Abstract
Sensory pathways are typically studied by starting at receptor neurons and following postsynaptic neurons into the brain. However, this leads to a bias in analyses of activity toward the earliest layers of processing. Here, we present new methods for volumetric neural imaging with precise across-brain registration to characterize auditory activity throughout the entire central brain of Drosophila and make comparisons across trials, individuals and sexes. We discover that auditory activity is present in most central brain regions and in neurons responsive to other modalities. Auditory responses are temporally diverse, but the majority of activity is tuned to courtship song features. Auditory responses are stereotyped across trials and animals in early mechanosensory regions, becoming more variable at higher layers of the putative pathway, and this variability is largely independent of ongoing movements. This study highlights the power of using an unbiased, brain-wide approach for mapping the functional organization of sensory activity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Calcium imaging data from this study (all segmented ROIs) are publicly available at https://doi.org/10.34770/gv6w-5351.
Code availability
The Matlab code for the neural imaging data processing pipeline is available at https://github.com/murthylab/FlyCaImAn.
References
Mann, K., Gallen, C. L. & Clandinin, T. R. Whole-brain calcium imaging reveals an intrinsic functional network in Drosophila. Curr. Biol. 27, 2389–2396.e4 (2017).
Chen, X. et al. Brain-wide organization of neuronal activity and convergent sensorimotor transformations in larval zebrafish. Neuron 100, 876–890.e5 (2018).
Bates, A. S. et al. The natverse, a versatile toolbox for combining and analysing neuroanatomical data. eLife 9, e53350 (2020).
Deutsch, D., Clemens, J., Thiberge, S. Y., Guan, G. & Murthy, M. Shared song detector neurons in Drosophila male and female brains drive sex-specific behaviors. Curr. Biol. 29, 3200–3215.e5 (2019).
von Schilcher, F. The role of auditory stimuli in the courtship of Drosophila melanogaster. Anim. Behav. 24, 18–26 (1976).
Clemens, J. et al. Connecting neural codes with behavior in the auditory system of Drosophila. Neuron 87, 1332–1343 (2015) ; erratum 97, 475 (2018).
Yoon, J. et al. Selectivity and plasticity in a sound-evoked male–male interaction in Drosophila. PLoS ONE 8, e74289 (2013).
Clemens, J. et al. Discovery of a new song mode in Drosophila reveals hidden structure in the sensory and neural drivers of behavior. Curr. Biol. 28, 2400–2412.e6 (2018).
Coen, P. et al. Dynamic sensory cues shape song structure in Drosophila. Nature 507, 233–237 (2014).
Coen, P., Xie, M., Clemens, J. & Murthy, M. Sensorimotor transformations underlying variability in song intensity during Drosophila courtship. Neuron 89, 629–644 (2016).
Göpfert, M. C. & Robert, D. The mechanical basis of Drosophila audition. J. Exp. Biol. 205, 1199–1208 (2002).
Yorozu, S. et al. Distinct sensory representations of wind and near-field sound in the Drosophila brain. Nature 458, 201–205 (2009).
Kamikouchi, A. et al. The neural basis of Drosophila gravity-sensing and hearing. Nature 458, 165–171 (2009).
Lai, J. S.-Y., Lo, S.-J., Dickson, B. J. & Chiang, A.-S. Auditory circuit in the Drosophila brain. Proc. Natl Acad. Sci. USA 109, 2607–2612 (2012).
Vaughan, A. G., Zhou, C., Manoli, D. S. & Baker, B. S. Neural pathways for the detection and discrimination of conspecific song in D. melanogaster. Curr. Biol. 24, 1039–1049 (2014).
Zhou, C. et al. Central neural circuitry mediating courtship song perception in male Drosophila. eLife 4, e08477 (2015).
Matsuo, E. et al. Organization of projection neurons and local neurons of the primary auditory center in the fruit fly Drosophila melanogaster. J. Comp. Neurol. 524, 1099–1164 (2016).
Patella, P. & Wilson, R. I. Functional maps of mechanosensory features in the Drosophila brain. Curr. Biol. 28, 1189–1203.e5 (2018).
Wang, K. et al. Neural circuit mechanisms of sexual receptivity in Drosophila females. Preprint at bioRxiv https://doi.org/10.1101/2020.08.07.241919 (2020).
