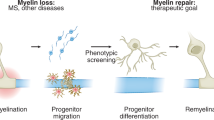
Abstract
The repair of inflamed, demyelinated lesions as in multiple sclerosis (MS) necessitates the clearance of cholesterol-rich myelin debris by microglia/macrophages and the switch from a pro-inflammatory to an anti-inflammatory lesion environment. Subsequently, oligodendrocytes increase cholesterol levels as a prerequisite for synthesizing new myelin membranes. We hypothesized that lesion resolution is regulated by the fate of cholesterol from damaged myelin and oligodendroglial sterol synthesis. By integrating gene expression profiling, genetics and comprehensive phenotyping, we found that, paradoxically, sterol synthesis in myelin-phagocytosing microglia/macrophages determines the repair of acutely demyelinated lesions. Rather than producing cholesterol, microglia/macrophages synthesized desmosterol, the immediate cholesterol precursor. Desmosterol activated liver X receptor (LXR) signaling to resolve inflammation, creating a permissive environment for oligodendrocyte differentiation. Moreover, LXR target gene products facilitated the efflux of lipid and cholesterol from lipid-laden microglia/macrophages to support remyelination by oligodendrocytes. Consequently, pharmacological stimulation of sterol synthesis boosted the repair of demyelinated lesions, suggesting novel therapeutic strategies for myelin repair in MS.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Human single-cell/nucleus RNA sequencing data were obtained from two previously published datasets, GSE118257 and GSE124335. All the data supporting the conclusions of the current study are presented in the figures (Fig. 4h–j contains visualizations of RNA-seq data and Extended Data Fig. 6a–e contains visualizations of RNA-seq data). If necessary, the data that support the findings of this study are available from the corresponding author upon reasonable request. There are no restrictions on data availability. Source data are provided with this paper.
Code availability
Code for data integration and analysis can be found at https://github.com/TSun-tech/MS_scRNAseq.
References
Dietschy, J. M. Central nervous system: cholesterol turnover, brain development and neurodegeneration. Biol. Chem. 390, 287–293 (2009).
Saher, G. et al. High cholesterol level is essential for myelin membrane growth. Nat. Neurosci. 8, 468–475 (2005).
Courtney, R. & Landreth, G. E. LXR regulation of brain cholesterol: from development to disease. Trends Endocrinol. Metab. 27, 404–414 (2016).
Reich, D. S., Lucchinetti, C. F. & Calabresi, P. A. Multiple sclerosis. N. Engl. J. Med. 378, 169–180 (2018).
Plemel, J. R., Liu, W. Q. & Yong, V. W. Remyelination therapies: a new direction and challenge in multiple sclerosis. Nat. Rev. Drug Discov. 16, 617–634 (2017).
Itoh, N. et al. Cell-specific and region-specific transcriptomics in the multiple sclerosis model: focus on astrocytes. Proc. Natl Acad. Sci. USA 115, E302–E309 (2018).
Voskuhl, R. R. et al. Gene expression in oligodendrocytes during remyelination reveals cholesterol homeostasis as a therapeutic target in multiple sclerosis. Proc. Natl Acad. Sci. USA 116, 10130–10139 (2019).
Jurevics, H. et al. Alterations in metabolism and gene expression in brain regions during cuprizone-induced demyelination and remyelination. J. Neurochem. 82, 126–136 (2002).
Lavrnja, I. et al. Expression profiles of cholesterol metabolism-related genes are altered during development of experimental autoimmune encephalomyelitis in the rat spinal cord. Sci. Rep. 7, 2702 (2017).
Cunha, M. I. et al. Pro-inflammatory activation following demyelination is required for myelin clearance and oligodendrogenesis. J. Exp. Med. 217, e20191390 (2020).
Yeung, M. S. Y. et al. Dynamics of oligodendrocyte generation in multiple sclerosis. Nature 566, 538–542 (2019).
Berghoff, S. A. et al. Dietary cholesterol promotes repair of demyelinated lesions in the adult brain. Nat. Commun. 8, 14241 (2017).
Spann, N. J. et al. Regulated accumulation of desmosterol integrates macrophage lipid metabolism and inflammatory responses. Cell 151, 138–152 (2012).
Meschkat, M. et al. White matter integrity requires continuous myelin synthesis at the inner tongue. Preprint at bioRxiv https://doi.org/10.1101/2020.09.02.279612 (2020).
Cantuti-Castelvetri, L. et al. Defective cholesterol clearance limits remyelination in the aged central nervous system. Science 359, 684–688 (2018).
Kotter, M. R., Li, W. W., Zhao, C. & Franklin, R. J. Myelin impairs CNS remyelination by inhibiting oligodendrocyte precursor cell differentiation. J. Neurosci. 26, 328–332 (2006).
Duewell, P. et al. NLRP3 inflammasomes are required for atherogenesis and activated by cholesterol crystals. Nature 464, 1357–1361 (2010).
Boven, L. A. et al. Myelin-laden macrophages are anti-inflammatory, consistent with foam cells in multiple sclerosis. Brain 129, 517–526 (2006).
Widenmaier, S. B. et al. NRF1 is an ER membrane sensor that is central to cholesterol homeostasis. Cell 171, 1094–1109 (2017).
Jakel, S. et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 566, 543–547 (2019).
Masuda, T. et al. Spatial and temporal heterogeneity of mouse and human microglia at single-cell resolution. Nature 566, 388–392 (2019).
