Abstract
Microglia become progressively activated and seemingly dysfunctional with age, and genetic studies have linked these cells to the pathogenesis of a growing number of neurodegenerative diseases. Here we report a striking buildup of lipid droplets in microglia with aging in mouse and human brains. These cells, which we call ‘lipid-droplet-accumulating microglia’ (LDAM), are defective in phagocytosis, produce high levels of reactive oxygen species and secrete proinflammatory cytokines. RNA-sequencing analysis of LDAM revealed a transcriptional profile driven by innate inflammation that is distinct from previously reported microglial states. An unbiased CRISPR–Cas9 screen identified genetic modifiers of lipid droplet formation; surprisingly, variants of several of these genes, including progranulin (GRN), are causes of autosomal-dominant forms of human neurodegenerative diseases. We therefore propose that LDAM contribute to age-related and genetic forms of neurodegeneration.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Change history
27 July 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41593-020-0682-y
31 January 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41593-020-0595-9
References
Aguzzi, A., Barres, B. A. & Bennett, M. L. Microglia: scapegoat, saboteur, or something else? Science 339, 156–161 (2013).
Mosher, K. I. & Wyss-Coray, T. Microglial dysfunction in brain aging and Alzheimer’s disease. Biochem. Pharmacol. 88, 594–604 (2014).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290.e17 (2017).
Krasemann, S. et al. The TREM2-APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581.e9 (2017).
Li, Q. et al. Developmental heterogeneity of microglia and brain myeloid cells revealed by deep single-cell RNA sequencing. Neuron 101, 207–223.e10 (2019).
Alzheimer, A. Über eine eigenartige Erkrankung der Hirnrinde. Allg. Z. Psychiatr. 64, 146–148 (1907).
Foley, P. Lipids in Alzheimer’s disease: a century-old story. Biochim. Biophys. Acta 1801, 750–753 (2010).
Fowler, S. D., Mayer, E. P. & Greenspan, P. Foam cells and atherogenesis. Ann. NY Acad. Sci. 454, 79–90 (1985).
Bozza, P. T. & Viola, J. P. Lipid droplets in inflammation and cancer. Prostaglandins Leukot. Essent Fatty Acids 82, 243–250 (2010).
den Brok, M. H., Raaijmakers, T. K., Collado-Camps, E. & Adema, G. J. Lipid droplets as immune modulators in myeloid cells. Trends Immunol. 39, 380–392 (2018).
Childs, B. G. et al. Senescent intimal foam cells are deleterious at all stages of atherosclerosis. Science 354, 472–477 (2016).
Castejon, O. J., Castellano, A., Arismendi, G. J. & Medina, Z. The inflammatory reaction in human traumatic oedematous cerebral cortex. J. Submicrosc. Cytol. Pathol. 37, 43–52 (2005).
Lee, S. C., Moore, G. R., Golenwsky, G. & Raine, C. S. Multiple sclerosis: a role for astroglia in active demyelination suggested by class II MHC expression and ultrastructural study. J. Neuropathol. Exp. Neurol. 49, 122–136 (1990).
Liu, L. et al. Glial lipid droplets and ROS induced by mitochondrial defects promote neurodegeneration. Cell 160, 177–190 (2015).
Shimabukuro, M. K. et al. Lipid-laden cells differentially distributed in the aging brain are functionally active and correspond to distinct phenotypes. Sci. Rep. 6, 23795 (2016).
Chang, P. K., Khatchadourian, A., McKinney, R. A. & Maysinger, D. Docosahexaenoic acid (DHA): a modulator of microglia activity and dendritic spine morphology. J. Neuroinflammation 12, 34 (2015).
Khatchadourian, A., Bourque, S. D., Richard, V. R., Titorenko, V. I. & Maysinger, D. Dynamics and regulation of lipid droplet formation in lipopolysaccharide (LPS)-stimulated microglia. Biochim. Biophys. Acta 1821, 607–617 (2012).
Harris, L. A., Skinner, J. R. & Wolins, N. E. Imaging of neutral lipids and neutral lipid associated proteins. Methods Cell. Biol. 116, 213–226 (2013).