Pnevmatikakis, E. A. et al. Simultaneous denoising, deconvolution, and demixing of calcium imaging data. Neuron 89, 285–299 (2016).
Aimon, S. et al. Fast near-whole-brain imaging in adult Drosophila during responses to stimuli and behavior. PLoS Biol. 17, e2006732 (2019).
Pnevmatikakis, E. A. & Giovannucci, A. NoRMCorre: an online algorithm for piecewise rigid motion correction of calcium imaging data. J. Neurosci. Methods 291, 83–94 (2017).
Giovannucci, A. et al. CaImAn an open source tool for scalable calcium imaging data analysis. eLife 8, e38173 (2019).
Yang, H. H. et al. Subcellular imaging of voltage and calcium signals reveals neural processing in vivo. Cell 166, 245–257 (2016).
Ito, K. et al. A systematic nomenclature for the insect brain. Neuron 81, 755–765 (2014).
Myronenko, A. & Song, X. Point set registration: coherent point drift. IEEE Trans. Pattern Anal. Mach. Intell. 32, 2262–2275 (2010).
Tootoonian, S., Coen, P., Kawai, R. & Murthy, M. Neural representations of courtship song in the Drosophila brain. J. Neurosci. 32, 787–798 (2012).
Azevedo, A. W. & Wilson, R. I. Active mechanisms of vibration encoding and frequency filtering in central mechanosensory neurons. Neuron 96, 446–460.e9 (2017).
Chang, A. E. B., Vaughan, A. G. & Wilson, R. I. A mechanosensory circuit that mixes opponent channels to produce selectivity for complex stimulus features. Neuron 92, 888–901 (2016).
Scheffer, L. K. et al. A connectome and analysis of the adult Drosophila central brain. Elife 9, e57443 (2020).
Lehnert, B. P., Baker, A. E., Gaudry, Q., Chiang, A.-S. & Wilson, R. I. Distinct roles of TRP channels in auditory transduction and amplification in Drosophila. Neuron 77, 115–128 (2013).
Senthilan, P. R. et al. Drosophila auditory organ genes and genetic hearing defects. Cell 150, 1042–1054 (2012).
Musall, S., Kaufman, M. T., Juavinett, A. L., Gluf, S. & Churchland, A. K. Single-trial neural dynamics are dominated by richly varied movements. Nat. Neurosci. 22, 1677–1686 (2019).
Stringer, C. et al. Spontaneous behaviors drive multidimensional, brainwide activity. Science 364, 255 (2019).
Baker, C. A. et al. Neural network organization for courtship song feature detection in Drosophila. Preprint at bioRxiv https://doi.org/10.1101/2020.10.08.332148 (2020).
Harris, D. T., Kallman, B. R., Mullaney, B. C. & Scott, K. Representations of taste modality in the Drosophila brain. Neuron 86, 1449–1460 (2015).
Naumann, E. A. et al. From whole-brain data to functional circuit models: the zebrafish optomotor response. Cell 167, 947–960.e20 (2016).
Haesemeyer, M., Robson, D. N., Li, J. M., Schier, A. F. & Engert, F. A brain-wide circuit model of heat-evoked swimming behavior in larval zebrafish. Neuron 98, 817–831.e6 (2018).
Frechter, S. et al. Functional and anatomical specificity in a higher olfactory centre. eLife 8, e44590 (2019).
Auer, T. O. & Benton, R. Sexual circuitry in Drosophila. Curr. Opin. Neurobiol. 38, 18–26 (2016).
Vogt, K. et al. Shared mushroom body circuits underlie visual and olfactory memories in Drosophila. eLife 3, e02395 (2014).
Jefferis, G. S. X. E. et al. Comprehensive maps of Drosophila higher olfactory centers: spatially segregated fruit and pheromone representation. Cell 128, 1187–1203 (2007).
Bennet-Clark, H. C. & Ewing, A. W. Pulse interval as a critical parameter in the courtship song of Drosophila melanogaster. Anim. Behav. 17, 755–759 (1969).
Murthy, M., Fiete, I. & Laurent, G. Testing odor response stereotypy in the Drosophila mushroom body. Neuron 59, 1009–1023 (2008).
Chiappe, M. E., Seelig, J. D., Reiser, M. B. & Jayaraman, V. Walking modulates speed sensitivity in Drosophila motion vision. Curr. Biol. 20, 1470–1475 (2010).