Cardeno, A. et al. Squalene targets pro- and anti-inflammatory mediators and pathways to modulate over-activation of neutrophils, monocytes and macrophages. J. Funct. Foods 14, 779–790 (2015).
Saher, G. et al. Therapy of Pelizaeus–Merzbacher disease in mice by feeding a cholesterol-enriched diet. Nat. Med. 18, 1130–1135 (2012).
Gylling, H. & Miettinen, T. A. Postabsorptive metabolism of dietary squalene. Atherosclerosis 106, 169–178 (1994).
Gudi, V., Gingele, S., Skripuletz, T. & Stangel, M. Glial response during cuprizone-induced de- and remyelination in the CNS: lessons learned. Front. Cell. Neurosci. 8, 73 (2014).
Muse, E. D. et al. Cell-specific discrimination of desmosterol and desmosterol mimetics confers selective regulation of LXR and SREBP in macrophages. Proc. Natl Acad. Sci. USA 115, E4680–E4689 (2018).
Hubler, Z. et al. Accumulation of 8,9-unsaturated sterols drives oligodendrocyte formation and remyelination. Nature 560, 372–376 (2018).
Miron, V. E. et al. M2 microglia and macrophages drive oligodendrocyte differentiation during CNS remyelination. Nat. Neurosci. 16, 1211–1218 (2013).
Bogie, J. F. et al. Myelin-derived lipids modulate macrophage activity by liver X receptor activation. PLoS ONE 7, e44998 (2012).
Lloyd, A. F. et al. Central nervous system regeneration is driven by microglia necroptosis and repopulation. Nat. Neurosci. 22, 1046–1052 (2019).
Olah, M. et al. Identification of a microglia phenotype supportive of remyelination. Glia 60, 306–321 (2012).
Kuhlmann, T. et al. An updated histological classification system for multiple sclerosis lesions. Acta Neuropathol. 133, 13–24 (2017).
Akula, M. K. et al. Protein prenylation restrains innate immunity by inhibiting Rac1 effector interactions. Nat. Commun. 10, 3975 (2019).
Araldi, E. et al. Lanosterol modulates TLR4-mediated innate immune responses in macrophages. Cell Rep. 19, 2743–2755 (2017).
Korner, A. et al. Inhibition of Δ24-dehydrocholesterol reductase activates pro-resolving lipid mediator biosynthesis and inflammation resolution. Proc. Natl Acad. Sci. USA 116, 20623–20634 (2019).
Yang, C. et al. Sterol intermediates from cholesterol biosynthetic pathway as liver X receptor ligands. J. Biol. Chem. 281, 27816–27826 (2006).
Brown, M. S. & Goldstein, J. L. The SREBP pathway: regulation of cholesterol metabolism by proteolysis of a membrane-bound transcription factor. Cell 89, 331–340 (1997).
Liebergall, S. R. et al. Inflammation triggers liver X receptor-dependent lipogenesis. Mol. Cell. Biol. 40, e00364 (2020).
Fellows Maxwell, K. et al. Oxysterols and apolipoproteins in multiple sclerosis: a 5 year follow-up study. J. Lipid Res. 60, 1190–1198 (2019).
Zmyslowski, A. & Szterk, A. Oxysterols as a biomarker in diseases. Clin. Chim. Acta 491, 103–113 (2019).
Safaiyan, S. et al. Age-related myelin degradation burdens the clearance function of microglia during aging. Nat. Neurosci. 19, 995–998 (2016).
Marschallinger, J. et al. Lipid-droplet-accumulating microglia represent a dysfunctional and proinflammatory state in the aging brain. Nat. Neurosci. 23, 194–208 (2020).
Thelen, K. M., Falkai, P., Bayer, T. A. & Lutjohann, D. Cholesterol synthesis rate in human hippocampus declines with aging. Neurosci. Lett. 403, 15–19 (2006).
Hendrickx, D. A. E. et al. Gene expression profiling of multiple sclerosis pathology identifies early patterns of demyelination surrounding chronic active lesions. Front. Immunol. 8, 1810 (2017).
Mailleux, J. et al. Active liver X receptor signaling in phagocytes in multiple sclerosis lesions. Mult. Scler. 24, 279–289 (2018).
Yu, S. et al. Dissociated sterol-based liver X receptor agonists as therapeutics for chronic inflammatory diseases. FASEB J. 30, 2570–2579 (2016).
Katz Sand, I. The role of diet in multiple sclerosis: mechanistic connections and current evidence. Curr. Nutr. Rep. 7, 150–160 (2018).
Sedaghat, F., Jessri, M., Behrooz, M., Mirghotbi, M. & Rashidkhani, B. Mediterranean diet adherence and risk of multiple sclerosis: a case–control study. Asia Pac. J. Clin. Nutr. 25, 377–384 (2016).
Beltrán, G., Buchelo, M. E., Aguilera, M. P., Belaj, A. & Jimenez, A. Squalene in virgin olive oil: screening of variability in olive cultivars. Eur. J. Lipid Sci. Tech. 118, 1250–1253 (2016).
Smith, T. J. Squalene: potential chemopreventive agent. Expert Opin. Investig. Drugs 9, 1841–1848 (2000).
Quinet, E. M. et al. Gene-selective modulation by a synthetic oxysterol ligand of the liver X receptor. J. Lipid Res. 45, 1929–1942 (2004).
