression in situ. Screening 46 candidate genes for astrocyte diversity across the mouse cortex, we identified superficial, mid and deep astrocyte identities in gradient layer patterns that were distinct from those of neurons. Astrocyte layer features, established in the early postnatal cortex, mostly persisted in adult mouse and human cortex. Single-cell RNA sequencing and spatial reconstruction analysis further confirmed the presence of astrocyte layers in the adult cortex. Satb2 and Reeler mutations that shifted neuronal post-mitotic development were sufficient to alter glial layering, indicating an instructive role for neuronal cues. Finally, astrocyte layer patterns diverged between mouse cortical regions. These findings indicate that excitatory neurons and astrocytes are organized into distinct lineage-associated laminae.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw bulk RNA-seq data are available at the Gene Expression Omnibus under the accession code GSE140822. The scRNA-seq data will be made available under https://holt-sc.glialab.org/sc. Other data are available as Supplementary Materials or from the corresponding author upon request.
Code availability
The code for spatial reconstruction of single-cell astrocyte RNA-seq can be found at https://github.com/vitkl/cortical_astrocyte_mapping. The SlideSegmenter code is available at https://bitbucket.org/alexmatlab/slidesegmenter/src/master. The Harmony image analysis scripts are provided as Supplementary Materials. Other code is available upon request.
References
Molyneaux, B. J., Arlotta, P., Menezes, J. R. L. & Macklis, J. D. Neuronal subtype specification in the cerebral cortex. Nat. Rev. Neurosci. 8, 427–437 (2007).
Freeman, M. R. & Rowitch, D. H. Evolving concepts of gliogenesis: a look way back and ahead to the next 25 years. Neuron 80, 613–623 (2013).
Doyle, J. P. et al. Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell 135, 749–762 (2008).
Chai, H. et al. Neural circuit-specialized astrocytes: transcriptomic, proteomic, morphological, and functional evidence. Neuron 548, 396–397 (2017).
John Lin, C.-C. et al. Identification of diverse astrocyte populations and their malignant analogs. Nat. Neurosci. 20, 396–405 (2017).
Boisvert, M. M., Erikson, G. A., Shokhirev, M. N. & Allen, N. J. The aging astrocyte transcriptome from multiple regions of the mouse brain. Cell Rep. 22, 269–285 (2018).
Lanjakornsiripan, D. et al. Layer-specific morphological and molecular differences in neocortical astrocytes and their dependence on neuronal layers. Nat. Commun. 9, 1623 (2018).
Molofsky, A. V. et al. Astrocyte-encoded positional cues maintain sensorimotor circuit integrity. Nature 509, 189–194 (2014).
Lein, E., Borm, L. E. & Linnarsson, S. The promise of spatial transcriptomics for neuroscience in the era of molecular cell typing. Science 358, 64–69 (2017).
Moffitt, J. R. et al. High-performance multiplexed fluorescence in situ hybridization in culture and tissue with matrix imprinting and clearing. Proc. Natl Acad. Sci. USA 113, 14456–14461 (2016).
Shah, S., Lubeck, E., Zhou, W. & Cai, L. In situ transcription profiling of single cells reveals spatial organization of cells in the mouse hippocampus. Neuron 92, 342–357 (2016).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Battich, N., Stoeger, T. & Pelkmans, L. Image-based transcriptomics in thousands of single human cells at single-molecule resolution. Nat. Methods 10, 1127–1133 (2013).
Nikouei, K., Muñoz-Manchado, A. B. & Hjerling-Leffler, J. BCL11B/CTIP2 is highly expressed in GABAergic interneurons of the mouse somatosensory cortex. J. Chem. Neuroanat. 71, 1–5 (2016).
Tasic, B. et al. Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016).
Sosunov, A. A. et al. Phenotypic heterogeneity and plasticity of isocortical and hippocampal astrocytes in the human brain. J. Neurosci. 34, 2285–2298 (2014).
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008).
Nakayama, N. et al. A novel chordin-like protein inhibitor for bone morphogenetic proteins expressed preferentially in mesenchymal cell lineages. Dev. Biol. 232, 372–387 (2001).