Fujiwara, T., Cruz, T. L., Bohnslav, J. P. & Chiappe, M. E. A faithful internal representation of walking movements in the Drosophila visual system. Nat. Neurosci. 20, 72–81 (2017).
Kim, A. J., Fitzgerald, J. K. & Maimon, G. Cellular evidence for efference copy in Drosophila visuomotor processing. Nat. Neurosci. 18, 1247–1255 (2015).
Batchelor, A. V. & Wilson, R. I. Sound localization behavior in Drosophila melanogaster depends on inter-antenna vibration amplitude comparisons. J. Exp. Biol. 222, jeb191213 (2019).
Rayshubskiy, A. et al. Neural control of steering in walking Drosophila. Preprint at bioRxiv https://doi.org/10.1101/2020.04.04.024703 (2020).
Zheng, Z. et al. A complete electron microscopy volume of the brain of adult Drosophila melanogaster. Cell 174, 730–743.e22 (2018).
Effertz, T., Wiek, R. & Göpfert, M. C. NompC TRP channel is essential for Drosophila sound receptor function. Curr. Biol. 21, 592–597 (2011).
Morley, E. L., Jonsson, T. & Robert, D. Auditory sensitivity, spatial dynamics, and amplitude of courtship song in Drosophila melanogaster. J. Acoust. Soc. Am. 144, 734 (2018).
Seelig, J. D. et al. Two-photon calcium imaging from head-fixed Drosophila during optomotor walking behavior. Nat. Methods 7, 535–540 (2010).
Moore, R. J. D. et al. FicTrac: a visual method for tracking spherical motion and generating fictive animal paths. J. Neurosci. Methods 225, 106–119 (2014).
Murthy, M. & Turner, G. Whole-cell in vivo patch-clamp recordings in the Drosophila brain. Cold Spring Harb. Protoc. 2013, 140–148 (2013).
Maimon, G., Straw, A. D. & Dickinson, M. H. Active flight increases the gain of visual motion processing in Drosophila. Nat. Neurosci. 13, 393–399 (2010).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Preibisch, S., Saalfeld, S. & Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinformatics 25, 1463–1465 (2009).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Park, M. & Pillow, J. W. Receptive field inference with localized priors. PLoS Comput. Biol. 7, e1002219 (2011).
Cachero, S., Ostrovsky, A. D., Yu, J. Y., Dickson, B. J. & Jefferis, G. S. X. E. Sexual dimorphism in the fly brain. Curr. Biol. 20, 1589–1601 (2010).
Rohlfing, T. & Maurer, C. R. Jr. Nonrigid image registration in shared-memory multiprocessor environments with application to brains, breasts, and bees. IEEE Trans. Inf. Technol. Biomed. 7, 16–25 (2003).
Chiang, A.-S. et al. Three-dimensional reconstruction of brain-wide wiring networks in Drosophila at single-cell resolution. Curr. Biol. 21, 1–11 (2011).
Peng, H. et al. BrainAligner: 3D registration atlases of Drosophila brains. Nat. Methods 8, 493–500 (2011).
Yu, J. Y., Kanai, M. I., Demir, E., Jefferis, G. S. X. E. & Dickson, B. J. Cellular organization of the neural circuit that drives Drosophila courtship behavior. Curr. Biol. 20, 1602–1614 (2010).
Acknowledgements
We thank J. Clemens for assistance on auditory stimuli delivery and linear modeling of calcium responses, as well as helpful discussions on auditory coding; A. Giovannucci for help with ROI segmentation of volumetric calcium signals and guidance with CaImAn toolbox usage; G. Jefferis and T. Rohlfing for help using the image registration toolbox CMTK and the neuroanatomy toolbox natverse; B. Cowley for help with linear modeling of calcium responses in behaving flies; A. Calhoun for help with analysis of the hemibrain connectome dataset; and T. Pereira for assistance with the CPD algorithm. We also thank G. Jefferis, S. Ahmed, C. Baker and J. Clemens for comments on the manuscript. D.A.P. was supported in part by a NSF Physics Frontier Center grant, and M.M. was supported by a NIH NINDS New Innovator award, NIH BRAIN Initiative R01s NS104899 and NS110060, and a Howard Hughes Medical Institute Faculty Scholar award.