Bachmanov, A. A., Reed, D. R., Beauchamp, G. K. & Tordoff, M. G. Food intake, water intake, and drinking spout side preference of 28 mouse strains. Behav. Genet. 32, 435–443 (2002).
Bligh, E. G. & Dyer, W. J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959).
Alizadeh, A. & Karimi-Abdolrezaee, S. Microenvironmental regulation of oligodendrocyte replacement and remyelination in spinal cord injury. J. Physiol. 594, 3539–3552 (2016).
Radzun, H. J. et al. Detection of a monocyte/macrophage differentiation antigen in routinely processed paraffin-embedded tissues by monoclonal antibody Ki-M1P. Lab. Invest. 65, 306–315 (1991).
Lampron, A. et al. Inefficient clearance of myelin debris by microglia impairs remyelinating processes. J. Exp. Med. 212, 481–495 (2015).
Bijland, S. et al. An in vitro model for studying CNS white matter: functional properties and experimental approaches. F1000Res 8, 117 (2019).
Bottenstein, J. E. & Sato, G. H. Growth of a rat neuroblastoma cell line in serum-free supplemented medium. Proc. Natl Acad. Sci. USA 76, 514–517 (1979).
Wang, X. & Roper, M. G. Measurement of DCF fluorescence as a measure of reactive oxygen species in murine islets of Langerhans. Anal. Methods 6, 3019–3024 (2014).
Lucchinetti, C. F. et al. Clinical and radiographic spectrum of pathologically confirmed tumefactive multiple sclerosis. Brain 131, 1759–1775 (2008).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: uniform manifold approximation and projection. J. Open Source Soft. 3, 861 (2018).
Wickham, H. ggplot2—Elegant Graphics for Data Analysis (Springer, 2009).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Acknowledgements
We thank A. Fahrenholz, T. Freerck, B. Sadowski, L. Fernandez Garcia-Agudo and M. Lim for technical support. We thank C. Stiles and J. Alberta (Harvard Medical School, Boston, USA), S. Ghandour (CNRS, Strasbourg, France) and H. J. Radzun (University Medical Centre, Göttingen, Germany) for the generous gifts of antibodies. We thank M. Schwaninger for providing Slco1c1-CreERT2 mice (University of Lübeck, Lübeck, Germany). This work was funded by the Deutsche Forschungsgemeinschaft (SA2014/2-1 to G.S.). G.S. is supported by the Wilhelm-Sander Stiftung and the Alzheimer Forschung Initiative. Myelinating cell culture work was funded by the UK MS Society (grant 38 to J.M.E.). K.A.N. is supported by a European Research Council Advanced Grant. The Adelson Medical Research Foundation supports M.S. and K.A.N. M.W.S. is supported by the BMBF (CMT-NET01GM1511C and CMT-NRG 01GM1605). P.S. is supported by the Studienstiftung des deutschen Volkes. M.P. is supported by the Sobek Foundation, the Ernst-Jung Foundation, the German Research Foundation (SFB992, SFB1160, Reinhart-Koselleck-Grant, TRR167) and the Ministry of Science, Research and Arts, Baden-Wuerttemberg (Sonderlinie ‘Neuroinflammation’) as well as by the BMBF-funded competence network of multiple sclerosis (KKNMS). F.O. is supported by the Deutsche Forschungsgemeinschaft (OD87/1-1, OD87/3-1). F.O. and L.H. are supported by the Deutsche Forschungsgemeinschaft (TRR274/1).
Author information
Authors and Affiliations
Contributions
S.A.B. and G.S. planned and designed the study. S.A.B. and L. Spieth were involved in all experiments. T.S. performed the reanalysis of human scRNA-seq datasets. L. Schlaphoff performed cell culture experiments. L.H. and F.O. performed the flow cytometry experiments with subsequent expression analyses. T.D. was involved in MACS cell isolation. D.E. and M.W.S. performed the HPLC serum analysis. J.N. performed the western blot analysis. T.R. performed the electron microscopy analysis. T.I. and P.S. performed lipid mass spectrometry. I.H. provided human tissue. C.S.N. and F.v.d.M. characterized and isolated human MS tissue lesions. R.S. and M.P. provided RNA-seq datasets. L.C.C. and M.S. were in involved in the lysolecithin experiments. J.W., W.M. and M.M. characterized mouse pathology. J.M.E. performed myelinating co-culture experiments. C.D. and A.O.S. performed light-sheet microscopy. S.A.B., J.M.E., K.A.N. and G.S. wrote and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.A.B. and G.S. are listed as inventors on pending patent claims (PCT/EP2020/084338) filed by MPG covering the application of squalene in demyelinating disorders.
Additional information
Peer review information Nature Neuroscience thanks Vittorio Gallo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Mouse models of demyelination and remyelination.
