Abstract
Understanding how genes, drugs and neural circuits influence behavior requires the ability to effectively organize information about similarities and differences within complex behavioral datasets. Motion Sequencing (MoSeq) is an ethologically inspired behavioral analysis method that identifies modular components of three-dimensional mouse body language called ‘syllables’. Here, we show that MoSeq effectively parses behavioral differences and captures similarities elicited by a panel of neuroactive and psychoactive drugs administered to a cohort of nearly 700 mice. MoSeq identifies syllables that are characteristic of individual drugs, a finding we leverage to reveal specific on- and off-target effects of both established and candidate therapeutics in a mouse model of autism spectrum disorder. These results demonstrate that MoSeq can meaningfully organize large-scale behavioral data, illustrate the power of a fundamentally modular description of behavior and suggest that behavioral syllables represent a new class of druggable target.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All datasets generated and/or analyzed during the current study will be available from the corresponding author upon reasonable request. The raw per-frame data, MoSeq per-frame labels and per-mouse behavioral summary data organized as NumPy arrays are stored in a Python pickle file and are available for download on an open-access basis via GitHub (https://github.com/dattalab/moseq-drugs).
Code availability
All code used in this manuscript will be made available on GitHub at https://github.com/dattalab/moseq-drugs.
References
Tinbergen, N. The Study of Instinct (Clarendon Press, 1951).
Dawkins, R. in Growing Points in Ethology (eds Bateson, P. P. G. & Hinde, R. A.) 7–54 (Cambridge Univ. Press, 1976).
Datta, S. R., Anderson, D. J., Branson, K., Perona, P. & Leifer, A. Computational neuroethology: a call to action. Neuron 104, 11–24 (2019).
Anderson, D. J. & Perona, P. Toward a science of computational ethology. Neuron 84, 18–31 (2014).
Mathis, A. et al. DeepLabCut: markerless pose estimation of user-defined body parts with deep learning. Nat. Neurosci. 21, 1281–1289 (2018).
Meyer, A. F., Poort, J., O’Keefe, J., Sahani, M. & Linden, J. F. A head-mounted camera system integrates detailed behavioral monitoring with multichannel electrophysiology in freely moving mice. Neuron 100, 46–60.e7 (2018).
Klaus, A. et al. The spatiotemporal organization of the striatum encodes action space. Neuron 95, 1171–1180.e7 (2017).
Pereira, T. D. et al. Fast animal pose estimation using deep neural networks. Nat. Methods 16, 117–125 (2019).
Wiltschko, A. B. et al. Mapping sub-second structure in mouse behavior. Neuron 88, 1121–1135 (2015).
Graving, J. M. et al. Fast and robust animal pose estimation. eLife 8, e47994 (2019).
Markowitz, J. E. et al. The striatum organizes 3D behavior via moment-to-moment action selection. Cell 174, 44–58.e17 (2018).
Crawley, J. N. Behavioral phenotyping of rodents. Comp. Med. 53, 140–146 (2003).
Crawley, J. N. Behavioral phenotyping strategies for mutant mice. Neuron 57, 809–818 (2008).
Crabbe, J. C. Genetics of mouse behavior: interactions with laboratory environment. Science 284, 1670–1672 (1999).
Wahlsten, D. et al. Different data from different labs: lessons from studies of gene–environment interaction. J. Neurobiol. 54, 283–311 (2002).
Egnor, S. E. R. & Branson, K. Computational analysis of behavior. Annu. Rev. Neurosci. 39, 217–236 (2016).
Berman, G. J., Choi, D. M., Bialek, W. & Shaevitz, J. W. Mapping the stereotyped behaviour of freely moving fruit flies. J. R. Soc. Interface https://doi.org/10.1098/rsif.2014.0672 (2014).
Fentress, J. C. & Stilwell, F. P. Grammar of a movement sequence in inbred mice. Nature 244, 52–53 (1973).
Berridge, K. C., Fentress, J. C. & Parr, H. Natural syntax rules control action sequence of rats. Behav. Brain Res. 23, 59–68 (1987).
Peñagarikano, O. et al. Absence of CNTNAP2 leads to epilepsy, neuronal migration abnormalities, and core autism-related deficits. Cell 147, 235–246 (2011).
Zetler, G. Haloperidol catalepsy in grouped and isolated mice. Pharmacology 13, 526–532 (1975).
Millichap, J. G. & Boldrey, E. E. Studies in hyperkinetic behavior. II. Laboratory and clinical evaluations of drug treatments. Neurology 17, 467–471 (1967).