Yu, Y., Ramachandran, P. V. & Wang, M. C. Shedding new light on lipid functions with CARS and SRS microscopy. Biochim. Biophys. Acta 1841, 1120–1129 (2014).
Bennett, M. L. et al. New tools for studying microglia in the mouse and human CNS. Proc. Natl Acad. Sci. USA 113, E1738–E1746 (2016).
Pluvinage, J. V. et al. CD22 blockade restores homeostatic microglial phagocytosis in ageing brains. Nature 568, 187–192 (2019).
Holtman, I. R. et al. Induction of a common microglia gene expression signature by aging and neurodegenerative conditions: a co-expression meta-analysis. Acta Neuropathol. Commun. 3, 31 (2015).
Chiu, I. M. et al. A neurodegeneration-specific gene-expression signature of acutely isolated microglia from an amyotrophic lateral sclerosis mouse model. Cell Rep. 4, 385–401 (2013).
Wang, Y. et al. TREM2 lipid sensing sustains the microglial response in an Alzheimer’s disease model. Cell 160, 1061–1071 (2015).
Hammond, T. R. et al. Single-cell RNA sequencing of microglia throughout the mouse lifespan and in the injured brain reveals complex cell-state changes. Immunity 50, 253–271.e6 (2019).
Namatame, I., Tomoda, H., Arai, H., Inoue, K. & Omura, S. Complete inhibition of mouse macrophage-derived foam cell formation by triacsin C. J. Biochem. 125, 319–327 (1999).
Hahn, O. et al. Dietary restriction protects from age-associated DNA methylation and induces epigenetic reprogramming of lipid metabolism. Genome Biol. 18, 56 (2017).
Chen, Z. et al. Lipopolysaccharide-induced microglial activation and neuroprotection against experimental brain injury is independent of hematogenous TLR4. J. Neurosci. 32, 11706–11715 (2012).
Kawabe, K., Takano, K., Moriyama, M. & Nakamura, Y. Lipopolysaccharide-stimulated transglutaminase 2 expression enhances endocytosis activity in the mouse microglial cell line BV-2. Neuroimmunomodulation 22, 243–249 (2015).
Yousef, H. et al. Aged blood impairs hippocampal neural precursor activity and activates microglia via brain endothelial cell VCAM1. Nat. Med. 25, 988–1000 (2019).
Morgens, D. W. et al. Genome-scale measurement of off-target activity using Cas9 toxicity in high-throughput screens. Nat. Commun. 8, 15178 (2017).
Kamkaew, A. et al. BODIPY dyes in photodynamic therapy. Chem. Soc. Rev. 42, 77–88 (2013).
Lee, J. et al. Adaptor protein sorting nexin 17 regulates amyloid precursor protein trafficking and processing in the early endosomes. J. Biol. Chem. 283, 11501–11508 (2008).
Liu, P. et al. S113R mutation in SLC33A1 leads to neurodegeneration and augmented BMP signaling in a mouse model. Dis. Model. Mech. 10, 53–62 (2017).
Huey, E. D. et al. Characteristics of frontotemporal dementia patients with a progranulin mutation. Ann. Neurol. 60, 374–380 (2006).
Tsika, E. et al. Parkinson’s disease-linked mutations in VPS35 induce dopaminergic neurodegeneration. Hum. Mol. Genet. 23, 4621–4638 (2014).
Yin, F. et al. Behavioral deficits and progressive neuropathology in progranulin-deficient mice: a mouse model of frontotemporal dementia. FASEB J. 24, 4639–4647 (2010).
Vieira-de-Abreu, A. et al. Cross-talk between macrophage migration inhibitory factor and eotaxin in allergic eosinophil activation forms leukotriene C(4)-synthesizing lipid bodies. Am. J. Respir. Cell Mol. Biol. 44, 509–516 (2011).
Hu, X., Xu, B. & Ge, W. The role of lipid bodies in the microglial aging process and related diseases. Neurochem. Res. 42, 3140–3148 (2017).