a, The cuprizone model of demyelination and remyelination exhibits some similarities to pattern III MS lesions25. Although the exact molecular cause of the cuprizone induced pathology remains elusive, copper chelation induced dysfunction of mitochondrial enzymes could selectively lead to apoptosis in oligodendrocytes (Praet et al., 2014). Cuprizone (bis-cyclohexanone oxaldihydrazone) is administered to adult mice as a 0.2% dietary supplement, which leads to a highly reproducible oligodendropathy followed by demyelination of distinct brain regions, predominantly the corpus callosum (CC). The graphical representation of myelination and microgliosis is based on histochemical stainings as shown in lower panels (further details to this model12,25 and Matsushima and Morell, 2001). Feeding mice with cuprizone induces progressive loss of mature oligodendrocytes (CAII) that is already detectable two to three days following cuprizone administration and accompanied by gliosis and blood-brain barrier impairment (Berghoff et al., 2017, Stumpf et al., 2019). Loss of oligodendrocytes induces demyelination (Gallyas) in the corpus callosum starting after 2-3 weeks of cuprizone treatment. Loss of mature oligodendrocytes to less than 10% of control numbers and maximum demyelination of the corpus callosum is achieved after 5 weeks of continuous cuprizone supplementation (5w, acute demyelination). During the 5 weeks, OPCs proliferate (OLIG2) and remain largely undifferentiated. Whether OPCs do not differentiate during this disease period or differentiate, attempt to remyelinate and subsequently die is unclear. Demyelination is accompanied by progressive microgliosis (MAC3, dashed gray line). Following the peak at 5w, microgliosis quickly resolves within the following week despite ongoing cuprizone administration, leading to moderate microgliosis at 6 weeks cuprizone treatment (about 30% of MAC3 levels at 5w). This coincides with a brief remission phase characterized by oligodendrocyte maturation. The corpus callosum remyelinates reaching levels of about 50% myelin content of untreated controls (6w, acute-phase remyelination). In case of ongoing cuprizone treatment, remyelinated myelin sheaths subsequently degenerate. Microglial numbers progressively decrease despite ongoing degeneration leading to residual numbers of microglia at 12 weeks cuprizone. At this stage, the corpus callosum is chronically demyelinated (12w, chronic demyelination). CAII numbers of ~30% of untreated controls might also reflect a second spontaneous but minor remyelination phase. When cuprizone is withdrawn at this point and mice are fed normal chow, remyelination commences and reaches ~20% myelin content of untreated controls after 2 weeks recovery (12 + 2 weeks, chronic-phase remyelination) (Scale 50 µm). b, Quantitative RT-PCR analysis determining the expression of oligodendrocyte and myelin related genes (Car2, Plp1, Olig2), and marker genes for microglia (Aif1) and astrocytes (Gfap) in corpus callosum samples during acute and chronic remyelination. Bars represent the means with individual data points normalized to the demyelination time point 5w cuprizone (n = 4 animals, set to 1, dotted line). Acute-phase (6w, n = 4 animals) and chronic-phase (12 + 2w, n = 4 animals) remyelination is reflected in increased transcripts of genes related to oligodendrocyte maturation and myelination (Car2, Plp1) in corpus callosum samples. The resolving microgliosis between 5w and 6w of cuprizone feeding is reflected in an about 40% drop of Aif1 transcripts. Asterisks mark significant changes, ***p < 0.001 (Student’s t-test, two-sided). c, The lysolecithin model of demyelination and remyelination in the spinal cord15 (Bjelobaba et al., 2018). Focal spinal cord demyelinating lesions are induced by stereotactic injection of 1 µl lysolecithin (1%) into the left and right ventro-lateral funiculus between Th3 and Th4. Demyelination starts within hours and proceeds until about 5-7 days post injection, which is paralleled by microgliosis and OPC proliferation. Then phagocyte activity progressively resolves resulting in low levels of activated microglia at 3 weeks post induction. After clearance of myelin debris, oligodendrocytes differentiate and remyelination starts after about 10 days post induction. Like other toxic models, pathology in the lysolecithin is not mediated by inflammatory immune cell infiltration, although some invasion of T- and B-lymphocytes occur and the blood-brain barrier is disrupted by the experimental procedure (Yazdi et al., 2015) (Scale 100 µm). In contrast to the cuprizone model, repeated demyelination does not result in limited remyelination and reduced number of OPCs (Penderis et al., 2003). d, MOG-EAE (experimental autoimmune encephalomyelitis) causes a monophasic inflammatory demyelinating disease and models predominantly inflammatory aspects of MS pathology (Constantinescu et al., 2011; Ransohoff, 2012). Mice are immunized with the MOG35-55 peptide (amino acid 35-55 of myelin oligodendrocyte protein) that induces expansion of autoreactive T lymphocytes during a preclinical phase. Already during this disease period, the blood-brain barrier is compromised (Paul et al., 2013) and some CNS immune cell entry and cytokine production occur. After 10-13 days post immunization clinical disabilities become evident characterized by ascending paralysis. Histopathology is more pronounced in spinal cord tissue than in brain. Lesions are characterized by marked infiltration of encephalitogenic effector T cells (CD3), as well as recruitment and activation of microglia/macrophages cells (Iba1) and demyelination (MBP). Tissue damage during this acute disease phase of EAE reaches a peak after several days followed by a chronic phase with milder but persistent disability. Remyelination events that are considered to be minor. Scale 50 µm.