Vainchtein, I. D. et al. Astrocyte-derived interleukin-33 promotes microglial synapse engulfment and neural circuit development. Science 359, 1269–1273 (2018).
Champliaud, M. F. et al. Gene characterization of sciellin (SCEL) and protein localization in vertebrate epithelia displaying barrier properties. Genomics 70, 264–268 (2000).
Ruzinova, M. B. & Benezra, R. Id proteins in development, cell cycle and cancer. Trends Cell Biol. 13, 410–418 (2003).
Batiuk, M. Y. et al. Molecularly distinct astrocyte subpopulations spatially pattern the adult mouse brain. Nat. Commun. https://doi.org/10.1038/s41467-019-14198-8 (in the press).
Halpern, K. B. et al. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature 542, 352–356 (2017).
Ge, W.-P., Miyawaki, A., Gage, F. H., Jan, Y.-N. & Jan, L. Y. Local generation of glia is a major astrocyte source in postnatal cortex. Nature 484, 376–380 (2012).
Alcamo, E. A. et al. Satb2 regulates callosal projection neuron identity in the developing cerebral cortex. Neuron 57, 364–377 (2008).
McKenna, W. L. et al. Mutual regulation between Satb2 and Fezf2 promotes subcerebral projection neuron identity in the developing cerebral cortex. Proc. Natl Acad. Sci. USA 112, 11702–11707 (2015).
Leone, D. P. et al. Satb2 regulates the differentiation of both callosal and subcerebral projection neurons in the developing cerebral cortex. Cereb. Cortex 25, 3406–3419 (2015).
Hartfuss, E. Reelin signaling directly affects radial glia morphology and biochemical maturation. Development 130, 4597–4609 (2003).
Kwan, K. Y., Sestan, N. & Anton, E. S. Transcriptional co-regulation of neuronal migration and laminar identity in the neocortex. Development 139, 1535–1546 (2012).
Boyle, M. P. et al. Cell-type-specific consequences of reelin deficiency in the mouse neocortex, hippocampus, and amygdala. J. Comp. Neurol 519, 2061–2089 (2011).
Cui, Y. et al. Astroglial Kir4.1 in the lateral habenula drives neuronal bursts in depression. Nature 554, 323–327 (2018).
Kelley, K. W. et al. Kir4.1-dependent astrocyte-fast motor neuron interactions are required for peak strength. Neuron 98, 306–307 (2018).
Blanco-Suarez, E., Liu, T.-F., Kopelevich, A. & Allen, N. J. Astrocyte-secreted chordin-like 1 drives synapse maturation and limits plasticity by increasing synaptic GluA2 AMPA receptors. Neuron 100, 1116–1132.e13 (2018).
Marshel, J. H. et al. Cortical layer–specific critical dynamics triggering perception. Science 365, eaaw5202 (2019).
Karreman, M. A. et al. Fast and precise targeting of single tumor cells in vivo by multimodal correlative microscopy. J. Cell Sci. 129, 444–456 (2016).
Velmeshev, D. et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science 364, 685–689 (2019).
Gong, S. et al. A gene expression atlas of the central nervous system based on bacterial artificial chromosomes. Nature 425, 917–925 (2003).
Gorski, J. A. et al. Cortical excitatory neurons and glia, but not GABAergic neurons, are produced in the Emx1-expressing lineage. J. Neurosci. 22, 6309–6314 (2002).
Wang, F. et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn. 14, 22–29 (2012).
Kerstens, H. M., Poddighe, P. J. & Hanselaar, A. G. A novel in situ hybridization signal amplification method based on the deposition of biotinylated tyramine. J. Histochem. Cytochem. 43, 347–352 (1995).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. BioMed. Cent. 14, R36 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. BioMed. Cent. 15, 550 (2014).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with Census. Nat. Methods 14, 309–315 (2017).
Lun, A. T. L., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res. 5, 2122 (2016).