Rambold, A. S., Cohen, S. & Lippincott-Schwartz, J. Fatty acid trafficking in starved cells: regulation by lipid droplet lipolysis, autophagy, and mitochondrial fusion dynamics. Dev. Cell 32, 678–692 (2015).
Ghoshal, N., Dearborn, J. T., Wozniak, D. F. & Cairns, N. J. Core features of frontotemporal dementia recapitulated in progranulin knockout mice. Neurobiol. Dis. 45, 395–408 (2012).
Currie, E., Schulze, A., Zechner, R., Walther, T. C. & Farese, R. V. Jr. Cellular fatty acid metabolism and cancer. Cell Metab. 18, 153–161 (2013).
Burke, A. C. & Huff, M. W. ATP-citrate lyase: genetics, molecular biology and therapeutic target for dyslipidemia. Curr. Opin. Lipidol. 28, 193–200 (2017).
Cantuti-Castelvetri, L. et al. Defective cholesterol clearance limits remyelination in the aged central nervous system. Science 359, 684–688 (2018).
Lee, S. J., Zhang, J., Choi, A. M. & Kim, H. P. Mitochondrial dysfunction induces formation of lipid droplets as a generalized response to stress. Oxid. Med. Cell Longev. 2013, 327167 (2013).
Nadjar, A. Role of metabolic programming in the modulation of microglia phagocytosis by lipids. Prostaglandins Leukot. Essent. Fatty Acids 135, 63–73 (2018).
Chinetti-Gbaguidi, G. et al. Human atherosclerotic plaque alternative macrophages display low cholesterol handling but high phagocytosis because of distinct activities of the PPARγ and LXRα pathways. Circ. Res. 108, 985–995 (2011).
Chandak, P. G. et al. Efficient phagocytosis requires triacylglycerol hydrolysis by adipose triglyceride lipase. J. Biol. Chem. 285, 20192–20201 (2010).
Jaitin, D. A. et al. Lipid-associated macrophages control metabolic homeostasis in a Trem2-dependent manner. Cell 178, 686–698.e14 (2019).
Nguyen, A. D et al. Progranulin in the hematopoietic compartment protects mice from atherosclerosis. Atherosclerosis 277, 145–154 (2018).
Evers, B. M et al. Lipidomic and transcriptomic basis of lysosomal dysfunction in progranulin deficiency . Cell Rep. 20, 2565–2574 (2017).
Gil-Perotin, S., Alvarez-Buylla, A. & Garcia-Verdugo, J. M. Identification and characterization of neural progenitor cells in the adult mammalian brain. Adv. Anat. Embryol. Cell Biol. 203, 1–101 (2009).
Bligh, E. G. & Dyer, W. J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B Meth. 57, 289–300 (1995).
Bohlen, C. J. et al. Diverse requirements for microglial survival, specification, and function revealed by defined-medium cultures. Neuron 94, 759–773.e8 (2017).
Deans, R. M. et al. Parallel shRNA and CRISPR–Cas9 screens enable antiviral drug target identification. Nat. Chem. Biol. 12, 361–366 (2016).
Acknowledgements
The authors thank the members of the Wyss-Coray laboratory for feedback and support throughout the study, W. Stoiber (University of Salzburg, Austria) and the Division of Optical and Electron Microscopy, University of Salzburg, Austria, for excellent assistance with EM images, and D. Channappa from the Department of Neurology and E. Plowey from the Department of Pathology, Stanford University, for providing human postmortem brain samples. CARS imaging was performed at the Microscopy Core Facility of the Institute of Molecular Biosciences, University of Graz, Austria. They also thank A. Enejder, the Heilshorn Biomaterials Group, Stanford University, for discussion and for reviewing the manuscript. This work was supported by the FWF Hertha-Firnberg Postdoctoral program no. T736-B24 (to J.M.), the PMU-FFF E-16/23/117-FEA (to T.K.F), the NIH grant R37 GM058867 (to C.R.B), the Stanford Neuroscience Institute Brain Rejuvenation Project Award and NIH Director’s New Innovator Award (1DP2HD084069-01) to M.C.B., the Department of Veterans Affairs (to T.W.-C.), the National Institute on Aging (DP1-AG053015 to T.W.-C.), the NOMIS Foundation (to T.W.-C.), the Glenn Foundation for Medical Research (to T.W.-C.), the Cure Alzheimer’s Fund (to T.W.-C.) and the Nan Fung Life Sciences Aging Research Fund (to T.W.-C.).