Extended Data Fig. 2 Generation and validation of cell specific cholesterol synthesis mutants.
a, Quantitative RT-PCR analysis determining the expression of housekeeping reference genes in corpus callosum or isolated cells from mice (n = 4 animals) treated with cuprizone (A astrocytes, EC endothelial cells, M microglia/macrophages, OL oligodendrocytes). b, Conditional mutants were generated by crossing SQSflox/flox mice to Cre driver lines. Fdft1 was inactivated by tamoxifen injections, and mice were analyzed at time points as indicated (middle panel). Fdft1 (SQS) expression in acutely isolated cells targeted by the conditional mutation shown as mean of fold values (OLcKO n = 4, OPCcKO n = 3, AcKO n = 3, McKO n = 3, ECcKO n = 3) normalized to cells from control mice (OLcKO-Ctrl n = 4, OPCcKO-Ctrl n = 3, AcKO-Ctrl n = 3, McKO-Ctrl n = 4, ECcKO-Ctrl n = 3; set to 1). Asterisks mark significant changes (Student’s t-test, two-sided). c, Body weight (OLcKO n = 15, OPCcKO n = 7, AcKO n = 5, McKO n = 10, ECcKO n = 5; OLcKO-Ctrl n = 11, OPCcKO-Ctrl n = 6, AcKO-Ctrl n = 5, McKO-Ctrl n = 9, ECcKO-Ctrl n = 6) and total serum cholesterol (OLcKO n = 7, OPCcKO n = 7, AcKO n = 5, McKO n = 5, ECcKO n = 8; OLcKO-Ctrl n = 4, OPCcKO-Ctrl n = 6, AcKO-Ctrl n = 5, McKO-Ctrl n = 5, ECcKO-Ctrl n = 5) of analyzed cell-specific cholesterol synthesis mutants in (b) at the age of 22 weeks. Bars represent mean of individual animals. Asterisks mark significant changes (Student’s t-test, two-sided). d, Evaluation of astrocyte-specific cholesterol synthesis mutants (Aldh1l1-CreERT2 driver line, Winchenbach et al., 2016), determining Fdft1 expression in isolated astrocytes, body weight and total serum cholesterol in mutants (isolated astrocytes n = 3 animals; body weight n = 9 animals, cholesterol n = 7 animals and Cre controls (isolated astrocytes n = 4 animals; body weight n = 3 animals, cholesterol n = 8 animals). Bars represent the means with individual data points normalized to controls. Asterisks mark significant changes (Student’s t-test, two-sided). e, Targeted sterol metabolism gene expression profile of isolated cells from the corpus callosum of n = 4 untreated WT animals. The relative expression heat map shows low transcript levels (blue) and high transcript levels (red) for each gene normalized to the mean of all samples. f, Quantification of histological stainings for myelination (Gallyas), mature oligodendrocytes (CAII), oligodendrocyte linage cells (Olig2), microgliosis (MAC3) and astrogliosis (GFAP) in Aldh1l1-CreERT2 astrocyte cholesterol synthesis mutants (n = 4 animals) as in (d) at 6 weeks cuprizone. Bars represent the means with individual data points normalized to controls (n = 4 animals) set to 100). ***p < 0.001, **p < 0.01, *p < 0.05.
Extended Data Fig. 3 Remyelination efficiency in conditional mutants.
a, Expression of cholesterol synthesis genes in isolated astrocytes from EAE lesions. Bars represent the means with individual data points (n = 4 animals) normalized to isolated cells from controls (n = 4 animals, set to 1). Asterisks mark significant changes (Student’s t-test, two-sided). b, (Upper panel) Generation of MBP conditional mutants (MBPcKO) and (Lower panel) expression of cholesterol synthesis genes in isolated astrocyte 12 month post tamoxifen administration. Bars represent the means with individual data points (n = 4 animals) normalized to isolated cells from control (n = 4 animals, set to 1). Asterisks mark significant changes (Student’s t-test, two-sided). c, Tamoxifen protocol to induced recombination during cuprizone paradigm. Tamoxifen was given either before cuprizone application to target SQS in astrocytes, microglia, OPCs and endothelial cells or before remyelination to target oligodendrocytes. d, TDTO reporter expression during remyelination to evaluate cell targeting in cuprizone fed mice by co-labeling with CAII for oligodendrocytes, OLIG2 for oligodendrocyte linage cells, GFAP for astrocytes and Iba1 for microglia (Scale 50 µm). e, Elevated beam testing of cholesterol synthesis mutants during chronic remyelination (OLcKO n = 8, OPCcKO n = 7, AcKO n = 5, McKO n = 5, ECcKO n = 7; OLcKO-Ctrl n = 8, OPCcKO-Ctrl n = 7, AcKO-Ctrl n = 5, McKO-Ctrl n = 5, ECcKO-Ctrl n = 8). Bars represent the means with individual data points of three testing sessions. Asterisks mark significant changes (Student’s t-test, two-sided). ***p < 0.001, **p < 0.01.
Extended Data Fig. 4 Ablation of microglial cholesterol synthesis does not affect demyelination in the cuprizone model.