Author information
Authors and Affiliations
Contributions
D.A.P. and M.M. designed the study. D.A.P. collected and analyzed data. S.Y.T. designed and built the two-photon imaging microscope. E.P. generalized ROI segmentation from 2D to 3D datasets. D.A.P. and M.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Neuroscience thanks Ioana Carcea, Elizabeth Hillman, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Spatial coverage of imaged activity and measurement of auditory-evoked activity.
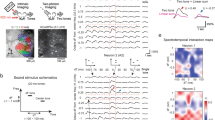
a, Data collection and processing. We collected data using the following protocols: a) short volumetric recordings (~10 min) of consecutive segments (from anterior to posterior axis), b) long volumetric recordings (~30 min) of selected brain areas (non-consecutive), and c) short and coarse volumetric recording (~15 min) of a single segment per fly that spans a large volume. For all these datasets we record a private whole brain structural volume (used later for registration). For all the data sets we recorded both tdTomato and GCaMP signal, and performed motion correction (using NoRMCorre) on each imaged segment using the tdTomato signal. These segments were then spatially resampled to have isotropic XY pixel size. This was followed by re-slicing of each Z-stack per segment (align time of all planes per Z-stack to the first plane imaged), and temporal resampling (aling Z-stack time relative to the start of the 1st stimulus and double the sampling rate). For protocol a), we stitched segments imaged consecutively (using NoRMCorre) obtaining a ‘volume’; for b) and c) each segment was treated as an independent volume. Volumes were mirrored to the right hemisphere, and the tdtomato signal was used for registration to the in vivo intersex atlas (IVIA). This is a two-step process, i) volumes per fly were registered to their own private whole brain volume (one per fly), and ii) whole brain volume was registered to the IVIA; registration i) and ii) were concatenated to map volumes to IVIA space. The GCaMP signal was used for ROI segmentation (via CaImAn), followed by identification of stimulus-modulated ROIs (see Extended Data Fig. 1d). b, Maximum projection (in each dimension) of segmented ROIs from all imaged volumes (n = 33 flies, 185,395 ROIs) - ROIs from the left hemisphere are mirrored (see Extended Data Fig. 1a) such that all ROIs are projected onto right hemisphere. ROIs cover the entirety of the D-V and A-P axes. Color scale indicates the number of flies with an ROI in each voxel. c, Number of ROIs across neuropils (see Fig. 1f,g and Supplementary Movies 1–4) sampled by ventral volumes or dorsal volumes (n = 33 flies, 185,395 ROIs). d, Method for identifying stimulus-modulated ROIs. Raw Ca++ signal (F(t)) is convolved with the stimulus history (f(t)) and a set of filters per stimulus type (q(τ)) to generate the predicted Ca+ + signal. Auditory modulation is measured by the cross-validated correlation scores (⍴F,g) between raw and predicted Ca++ signals. Correlation of shuffled Ca++ signal (sF(t)) to predicted signal (g(t)) is used to generate the null-distribution of correlation scores (⍴sF,g), which is used to determine significance. e, Distribution of Pearson correlation coefficients of shuffled-vs-predicted signals (⍴sF,g) and raw-vs-predicted signal (⍴F,g) across all flies imaged (n = 33 flies). Unlike the distribution of ⍴sF,g, ⍴F,g has a distribution with a long tail of positive correlation scores. ROIs within the positive tail and outside the null distribution are considered to have significant stimuli modulation and selected as auditory ROIs. f, Frequency of auditory ROIs separated by whether the ROI was segmented from a dorsal or ventral volume (n = 33 flies).
Extended Data Fig. 2 Building the in vivo intersex atlas (IVIA) and registering it to fixed-brain atlas (IBNWB).