Ebenezer, I. S. Neuropsychopharmacology and Therapeutics (Wiley, 2015).
Duncan, G. E., Zorn, S. & Lieberman, J. A. Mechanisms of typical and atypical antipsychotic drug action in relation to dopamine and NMDA receptor hypofunction hypotheses of schizophrenia. Mol. Psychiatry 4, 418–428 (1999).
Roth, B. L., Sheffler, D. J. & Kroeze, W. K. Magic shotguns versus magic bullets: selectively non-selective drugs for mood disorders and schizophrenia. Nat. Rev. Drug Discov. 3, 353–359 (2004).
McOmish, C. E., Lira, A., Hanks, J. B. & Gingrich, J. A. Clozapine-induced locomotor suppression is mediated by 5-HT2A receptors in the forebrain. Neuropsychopharmacolgy 37, 2747–2755 (2012).
Volkow, N. D. et al. Effects of modafinil on dopamine and dopamine transporters in the male human brain: clinical implications. JAMA 301, 1148–1154 (2009).
Zolkowska, D. et al. Evidence for the involvement of dopamine transporters in behavioral stimulant effects of modafinil. J. Pharmacol. Exp. Ther. 329, 738–746 (2009).
Alarcón, M. et al. Linkage, association, and gene-expression analyses identify CNTNAP2 as an autism-susceptibility gene. Am. J. Hum. Genet. 82, 150–159 (2008).
Rodenas-Cuadrado, P., Ho, J. & Vernes, S. C. Shining a light on CNTNAP2: complex functions to complex disorders. Eur. J. Hum. Genet. 22, 171–178 (2014).
Brunner, D. et al. Comprehensive analysis of the 16p11.2 deletion and null Cntnap2 mouse models of autism spectrum disorder. PLoS ONE 10, e0134572 (2015).
So, H.-C. et al. Analysis of genome-wide association data highlights candidates for drug repositioning in psychiatry. Nat. Neurosci. 20, 1342–1349 (2017).
Ferreri, F. et al. The in vitro actions of loxapine on dopaminergic and serotonergic receptors. Time to consider atypical classification of this antipsychotic drug? Int. J. Neuropsychopharmacol. 21, 355–360 (2018).
Datta, S. R. Q&A: understanding the composition of behavior. BMC Biol. 17, 44 (2019).
Brown, A. E. X., Yemini, E. I., Grundy, L. J., Jucikas, T. & Schafer, W. R. A dictionary of behavioral motifs reveals clusters of genes affecting Caenorhabditis elegans locomotion. Proc. Natl Acad. Sci. USA 110, 791–796 (2013).
Berman, G. J., Choi, D. M., Bialek, W. & Shaevitz, J. W. Mapping the stereotyped behaviour of freely moving fruit flies. J. R. Soc. Interface 11, 20140672 (2014).
Vogelstein, J. T. et al. Discovery of brainwide neural-behavioral maps via multiscale unsupervised structure learning. Science 344, 386–392 (2014).
Swinney, D. C. & Anthony, J. How were new medicines discovered? Nat. Rev. Drug Discov. 10, 507–519 (2011).
Hendriksen, H. & Groenink, L. Back to the future of psychopharmacology: a perspective on animal models in drug discovery. Eur. J. Pharmacol. 759, 30–41 (2015).
Witt, P. N. Drugs alter web-building of spiders: a review and evaluation. Behav. Sci. 16, 98–113 (1971).
Bishop, C. M. Pattern Recognition and Machine Learning (Springer, 2006).
McLachlan, G. J. Discriminant Analysis and Statistical Pattern Recognition (Wiley, 2004).
Fukunaga, K. & Olsen, D. R. An algorithm for finding intrinsic dimensionality of data. IEEE Trans. Computers 20, 176–183 (1971).
Acknowledgements
We thank members of the Datta Lab for helpful comments on the manuscript. We thank O. Mazor and P. Gorelik from the Research Instrumentation Core Facility for engineering support, S. Knemeyer for mouse illustrations and C. Ashton for technical assistance. Core facility support is provided by NIH grant P30 HD18655. Pilot experiments for this paper were supported by Hoffman LaRoche. S.R.D. is supported by the National Institutes of Health (U24NS109520, RO11DC016222, U19NS113201 and RO1NS114020), by a SFARI grant from the Simons Foundation and by the Simons Collaboration on the Global Brain.