a, Generation and cuprizone treatment paradigm of phagocyte-specific Abca1/Abcg1 cholesterol export mutants (ABC/McKO). b, Representative images from control (n = 3 animals) and SQS/McKO mice (n = 3 animals) after 5w cuprizone (acute demyelination), showing myelination (ultrastructure, scale 1 µm), oligodendrocytes (CAII), oligodendrocyte lineage cells (OLIG2) and microgliosis (MAC3) (Scale 50 µm) with corresponding quantification on the right. Asterisks mark significant changes (Student’s t-test, two-sided). c, Fdft1 expression in SQS-KO BMDMs cultures that were recombined in vitro (n = 4 cultures). Data normalized to control cells (n = 3 cultures, set to 1). Asterisks mark significant changes (Student’s t-test, two-sided). d, Cholesterol labeling in SQS-KO BMDMs (Ctrl n = 3, SQS-KO n = 4 cultures) cultured in lipid-depleted media by live staining with BCtheta (Scale 10 µm). e, In vitro Fluoromyelin phagocytosis in control and SQS-KO BMDM cultures after a 4 h challenge with myelin (Scale 10 µm). Internalized myelin (fluoromyelin) was quantified by co-labeling for Lamp1-positive endo/lysosomes (Ctrl n = 7 cultures, SQS-KO n = 4 cultures; 600-1050 cells). f, Mean gene expression related to NLRP3 inflammasome and cholesterol efflux (Abca1) in isolated phagocytes from SQS/McKO mice (n = 5 animals) normalized to cuprizone control (n = 3 animals). Asterisks mark significant changes (Student’s t-test, two-sided). g,Abca1 and Abcg1 expression in ABC-KO BMDMs cultures that were recombined in vitro (n = 4 cultures). Data normalized to control cells (n = 4 cultures, set to 1). Asterisks mark significant changes (Student’s t-test, two-sided). h, In vitro survival of BMDM following myelin (10 µg/ml) treatment (n = 17 cultures from control mice; n = 16 from SQS mutants and ABC mutants, n = 12 from APO knockout mice; each with 251-351 cells). About 50% of SQS mutant phagocytes died within 16 h of incubation with myelin, with morphology attributable to pyroptosis (Video S1). Of note, in two lipid export mutants, that is ABCA1/ABCG1 and ApoE-deficient phagocytes, pyroptotic cell death was even more pronounced, whereas control phagocyte numbers did not change. Asterisks mark significant changes (one-way ANOVA with Tukey’s post test). ***p < 0.001, **p < 0.01, *p < 0.05.
Extended Data Fig. 5 EAE pathology in SQS/McKO animals.
a, Representative pictures showing gating strategy to analyze lymphocytes and monocytes by flow cytometry. b, Flow cytometry plots depicting CD4 + T cells (left), CD8 + T cells (middle) and myeloid subpopulations (right) consisting of macrophages (CD11b + /CD45high) and microglia (CD11b + /CD45low). c, Flow cytometric quantification of inflammatory cell abundance at chronic EAE of SQS/McKO (n = 8 animals) and control (n = 8 animals) spinal cord samples. Asterisks mark significant changes (Student’s t-test, two-sided). d, Light sheet microcopy of SQS/McKO (n = 3 animals) and control (n = 3 animals) spinal cord samples stained for Iba1+ phagocytes (red) and autofluorescence imaging. Arrows point to circular areas with low optical density suggestive of focal edema that was only occasionally observed in EAE controls (Scale 100 µm). e, Quantitative RT-PCR analysis determining the expression housekeeping reference genes in tissue and isolated CD11b + cells from cuprizone treated SQS/McKO (n = 5 animals), cuprizone control mice (n = 4 animals), EAE (n = 6 animals) and EAE control mice (n = 5 animals). ***p < 0.001, **p < 0.01, *p < 0.05.
Extended Data Fig. 6 LXR target gene upregulation and desmosterol accumulation in demyelinating mouse models.
a, Quantitative RT-PCR analysis determining expression of Abca1 and Apoe in isolated CD11b + cells from chronic EAE (left panel, n = 4 animals), MBPcKO (middle panel, n = 4 animals) and lysolecithin (right panel, n = 3 animals) lesions. Bars represent the means with individual data points normalized to isolated cells from control (EAE n = 4 animals, MBPcKO-Ctrl n = 4 animals, Lysolecithin Ctrl n = 4 animals, set to 1) (Student’s t-test, two-sided). b, Relative abundance of lanosterol and cholesterol normalized to cell number in isolated CD11b + myeloid cells during acute cuprizone-mediated demyelination (5w cuprizone), measured by lipid mass spectroscopy (Ctrl n = 3 animals, cuprizone n = 4 animals) (Student’s t-test, two-sided). c, Relative expression of Nrf1/Nfe2l1 in isolated microglia isolated from control mice and diseased mice (n = 4 mice per group) after feeding cuprizone for 6 weeks (left), 12 + 2 weeks (middle) and 28d after induction of EAE (right) (Student’s t-test, two-sided). d, Relative abundance of desmosterol in corpus callosum during acute demyelination (5w cuprizone) of SQS/McKO animals (n = 5 animals) and cuprizone controls (n = 5 animals) measured by lipid mass spectrometry. Data normalized to non-disease controls (n = 3 animals, set to 1) (one-way ANOVA with Tukey’s post test). e, Relative abundance of desmosterol in spinal cord tissue during chronic EAE (28 dpi) of SQS/McKO animals (n = 4 animals) and EAE controls (n = 3 animals) normalized to untreated controls (n = 4 animals, set to 1) measured by lipid mass spectroscopy (one-way ANOVA with Tukey’s post test). f, Total ion chromatogram, selected ion mass chromatograms and spectra from representative human tissue sample and commercial standards analyzed by GC-MS. Time axes of chromatograms are broken to show only the elution of sterols. Lower panel shows background subtracted (adjacent to peak) mass spectra of N-Methyl-N (trimethylsilyl) trifluoroacetamide (MSTFA)-derivated metabolites. Asterisks mark significant changes, ***p < 0.001, **p < 0.01, *p < 0.05.