Author information
Authors and Affiliations
Contributions
J.M. and T.W.-C. conceptualized and designed the study, analyzed and interpreted data, and wrote the manuscript. J.M. and S.E.L. designed the figures. J.M., T.I., M.Z., S.E.L. and J.V.P. acquired the data. J.M. performed the electron microscopy experiments. J.M. and M.Z. performed histology and organotypic slice culture experiments. J.M., T.I. and M.Z. performed the cell culture experiments. J.M., T.I. and S.E.L performed the RNA-seq experiments. J.V.P., M.Z. and J.M. performed the stereotactic procedures. V.M. conducted the in vivo LPS injections and provided Grn−/− mouse brain sections. J.M., M.S.H. and D.W.M. generated and analyzed the CRISPR–Cas9 screen data. J.M. and B.L. analyzed the RNA-seq data. J.T., T.K.F. and O.H. performed the mass spectrometry experiments. J.M. and H.W. performed the CARS imaging. J.K. and C.R.B. designed and produced the methylated BODIPY derivatives. M.C.B and L.A. reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
T.W.-C., J.M., C.R.B. and M.S.H. are co-inventors on a patent application related to the work published in this paper. All other authors have no competing interests.
Additional information
Peer review information Nature Neuroscience thanks Staci Bilbo, Serge Rivest, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Lipid droplet accumulating microglia are abundant in the hippocampus but rare in other brain regions of aged mice.
a–d, Representative confocal images of the cortex (a), thalamus (b), corpus callosum (c) and hippocampal dentate gyrus (d) from 20-month old male mice stained for BODIPY+ (lipid droplets) and Iba1+ (microglia). Scale bar: 20 μm. Arrows point towards BODIPY+ lipid droplets. e, Quantification of BODIPY+/Iba1+ cells. n = 4 mice per group. One-way ANOVA followed by Tukey’s post hoc test. Error bars represent mean ± SD. ***P< 0.001.
Extended Data Fig. 2 LDAM have a unique transcriptional signature that minimally overlaps with published gene expression profiles of microglia in aging and neurodegeneration.
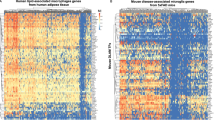
a,b, IPA pathway analysis of genes that are significantly upregulated (a) or downregulated (b) in LD-hi microglia in aging. Analysis based on top 100 down- and up-regulated genes (Fisher’s exact test, Benjamini-Hochberg FDR). c-g, Expression plots comparing RNA-Seq data of LDAM (see Fig. 2) with published RNA-Seq data of microglia in aging (c), AD (d), ALS (e), disease-associated microglia (DAM) (f) and neurodegenerative microglia (MGnD) (g). Data are expressed as signed fdr, i.e the product of log2 FC and log10 fdr. h, Paired dot plot showing FPKM values of LD-lo and LD-hi microglia for ApoE (paired Student’s t-test; P= 0.423). Dotted lines connect LD-lo and LD-hi microglia sorted from the same samples. i, Heatmap showing expression changes of LDAM genes (genes differentially expressed in LD-hi microglia in aging) in LD-hi microglia from GRN-/- mice, from LPS treated mice, and in microglia clusters revealed by Li et al. (2019) and Hammond et al. (2019)15,16. Sample size in a,b,h: n = 3 samples per group. Each sample is a pool of microglia from the hippocampi of 3 mice. LD, lipid droplet.