Author information
Authors and Affiliations
Contributions
A.B.W., T.T., R.E.P. and S.R.D. conceived and designed experiments. A.B.W., T.T., A.Z., R.A., R.E.P. and J.K. acquired behavioral data and performed data pre-processing. A.B.W., T.T., A.Z., R.A., J.E.M., W.F.G. and M.J.J. carried out data analyses. A.B.W. and S.R.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests: A.B.W., M.J.J. and S.R.D. are co-founders of Syllable Life Sciences, Inc. A.B.W. and S.R.D. are co-authors on awarded patents WO2013170129A1 and US10025973B2, which describe behavioral methods used herein.
Additional information
Peer review information Nature Neuroscience thanks Ann Kennedy, Paul Kenny, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
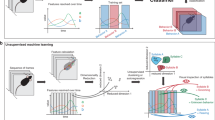
Extended Data Fig. 1 Workflow for classifying drug effects in mice using scalars and MoSeq.
Depth cameras are used to capture 3D video data encapsulating mouse postural dynamics in the open field. These data are saved locally before being uploaded to the cloud, where the videos are denoised and aligned. The image of the mouse is then extracted from the larger image; at this step, scalar behavioral metrics (like the position of the mouse within the arena, or its velocity) are computed. After extraction, aligned 3D mouse images are analyzed either locally or in the cloud, depending upon resource demands. 3D mouse images are compressed by PCA (for ease of computation), then these data are used to train an AR-HMM (as in Wiltschko et al9). The output of this training procedure is the optimal set of behavioral syllables that describe the 3D pose dynamics observed within the experiment (each of which is described as an autoregressive process through pose space). Every frame of the imaging data is then labeled with behavioral syllable MoSeq considers most likely, thereby revealing the behavioral grammar that governs the transitions from any given syllable to any other syllable. Herein, each mouse is characterized by a MoSeq behavioral summary that includes only information about how often each behavioral syllable is expressed during the experiment (without consideration of the syllable transition matrix), whereas the scalar summary includes a wide variety of data describing the mouse’s behavioral comportment (including height, length, speed, position). These MoSeq and scalar behavioral summaries are then submitted to linear classifiers to predict the identity of the drug, drug and dose, or drug class to which each mouse was exposed.
Extended Data Fig. 2 Tuning MoSeq parameters.
a, Scanning the MoSeq kappa parameter (which sets the timescale at which syllables are identified) reveals a value at which the modal syllable length matches the model-free block length identified by changepoints analysis (see Methods). b, The mode of the syllable duration distribution established by MoSeq, given the kappa established in a, matches that for the model-free changepoint distribution. c, Ninety percent of the total frames are explained by 92 behavioral syllables; for the sake of simplicity herein we analyze the top 90 syllables.
Extended Data Fig. 3 Cladogram of syllables with representative illustrations.
A cladogram describing behavioral relationships among syllables was computed using hierarchical clustering performed on the autoregressive matrices describing all syllables (see Methods). Nine general behavioral categories were identified after visual inspection and given natural language names. Illustrations are representative of syllables in each category.
Extended Data Fig. 4 MoSeq outperforms scalar metrics at all-vs-all drug discrimination.
a, Normalized confusion matrices as in Fig. 3a, but computed for all drug/dose combinations. For the shuffled control (bottom row), syllable labels were shuffled on a per-mouse basis to compute a baseline of expected random performance. Heat map indicates classification successes and errors (see Methods for summary definitions). b, Mean precision-recall curves for all drugs and doses, computed for each behavioral summary type. c, The Fukunaga and Olsen method43 was used to estimate the effective dimensionality of both scalar and MoSeq summaries; this analysis demonstrated that that MoSeq has a higher effective dimensionality than scalars (34 versus 26 dimensions), using a threshold value of 0.01 (see Methods).
Extended Data Fig. 5 Adding additional information to behavioral summaries or altering summary dimensionality does not improve performance.
a, Additional information was added to the MoSeq and scalar behavioral summaries used to predict drug identity. For “MoSeq+ +,” the empirical transition matrix derived from the syllable label sequence was calculated, flattened, and concatenated to the syllable usage frequency information. For “Scalars+ +,” histograms of mouse acceleration, the mouse’s heading, the area contained by the mouse’s body contour, the ellipticity of the best-fit ellipse around the mouse’s contour, and the mouse’s width were added to the initial scalar behavioral summary. b, The granularity of the bins used to generate scalar behavioral summaries was systematically varied; bin size did not affect classification performance. c, To ensure that the higher dimensionality of the scalar summaries did not adversely affect performance, behavioral summaries containing scalars were also subjected to PCA to assess the consequences of dimensionality reduction (keeping the number of dimensions required to capture 95 percent of the variance; for scalars this is 33 dimensions); although performance was modestly improved, performance did not equal that observed for MoSeq.