a, Generation of in vivo intersex atlas (IVIA): images of male (n = 5) and female (n = 8) brains expressing membranal tdTomato pan-neuronally are registered to a seed brain (reference image). Images are then transformed to generate an average model image, which, after five iterations, produces the in vivo intersex atlas (IVIA). b, IVIA registration accuracy. Left: 3D-rendering of traced tracts of Dsx+ neurons (pC2m, pC2l), and Fru+ neuron pMP7. Right: Per-axis jitter (X, Y, and Z) between matched traced tracts of pC2m, pC2l, and pMP7 across flies (n = 10, 12, and 7 brains, respectively). c, Example imaged tdTomato volumes (red scale) from different flies registered to the IVIA (right-most column, gray scale). Middle column is the intermediate registration of volumes to their own private whole brain atlas (as in Extended Data Fig. 1a). 45 out of 48 flies were successfully registered to the IVIA. d, Central brain neuropils and tracts used for point set registration to morph the in vivo intersex atlas (IVIA) to IBNWB fixed brain atlas. Points from meshes of segmented antennal lobe (AL), mushroom body (MB) (includes mushroom body lobes, peduncle and calyx), protocerebral bridge (PB), antennal mechanosensory and motor center commissure (AMMCC), anterior optic tract (AOT), great commissure (GC), wedge commissure (WEDC), posterior optic commissure (POC), lateral antennal lobe tract (lALT), posterior cerebro-cervical fascicle (pCCF), superior saddle commissure (sSADC), and whole central brain were used to generate IVIA-to-IBNWB and IBNWB-to-IVIA transformations. e, Overlay of IVIA (red) and registered IBNWB (in IVIA space, green) at different depths (90, 130, 180, 220, 260, and 280 µm). 0 μm is the most anterior section of the brain and 300 um the most posterior. f,g, Atlas-to-atlas registration accuracy measured using pC1 stalks from IBNWB and IVIA. (f) pC1 traces from IBNWB and IVIA atlases; black traces are single pC1 neurons (from IBNWB or IVIA, n = 70 and 20 pC1 traces respectively) and red trace is the mean reference pC1 stalk. (g) Between-atlas registration accuracy; IVIA-to-IBNWB transformation increases the jitter across all axes from the reference mean pC1 trace by ~2.24 µm, while IBNWB-to-IVIA transformation increases the jitter by ~2.8 μm.
Extended Data Fig. 3 Auditory responses in neuropils and neurite tracts, and to additional song stimuli: natural song and fast pulses.
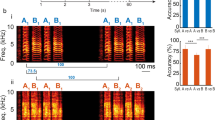
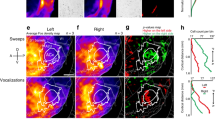
a, Median ROI responses (across 6 trials) to pulse, sine, and white noise stimuli (n = 33 flies, 19,036 ROIs) as in Fig. 2a, but without z-scoring ΔF/F signal. b, Spatial distribution of auditory activity across sexes. Maximum projections (from two orthogonal views) of the density of auditory ROIs in female (n = 17) or male flies (n = 16). c, Maximum projection (from two orthogonal views) of the density of i) auditory ROIs outside neuropils or neurite tracts (n = 33 flies, 4,346 ROIs), and ii) auditory ROIs within neuropils or neurite tracts (n = 33 flies, 14,658 ROIs). Color scale for (B) and (C) is the number of flies with an auditory ROI per voxel. d, Percentage of auditory ROIs within and outside neuropils and tracts, and beyond the midline. e, Spectral profile of auditory stimuli - pulse (Pslow), sine, and white noise - used to classify response types in Fig. 3a, and their spectral features. f, Spectral profile of natural song stimulus used. g, Distribution of stimuli preference (to pulse, sine, white noise, and natural song) across auditory ROIs (n = 5 flies, 2,258 ROIs). Only ~20% of auditory ROIs prefer natural song. Preference is defined by the stimulus that drives the maximum absolute response (at least 15% greater than the second highest response), as in Fig. 3f. h, Responses from auditory ROIs that prefer natural song (n = 451 ROIs). Each row is the median z-scored ΔF/F response across 6 across trials. Activity is plotted as the change in the s.d. of the ΔF/F signal. i, Spatial distribution of natural song preferring ROIs. Images are the maximum projection (from two orthogonal views) of the density of auditory ROIs with preference for natural song throughout the central brain (n = 5, 451 ROIs). j, Spectral profile of Pfast and Pslow stimuli. k, Distribution of stimuli preference to Pslow, Pfast, or broad preference for both pulse types across auditory ROIs (n = 2 flies, 2,193 ROIs). Only ~4% of auditory ROIs prefer Pfast. Preference is defined as in (G). l, Responses from auditory ROIs that prefer Pfast (n = 106 ROIs). Pfast preferring ROIs also show strong responses to Pslow. Conventions same as in (H). m, Fraction of voxels with auditory activity by central brain neurite tracts; percentages averaged across 33 flies (a minimum of 4 flies with auditory activity in a given tract was required for inclusion). Red represents tracts that were clearly distinguishable from neuropil by visual inspection (see (N)). n, For three flies, pixels with auditory activity (red) overlaid on time-averaged GCaMP6 fluorescence (grayscale). Several neurite tracts are indicated (planes from different depths are arbitrarily selected for each fly to highlight ROIs contained within neurite tracts).