Acknowledgements
We thank B. Barres and S. Teichmann for helpful discussions and comments. We thank B. Lynch and X.-J. Ma for advice on RNAScope, as well as R. Sawkins and J. Hutt for technical support on imaging. We also thank E. Olson (SUNY Upstate University) for providing Reeler mice and R. Marcucio (UCSF) for providing the Satb2-flox mice. The authors were supported by the Life Sciences Research Fellowship and the Howard Hughes Medical Institute (O.A.B.), the Wellcome Trust (T.B.), National Institute for Health Research (NIHR) Academic Clinical Fellowship and a Wellcome Trust PhD for Clinicians Fellowship (A.M.H.Y.) and Stichting Alzheiemer Onderzoek (A.M.). The present study was supported by the Paul G. Allen Foundation Distinguished Investigator Program (E.M.U. and D.H.R.), the Loulou Foundation, the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation (D.H.R., D.G. and G. C.), BRAIN initiative (grant no. 1U01 MH105991 to D.G.) and National Institutes of Health (grant nos. 1R01 MH109912 to D.G. and P01NS08351 to D.H.R.), NIHR and the European Union Seventh Framework (to P.H.), National Institute of Neurological Disorders and Stroke Informatics Center for Neurogenetics and Neurogenomics (grant no. P30 NS062691 to G.C.), Wellcome Trust core support (M.H. and O.A.B.), European Research Council (grant no. 281961 to M.G.H.), Fonds Wetenschappelijk Onderzoek (grant nos. G066715N and 1523014N to M.G.H.), Stichting Alzheimer Onderzoek (S no. 16025 to M.G.H.) and VIB Institutional Support and Tech Watch funding (to M.G.H.), Howard Hughes Medical Institute and the Wellcome Trust (to D.H.R.).
Author information
Authors and Affiliations
Contributions
O.A.B. and D.H.R. conceived the study. O.A.B. planned the experiments. O.A.B., T.B., S.H. and K.R. performed histology and imaging experiments. O.A.B. performed the image analysis. K.P. and J.S. contributed to the histology and imaging pipeline. D.P. and O.A.B. analyzed neuron gene expression data to identify subtypes. A.B. wrote the SlideSegmenter software. A.M.H.Y., P.H., M.F.P. and E.H. provided human tissue. O.A.B. and L.B.H. performed layer astrocyte purification. O.A.B. and R.K. analyzed RNA-seq data. G.S., D.H.G. and E.M.U. supervised analysis of neuron smFISH and astrocyte RNA-seq data. A.M., M.B. and M.G.H. generated the astrocyte scRNA-seq data. V.K. and M.H. analyzed the scRNA-seq data and performed spatial reconstruction analysis. K.S. and S.C. supported mouse work and genotyping. O.A.B. and D.H.R. wrote the manuscript with feedback from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Neuroscience thanks F. Guillemot, N. Sestan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–22 and Supplementary Tables 6 and 7.
Supplementary Table
Supplementary Tables 1–5 and 8–11.
Rights and permissions
About this article
Cite this article
Bayraktar, O.A., Bartels, T., Holmqvist, S. et al. Astrocyte layers in the mammalian cerebral cortex revealed by a single-cell in situ transcriptomic map. Nat Neurosci 23, 500–509 (2020). https://doi.org/10.1038/s41593-020-0602-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-0602-1
This article is cited by
-
Distinct forebrain regions define a dichotomous astrocytic profile in multiple system atrophy
Acta Neuropathologica Communications (2024)
-
SRT-Server: powering the analysis of spatial transcriptomic data
Genome Medicine (2024)
-
Brain stars take the lead during critical periods of early postnatal brain development: relevance of astrocytes in health and mental disorders
Molecular Psychiatry (2024)
-
Morphological diversification and functional maturation of human astrocytes in glia-enriched cortical organoid transplanted in mouse brain
Nature Biotechnology (2024)
-
Multicore fiber optic imaging reveals that astrocyte calcium activity in the mouse cerebral cortex is modulated by internal motivational state
Nature Communications (2024)