Extended Data Fig. 3 LPS treatment induces lipid droplet formation in microglia and in BV2 cells.
a,b, 3-month-old male mice were given intraperitoneal (i.p.) injections of LPS (1 mg/kg BW) for four days. Representative confocal images of BODIPY+ and Tmem119+ in the hippocampus (a) and of BODIPY and Iba1 staining in the cortex, corpus callosum, and thalamus (b). c-e, Lipidome profiling of lipid droplets from LPS-treated BV2 cells, primary microglia, and liver tissue. c, Pie charts showing that the lipid composition of lipid droplets from young and aged microglia is highly similar, but differs between young and aged liver tissue. d,e, Distribution of MAG chain lengths (d) and TAG saturation levels (e) of lipid droplets isolated from LPS-treated BV2 cells and from microglia and liver tissue from aged mice. young= 5-month-old male mice, old= 20-month-old male mice; n = 4 mice per group. Data in a-b were replicated in at least two independent experiments. Error bars represent mean ± s.e.m. Scale bars, 20 μm.
Extended Data Fig. 4 Aged plasma induces lipid droplet formation in BV2 cells.
a, Representative micrographs of BODIPY+ staining and of phagocytosis of of pHrodo red Zymosan in BV2 cells treated with 5% plasma from young (3-months) and aged (18-months) mice for 12 hours. Scale bars, 5 μm. b, Quantification of BODIPY+ staining in BV2 cells treated with young and aged plasma. c,d, Quantification of Zymosan uptake in BV2 cells treated with young and aged plasma (c), and in aged plasma treated BODIPY-low and BODIPY-high cells (d). Statistical tests: two-sided Student’s t-test. Error bars represent mean ± SD. *P< 0.05, ***P< 0.001.
Extended Data Fig. 5 Lipid droplet containing microglia in the cortex, corpus callosum, and thalamus of GRN-/- mice.
a-c, Representative confocal images of BODIPY+ (lipid droplets) and Iba1+ (microglia) in the cortex (a), corpus callosum (b), (c) and thalamus from 9-month-old male GRN-/- mice. BODIPY+/Iba1+ cells were frequently found in the thalamus and were detected to a lesser extent in cortex and corpus callosum. Data were replicated in at least three independent experiments.
Extended Data Fig. 6 Expression changes of LDAM genes in lipid droplet-rich microglia from normal aging, GRN-/- and LPS-treated mice.
a, Heatmap showing expression changes of LDAM genes (genes differentially expressed in LD-hi microglia in aging; 692 genes) in LD-hi microglia from GRN-/- mice and from LPS treated mice.
Extended Data Fig. 7 LDAM show signs of metabolic alterations.
a, Paired dot plot showing FPKM values of LD-lo and LD-hi microglia for ACLY (data obtained from RNA-Seq analysis, see Fig. 2). Dotted lines connect LD-lo and LD-hi microglia sorted from the same samples. P=b, NAD colorimetric assay showing the NAD+/NADH ratio of primary hippocampal microglia from 3-month old mice (young) and of LD-lo and LD-hi primary microglia from 20-month old male mice. Experiments were performed two times in technical triplicates. n=3 mice per group per experiment. Statistical tests: paired two-sided Student’s t-test (a) one-way ANOVA (b) followed by Tukey’s post hoc test. Horizontal lines in the box plots indicate medians, box limits indicate first and third quantiles, and vertical whisker lines indicate minimum and maximum values. *P< 0.05, ***P< 0.001.
Supplementary information
Supplementary Tables
Supplementary Tables 1 and 2.
Rights and permissions
About this article
Cite this article
Marschallinger, J., Iram, T., Zardeneta, M. et al. Lipid-droplet-accumulating microglia represent a dysfunctional and proinflammatory state in the aging brain. Nat Neurosci 23, 194–208 (2020). https://doi.org/10.1038/s41593-019-0566-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0566-1
This article is cited by
-
Emerging role of senescent microglia in brain aging-related neurodegenerative diseases
Translational Neurodegeneration (2024)
-
Progranulin haploinsufficiency mediates cytoplasmic TDP-43 aggregation with lysosomal abnormalities in human microglia
Journal of Neuroinflammation (2024)
-
Changes in lipid metabolism track with the progression of neurofibrillary pathology in tauopathies
Journal of Neuroinflammation (2024)
-
Neuroimmunology and ageing – the state of the art
Immunity & Ageing (2024)
-
Nanotechnology for microglial targeting and inhibition of neuroinflammation underlying Alzheimer’s pathology
Translational Neurodegeneration (2024)