Extended Data Fig. 4 Auditory activity is present in olfactory pathway neurons and absent in deaf flies.
a,b, Auditory responses from (A) putative individual antennal lobe projection neurons (PNs), and (B) putative individual mushroom body Kenyon cells (KCs) from pan-neuronal recordings. Time-averaged GCaMP6s signal is shown in grayscale (over the entire experiment). Pixels belonging to individual ROIs and their corresponding time traces (median z-scored ΔF/F responses across 6 trials to pulse, sine, and white noise stimuli) are indicated in different colors. ROIs were drawn manually over the location of PN or KC somas from two independently imaged flies. Activity scale bar unit is s.d. c,d, Auditory responses from (C) antennal lobe projection neurons, and (D) mushroom body Kenyon cells using cell type specific genetic lines (GH146-Gal4 and OK107-Gal4, respectively). Time-averaged GCaMP6s signal is shown in grayscale (over the entire experiment) - the compartments where activity was recorded from are indicated (data was collected and processed as we did for pan-neuronal data (protocol a) in Extended Data Fig. 1a) but the field of view was restricted to the region defined by the orange and purple boxes). Auditory ROIs are detected in all imaged compartments of antennal lobe projection neurons (n = 8 flies, 1,133 ROIs) and Kenyon cells (n = 4 flies, 154 ROIs). Each row is the median across 6 trials - all responses are z-scored and therefore plotted as the change in the s.d. of the ΔF/F signal. e, Number of detected auditory ROIs per fly in wild type (n = 33 flies) and iav1 flies (n = 4 flies). f, Maximum projection (from two orthogonal views) of the density of auditory ROIs in iav1 flies (all 21 ROIs come from 1 out of 4 flies imaged). These few ROIs were located in the mushroom body calyx (MB-CA), posterior VLP (PVLP), and the posterior lateral protocerebrum (PLP). Color scale is the number of flies with an auditory ROI per voxel. g, Distribution of mean ΔF/F values (during stimuli) for auditory ROIs in wild type (n = 33 flies) and iav1 flies (n = 4 flies).
Extended Data Fig. 5 Auditory responses are not sexually dimorphic and Auditory response types have distinct spatial distributions throughout the central brain.
a, Hierarchical clustering of auditory responses into 18 distinct response types, and divided by sex. The mean response across ROIs belonging to each response type is shown. All responses are z-scored (as in Fig. 3a), and therefore plotted as the s.d. of the responses over time. Color code is the same as in Fig. 3a. b, Probabilities of neuropil voxels with a given response type (separated by sex) in two neuropils, the gnathal ganglion (GNG) and the lateral accessory lobe (LAL). These are the only two neuropils with sex differences - males have a slightly higher probability of response type 18 activity in the GNG, while females have a slightly higher probability of response type 13 activity in the LAL. Each dot is the probability for one fly. c, Sex-related differences in all response types across all neuropils. Each dot is the effect size (see Methods) of the difference in probability for each response type across sexes, color code is the same as (A). Neuropils with the greatest effect size are the GNG (response type 18) and LAL (response type 13). However all differences are not significant (p > 0.05). Statistical significance was determined using two-tailed two-sample t-test with Benjamini-Hochberg FDR correction. Neuropils with no auditory activity are indicated in gray font. d, Spatial distribution of excitatory vs inhibitory auditory ROI responses across the central brain. Images are the maximum projection (from two orthogonal views) of the density of excitatory and inhibitory responses across flies (n = 33 flies). e, Spatial distribution of auditory activity belonging to each response type (see Fig. 3a) across the central brain. Images are the maximum projection (from two orthogonal views) of the density of responses across flies for each response type (n = 33 flies). Color scale for (D) and (E) is the number of flies with an auditory ROI belonging to each category (excitatory, inhibitory, or response type) per voxel.