Extended Data Fig. 7 Identity of immune cell cluster in human MS scRNA-seq datasets and characterization of analyzed human MS lesions.
a, Distribution of immune cells from control and MS patients (left) and k-means clustering (right) of cells (resolution 0.1) on UMAP plot of merged external scRNA-seq datasets. b, Frequency distribution of cells from control and MS patients in each cluster. c, Expression of selected marker genes in T cells (CD3) and non-microglia/macrophages (FSP1, Fibroblast-Specific Protein-1) in cluster 4 of sc-RNAseq datasets. d, Expression of selected marker genes in myeloid cells (AIF1, CX3CR1), oligodendrocytes (PLP1) and DHCR24 expressing cells in analyzed immune cells of sc-RNAseq datasets. e, Expression of genes related to cholesterol export in analyzed immune cells of sc-RNAseq datasets. f, Histological characterization of isolated lesions from individual patients. Dotted line indicates lesion rim defined by myelin staining (LFB/PAS).Center activity (square) was determined by presents of phagocytic cells (KiM1P) (Scale 500 µm). g, Tissue water (g/g dry tissue) of control white (n = 3 patients) matter and MS lesion material (n = 4 patients). R: Active lesion rim (n = 5 lesions); AC: Active lesion center (n = 3 lesions); IC: Inactive lesion center (n = 4 lesions). Asterisks mark significant changes (one-way ANOVA with Holm-Sidak post test). **p < 0.01,*p < 0.05. h, Relative abundance of cholesterol normalized to input tissue weight and standard (controls set to 1) of control white (n = 3 patients) matter and MS lesion material (n = 4 patients). R: Active lesion rim (n = 5 lesions); AC: Active lesion center (n = 3 lesions); IC: Inactive lesion center (n = 4 lesions). Asterisks mark significant changes (one-way ANOVA with Holm-Sidak post test). ***p < 0.001, **p < 0.01, *p < 0.05.
Extended Data Fig. 8 Inflammatory response in wild type and SQS-KO BMDMs challenged with LPS.
a, Mean ROS levels in control (left) and SQS-KO BMDMs (right) stimulated with or without LPS and treated with or without squalene (100 µM) (n = 8 cultures per condition). Data normalized to untreated control (set to 1). Asterisks mark significant changes (two-way ANOVA with Holm-Sidak post test). b-f, Relative expression of cholesterol efflux (b, c) and inflammatory (d-f) genes in control and SQS-KO BMDMs treated with myelin+LPS with or without squalene (as depicted in Fig. 5c), normalized to untreated WT cultures(n = 4 cultures) (two-way ANOVA with Holm-Sidak post test). Asterisks mark significant changes, ***p < 0.001, **p < 0.01, *p < 0.05.
Extended Data Fig. 9 Squalene affects serum squalene levels in lysolecithin and EAE models of MS but not in cuprizone-treated mice.
a, Peripheral serum cholesterol profile of control (n = 5 animals) and squalene-treated WT (n = 6 animals) animals 14 days post injection with lysolecithin. Measurement shows total cholesterol (TC), low-density lipoprotein cholesterol (LDL) and high-density lipoprotein cholesterol (HDL). Dotted lines indicate values of non-diseased WT mice (Student’s t-test, two-sided). b, Serum squalene concentration measured by HPLC in control and squalene-treated lysolecithin animals treated as in (a) (n = 3 animals each group) (Student’s t-test, two-sided). c, Histopathological quantification of the lesion size and MAC3 + microglia/macrophages in the lesions from control (n = 9 lesions from 5 animals) and squalene treated animals (n = 11 lesions from 6 animals) at 14 dpi lysolecithin (Scale 100 µm) (Student’s t-test, two-sided). d, Scheme depicting induced remyelination paradigm in the cuprizone model12. Mice were fed cuprizone for 4 weeks after which mice received chow supplemented with vehicle or squalene for one week (dotted line) followed by analysis. The gray line depicts the degree of myelination in case of continued cuprizone feeding. e, Peripheral serum cholesterol profile determining total cholesterol (TC), low-density lipoprotein cholesterol (LDL) and high-density lipoprotein cholesterol (HDL) in control (n = 5 animals) and cuprizone (n = 4 animals) mice treated as in (d). Dotted lines indicate values of non-disease WT mice (Student’s t-test, two-sided). f, Serum squalene concentration measured by HPLC in control and squalene-treated lysolecithin animals treated as in (d) (n = 3 animals each group). Dotted line indicates values of non-disease controls. g, Histological quantification of myelination (Gallyas), mature oligodendrocytes (CAII), oligodendrocyte linage cells (OLIG2), microgliosis (MAC3) and astrogliosis (GFAP) at induced remyelination with squalene supplementation (n = 3 animals) compared to control (n = 7 animals) in mice treated as in (d). h, Peripheral serum cholesterol profile of control (n = 10 animals) and squalene-treated (n = 11 animals) EAE animals during peak of disease (16 dpi). Measurement shows total cholesterol (TC), low-density lipoprotein cholesterol (LDL) and high-density lipoprotein cholesterol (HDL). Dotted lines indicate values of non-disease WT mice. i, Serum squalene concentration of control (n = 2 animals) and squalene-treated (n = 3 animals) EAE animals during peak of disease (16 dpi) measured by HPLC. j, Clinical score ± SEM of control (n = 6) and prophylactically squalene-treated (n = 6) EAE animals analyzed at peak of disease (16 dpi). k, Number of CD4 + and CD8 + T cells, CD11b/CD45high macrophages and CD11b/CD45low microglia per gram spinal cord determined by flow cytometry at peak of EAE (16 dpi) in control (n = 6 animals) and squalene-treated animals (n = 6 animals) (Student’s t-test, two-sided). l, Quantification of blood inflammatory cells by flow cytometry from animals as in (j) during peak of EAE disease (n = 6 animals each group). m, Analysis of inflammatory cells in spinal cord tissue by flow cytometry from control (n = 6 animals) and therapeutically squalene-treated (n = 5 animals) EAE animals (Fig. 6e) (Student’s t-test, two-sided). n, Relative abundance of sterol intermediates in spinal cord of squalene-treated EAE animals (n = 4 animals) normalized to untreated EAE controls (n = 4 animals, set to 1) (Student’s t-test, two-sided). o, Quantitative RT-PCR analysis determining the expression housekeeping reference genes in isolated CD11b + cells from spinal cord from EAE control (n = 6 animals) and EAE mice treated with squalene (n = 6 animals). Bar graphs represent means with individual data points. Asterisks mark significant changes, ***p < 0.001, **p < 0.01, *p < 0.05.