Extended Data Fig. 6 Exploring behavioral similarities elicited by specific drug/dose pairs.
Average cosine distance ±1 standard deviation of mice given the same drug/dose pair (blue) and mice given different drug/dose pairs (red) using either scalar- (top) or MoSeq-based behavioral summaries (bottom). The difference observed between mice given the same drug/dose pair and different drug/dose pairs is uniformly larger when behavior is summarized using MoSeq when compared to scalars. Inset: summary of mean within- and between- class differences and their ratio for either scalar- and MoSeq-based analysis. MoSeq shows larger differences (two-sided paired t-test, p < 0.05, stars indicate statistically significant differences between MoSeq and scalars).
Extended Data Fig. 7 MoSeq captures the behavioral variability of individual mice.
To test whether the cosine distances that separate individual mice within a treatment class reflect individual variability or technical noise, we subsampled the data from each individual mouse and then asked how these sub-samples of each individual mouse compared to each other; observing low variability in these sub-samples would be consistent with each individual mouse expressing a stable set of behavioral syllables within an experiment, and with the within-condition variability observed across mice reflecting differences in individual mouse responses to a given drug and dose. In specific, within-mouse variability of MoSeq was assessed by randomly picking 1000 frames (with replacement) of the 3D imaging data (which for each mouse was constituted of approximately 36,000 frames), identifying the syllable associated by MoSeq with that frame, and then using those syllable labels to compute overall syllable usages; this procedure is roughly equivalent to randomly choosing less than one third of the syllables to quantify the pattern of syllable usage within a mouse. We repeated this procedure 100 times, and by computing cosine distances between each sub-sample within-mouse variability could be assessed. The bootstrapped estimate of individual variability (Resampled Within Mouse) was lower than the treatment-induced variability (Within tTreatment), as measured by the cosine distance between all pairs of mice given the same treatment, and was also lower than the cosine distance between pairs of mice given different treatments (Between Treatment). Thus the observed within-treatment variability reflects stable differences in behavior expressed by individual mice.
Extended Data Fig. 8 MoSeq distinguishes the behavioral effects of drugs independent of effects on mouse movement speed.
a, Similar as Fig. 3a, but classifying drug/dose identity instead of drug identity, across the entire risperidone, haloperidol, clozapine dose-response experiment. Many significant syllables that differentiated drug-treated mice from controls were, by inspection, behaviors like grooming or rearing that do not include significant two-dimensional velocity components (data not shown). b, Syllable usages for all mice and all drug/dose combinations (top), doses which resulted in slow mouse movement speed (middle) or moderate movement speed (bottom). Slow and medium speeds (relative to normal) were identified via a Gaussian Mixture Model (mean centroid speed of saline control mouse = 74 mm/sec; “medium speed” = 54 mm/sec; “slow speed” = 24 mm/sec; see Methods). Significant differential syllable usage for each drug versus control indicated with an asterisk (Kruskal-Wallis and post-hoc Dunn’s two-sided test with permutation, with Benjamini/Hochberg FDR with alpha = 0.05).
Extended Data Fig. 9 High classification performance by MoSeq is supported by a limited set of syllables.
Sparsification reveals the number of syllables required to correctly distinguish each drug, as assessed by F1 scores emerging from linear classifiers trained on subsets of syllables (see Methods).
Supplementary information
Supplementary Information
Supplementary Figs. 1–6, Supplementary Tables 1–4 and Supplementary References.
Rights and permissions
About this article
Cite this article
Wiltschko, A.B., Tsukahara, T., Zeine, A. et al. Revealing the structure of pharmacobehavioral space through motion sequencing. Nat Neurosci 23, 1433–1443 (2020). https://doi.org/10.1038/s41593-020-00706-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-00706-3
This article is cited by
-
Development of a 3D tracking system for multiple marmosets under free-moving conditions
Communications Biology (2024)
-
Spontaneous behaviour is structured by reinforcement without explicit reward
Nature (2023)
-
Neural circuits regulating prosocial behaviors
Neuropsychopharmacology (2023)
-
Automated maternal behavior during early life in rodents (AMBER) pipeline
Scientific Reports (2023)
-
Defensive responses: behaviour, the brain and the body
Nature Reviews Neuroscience (2023)