Extended Data Fig. 6 Similarity of auditory activity between neuropils, and compartmentalization within neuropils.
a, Pairwise Pearson’s correlation coefficients of response type distributions (as in Fig. 3b) between neuropils. Neuropils are ordered based on the hierarchical clustering of the correlation matrix. Only positive correlation values are plotted for clarity. b, Spatial distribution of auditory responses within selected olfactory (AL, MB-ML, MB-VL, MB-PED, MB-CA, and LH), visual (PVLP, and PLP) and mechanosensory neuropils (AVLP). Auditory activity is spatially restricted within each neuropil. Voxel density was calculated from 19, 15, 17, 17, 22, 19, 18, 30, and 25 flies for the AL, MB-ML, MB-VL, MB-PED, MB-CA, LH, PVLP, PLP, and AVLP respectively.
Extended Data Fig. 7 Neuropil subregions imaged, definition of stimulus tuning, and tuning for additional song features.
a, Neuropil volume and subregion of neuropil imaged in Fig. 4. Top row, location of the neuropil imaged (dark grey surface) relative to the central brain (light grey surface). Middle and bottom rows show maximum projections (from two orthogonal views) of imaged neuropil subvolume (orange surface) and the distribution of ROI stimulus selectivity (based on data in Fig. 3b). The neuropil subvolume (orange) is the volume imaged in at least 2 flies. This volume represents 24.4, 46.2, 10.5, 83, 39.8, 99.3, 63.6, and 91.8 % of the AMMC, SAD, GNG, WED, AVLP, PVLP, PLP and LH, respectively. b, Example responses from one ROI to song feature stimuli (see Fig. 6a). Top panel, Median (across trials) and z-scored responses to each song feature (responses are baseline subtracted, and baseline is defined as activity −4 to −0.25 seconds before stimulus onset). Orange timepoints correspond to activity during the stimulus. Bottom panel, response magnitudes (80th or 20th percentile of activity - for excitatory or inhibitory responses - during stimulus plus 2 seconds after) to each song feature calculated from the top trace (that is the tuning curve for this ROI). All responses are z-scored, so responses are plotted as the s.d. of ΔF/F value. c, Tuning types as in Fig. 6c, but plotting additional responses to pulses and sines of different amplitudes, varying pulse and sine train durations, and also to white noise and natural song (these additional responses were not used to cluster responses). Thick traces are the mean response magnitudes (calculated as in (B)) across all ROIs within each tuning type and shading is the standard deviation (1,783, 513, 739, 1,682, 2,410, 2,321 and 1424 ROIs for each tuning type 1–7 respectively, n = 21 flies). Responses are plotted as the s.d. of ΔF/F value. d, Auditory responses to different sine and pulse intensities, sorted by tuning type. Response magnitudes (calculated as in (B), but 80th or 20th percentile of the activity is measured during stimuli only) per ROI are normalized to the response to 0.5 mm/s stimuli. Thick traces are the mean normalized response magnitude, and shading is the s.e.m (ROI number per tuning type is the same as in (C)). e, Auditory responses to different sine and pulse train durations. Response magnitudes (calculated as in (D)) per ROI are normalized to the response to 2 seconds stimuli. Conventions same as in (D).
Extended Data Fig. 8 Across-trial and across-individual comparisons of auditory activity by neuropil.
a, Comparison of across-trial variability index (see Fig. 5b) and response magnitude. Each dot is the mean variability index and response magnitude (80th or 20th percentile - for excitatory or inhibitory responses - of z-scored ΔF/F from stimuli onset until 5 seconds after stimulus offset) per neuropil for each fly (across all ROIs). Neuropils are color coded according to the legend. Variability index is inversely correlated with response magnitude. b, Comparison of across-trial variability (see Fig. 5b) and across-individual similarity (see Fig. 5d). Each dot is the mean across-trial variability index and mean across-individual similarity index per neuropil. Groups of neuropils are color coded according to the legend. Early mechanosensory neuropils have low across-trial variability and high across-individual similarity. c, Robustness of differences in similarity index across neuropils to the number clusters selected for hierarchical clustering of response types (see Fig. 3a). Similarity index per neuropil (as in Fig. 5d) is calculated for different numbers of clusters (from 10 to 26 - in Fig. 3 we used 18 types (red)). d, No systematic difference in distribution of time-average fluorescence (over the entire experiment) across individuals. Left: Histograms of time-averaged fluorescence per individual, cyan and magenta correspond to male (n = 16) and female (n = 17) flies. Right: Median fluorescence per individual. Black dot corresponds to the mean (of median fluorescence) across flies.