Extended Data Fig. 10 Working model.
a, Abca1 expression following squalene (100 µM, left) and DMHCA (10 µM, right) treatment in LPS activated BMDMs (n = 3 cultures) (Student’s t-test, two-sided). b, Expression of inflammatory genes (Tnf, Nos2) in LPS (10 ng/ml) activated WT BMDMs (n = 3 cultures) 24 h after DMHCA (10 µM) treatment. Data normalized to untreated control (n = 3 cultures, set to 1) (two-way ANOVA with Holm-Sidak post test). c, Quantitative RT-PCR analysis determining the expression housekeeping reference genes in spinal cord from EAE mice treated with DMHCA (n = 6 animals) and squalene (n = 6 animals). d, Fold expression profiles of inflammatory genes in CD11b + cells isolated from spinal cord of EAE animals treated with DMHCA (n = 6 animals) and squalene (n = 6 animals) (normalized to vehicle-treated EAE mice, n = 6 animals). Asterisks indicate significant differences between DMHCA and DMHCA/squalene co-treatment (Student’s t-test, two-sided). e, Resulting working model of the relationship between sterol metabolism in microglia/macrophages in an inflammatory demyelinating lesion and myelin repair. (1) Phagocytosis of lipid-rich myelin increases the cellular cholesterol content in microglia/macrophages, which (2) inhibits sterol synthesis by negative feedback regulation. (3) In demyelinating lesions, pro-inflammatory mediators counteract this inhibition and induce sterol synthesis. NRF1/NFE2L1 likely senses intracellular cholesterol excess and represses Dhcr24, leading to the synthesis of the sterol intermediate desmosterol. (4) Increasing desmosterol levels activate LXR-signaling. (5) This leads to LXR-mediated export of cholesterol via ABC transporters (cholesterol recycling) and (6) dampening of the pro-inflammatory phenotype of microglia/macrophages. (7) Resolved inflammation and increased disposition of cholesterol facilitates remyelination by oligodendrocytes. f, In case of normal repair of demyelinated lesions, microglia/macrophages are capable of lipid/cholesterol recycling and switching to an anti-inflammatory signature that depends on desmosterol synthesis. We speculate that this conserved mechanism mediates ‘silent’ regeneration in asymptomatic individuals and in the preclinical phase of MS. When endogenous sterol synthesis is impaired as in our SQS conditional mutants or presumably also during aging or advanced disease, these repair strategies fail leading to cholesterol overloading of microglia/macrophages, persistent inflammation and impaired regeneration. Dietary squalene supplementation supports sterol synthesis. This increases desmosterol levels in microglia/macrophages that facilitates to resolve the local inflammation and to recycle lipid/cholesterol. In addition, squalene promotes cholesterol synthesis by oligodendrocytes for remyelination that collectively enhances repair. Asterisks mark significant changes, ***p < 0.001, **p < 0.01, *p < 0.05.
Supplementary information
Supplementary Information
Supplementary Tables 1–12.
Supplementary Table 13
Exact P values of all comparisons.
Supplementary Video 1
Survival of control and genetically modified BMDMs.
Supplementary Video 2
Light-sheet microscopy of microglia in EAE control.
Supplementary Video 3
Light-sheet microscopy of microglia in SQS/McKO.
Supplementary Video 4
Light-sheet microscopy of microglia in mice with EAE.
Supplementary Video 5
Light-sheet microscopy of microglia in squalene-treated mice with EAE.
Supplementary Software
Code for data integration and analysis in R.
Source data
Source Data Fig. 2
Full scans of unprocessed immunoblots.
Rights and permissions
About this article
Cite this article
Berghoff, S.A., Spieth, L., Sun, T. et al. Microglia facilitate repair of demyelinated lesions via post-squalene sterol synthesis. Nat Neurosci 24, 47–60 (2021). https://doi.org/10.1038/s41593-020-00757-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-00757-6
This article is cited by
-
LXR agonism for CNS diseases: promises and challenges
Journal of Neuroinflammation (2024)
-
Microglia regulation of central nervous system myelin health and regeneration
Nature Reviews Immunology (2024)
-
Abcd1 deficiency accelerates cuprizone-induced oligodendrocyte loss and axonopathy in a demyelinating mouse model of X-linked adrenoleukodystrophy
Acta Neuropathologica Communications (2023)
-
Cholesterol-dependent amyloid β production: space for multifarious interactions between amyloid precursor protein, secretases, and cholesterol
Cell & Bioscience (2023)
-
Novel CH25H+ and OASL+ microglia subclusters play distinct roles in cerebral ischemic stroke
Journal of Neuroinflammation (2023)