Extended Data Fig. 9 Distribution of 18 auditory response types per neuropil, separated by flies.
Response type distributions, similar to Fig. 5c, for mechanosensory, visual, olfactory, central and lateral complex neuropils. For each neuropil, male and female flies are indicated in cyan or magenta, respectively.
Extended Data Fig. 10 Behavior of head-fixed flies and minimal auditory responses in the Drosophila ventral nerve cord.
a, Distribution of across-trial variability of auditory ROIs in behaving (data used in Fig. 6, n = 7 flies, 4,560 ROIs) and non-behaving flies (data used in Fig. 5b, n = 33 flies, 19,036 ROIs). b, Distribution of speed across flies. Each gray line is the speed distribution per fly (n = 7 flies), and black line is the mean across animals. c, Correlation between linearly predicted velocities (based on stimuli history) and actual or shuffled (see Methods) velocities. Each black or gray line is the probability per fly (n = 7 flies). d, Schematic of VNC functional imaging during auditory stimulation. Similar to Fig. 1a, but the dorsal side of the thorax is dissected to expose the dorsal side of the first and second segment of the VNC depicted inside the orange rectangle. e, Frequency of auditory ROIs detected in the brain (n = 33 flies, 185,395 ROIs) and the VNC (n = 8 flies, 39,580 ROIs) using the same criteria as in Extended Data Fig. 1d. f, Distribution of Pearson correlation coefficients of raw ROI activity to predicted ROI activity (based on stimuli history, as in Extended Data Fig. 1e) for ROIs recorded from the brain (n = 33 flies, 185,395 ROIs) and the VNC (n = 8 flies, 39,580 ROIs). g, Responses of VNC auditory ROIs to pulse, sine, and white noise stimuli (n = 8 flies, 39,580 ROIs). Each row is the across-trial median and z-scored ΔF/F response to each stimulus (6 trials per stimulus). ΔF/F units are in s.d.
Supplementary information
Supplementary Table 1
Full names and abbreviations of brain neuropils and tracts.
Supplementary Table 2
Parameters for all auditory stimuli.
Supplementary Video 1
ΔF/F0 time-series movie (maximum intensity projected over the entire z axis of the Drosophila brain) of a male dorsal quadrant. Responses to pulse (250 Hz), sine (150 Hz) and white noise are shown. The pseudocolor scale is from 0 to 2 ΔF/F0. The movie speed is sped up 5×.
Supplementary Video 2
ΔF/F0 time-series movie of a male ventral quadrant. Conventions are the same as for Supplementary Video 1.
Supplementary Video 3
ΔF/F0 time-series movie of a female dorsal quadrant. Conventions are the same as for Supplementary Video 1.
Supplementary Video 4
ΔF/F0 time-series movie of a female ventral quadrant. Conventions are the same as for Supplementary Video 1.
Supplementary Video 5
A z-stack of the probability density of auditory ROIs (number of flies containing an auditory ROI at each voxel) throughout the central brain (n = 33 flies, 19,036 ROIs). Mechanosensory neuropils have gray contours, while the rest of the brain neuropils imaged have red contours. The pseudocolor scale is from 0 to 15 flies per voxel.
Rights and permissions
About this article
Cite this article
Pacheco, D.A., Thiberge, S.Y., Pnevmatikakis, E. et al. Auditory activity is diverse and widespread throughout the central brain of Drosophila. Nat Neurosci 24, 93–104 (2021). https://doi.org/10.1038/s41593-020-00743-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-00743-y
This article is cited by
-
Flexible circuit mechanisms for context-dependent song sequencing
Nature (2023)
-
The spatial and temporal structure of neural activity across the fly brain
Nature Communications (2023)
-
Differential mechanisms underlie trace and delay conditioning in Drosophila
Nature (2022)
-
Imaging whole-brain activity to understand behaviour
Nature Reviews Physics (2022)
-
Large-scale neural recordings call for new insights to link brain and behavior
Nature Neuroscience (2022)