Abstract
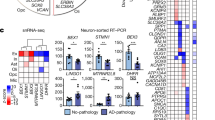
There is currently little information available about how individual cell types contribute to Alzheimer’s disease. Here we applied single-nucleus RNA sequencing to entorhinal cortex samples from control and Alzheimer’s disease brains (n = 6 per group), yielding a total of 13,214 high-quality nuclei. We detail cell-type-specific gene expression patterns, unveiling how transcriptional changes in specific cell subpopulations are associated with Alzheimer’s disease. We report that the Alzheimer’s disease risk gene APOE is specifically repressed in Alzheimer’s disease oligodendrocyte progenitor cells and astrocyte subpopulations and upregulated in an Alzheimer’s disease-specific microglial subopulation. Integrating transcription factor regulatory modules with Alzheimer’s disease risk loci revealed drivers of cell-type-specific state transitions towards Alzheimer’s disease. For example, transcription factor EB, a master regulator of lysosomal function, regulates multiple disease genes in a specific Alzheimer’s disease astrocyte subpopulation. These results provide insights into the coordinated control of Alzheimer’s disease risk genes and their cell-type-specific contribution to disease susceptibility. These results are available at http://adsn.ddnetbio.com.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All single-cell RNA sequencing data are available from the Gene Expression Omnibus (GEO) under the accession number GSE138852. Data can also be visualized via the interactive web application at adsn.ddnetbio.com. Single-cell gene expression data and metadata can also be downloaded directly via adsn.ddnetbio.com.
Code availability
Code is available from the authors by reasonable request.
References
Huang, K.-L. et al. A common haplotype lowers PU.1 expression in myeloid cells and delays onset of Alzheimer’s disease. Nat. Neurosci. 20, 1052–1061 (2017).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Zhang, B. et al. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 153, 707–720 (2013).
Füger, P. et al. Microglia turnover with aging and in an Alzheimer’s model via long-term in vivo single-cell imaging. Nat. Neurosci. 20, 1371–1376 (2017).
Whitehouse, P. J. et al. Alzheimer’s disease and senile dementia: loss of neurons in the basal forebrain. Science 215, 1237–1239 (1982).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Mathys, H. Single-cell transcriptomic analysis of Alzheimer’s disease.Nature 570, 332–337 (2019).
Wilhelmsson, U. et al. Injury leads to the appearance of cells with characteristics of both microglia and astrocytes in mouse and human brain. Cereb. Cortex 27, 3360–3377 (2017).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Darmanis, S. et al. A survey of human brain transcriptome diversity at the single cell level. Proc. Natl Acad. Sci. USA 112, 7285–7290 (2015).
Miceli, F., et al. KCNQ3-Related Disorders. in Gene Reviews (ed. Adam, M. P.) https://www.ncbi.nlm.nih.gov/books/NBK201978/ (Univ. Washington, 2014).
Ebermann, I. et al. GPR98 mutations cause Usher syndrome type 2 in males. J. Med. Genet. 46, 277–280 (2009).
Grubman, A., Choo, X. Y., Chew, G., Ouyang, J. F. & Sun, G. Mouse and human microglial phenotypes in Alzheimer’s disease are controlled by amyloid plaque phagocytosis through Hif1α. Preprint at biorXiv https://www.biorxiv.org/content/10.1101/639054v1 (2019).
Veereshwarayya, V., Kumar, P., Rosen, K. M., Mestril, R. & Querfurth, H. W. Differential effects of mitochondrial heat shock protein 60 and related molecular chaperones to prevent intracellular β-amyloid-induced inhibition of complex IV and limit apoptosis. J. Biol. Chem. 281, 29468–29478 (2006).
De Strooper, B. & Karran, E. The cellular phase of Alzheimer’s disease. Cell 164, 603–615 (2016).
Xie, H. et al. Rapid cell death is preceded by amyloid plaque-mediated oxidative stress. Proc. Natl Acad. Sci. USA 110, 7904–7909 (2013).
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017).
Krasemann, S. et al. The TREM2-APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581.e9 (2017).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290.e17 (2017).
De Rossi, P. et al. Predominant expression of Alzheimer’s disease-associated BIN1 in mature oligodendrocytes and localization to white matter tracts. Mol. Neurodegener. 11, 59 (2016).
Savvaki, M. et al. The expression of TAG-1 in glial cells is sufficient for the formation of the juxtaparanodal complex and the phenotypic rescue of tag-1 homozygous mutants in the CNS. J. Neurosci. 30, 13943–13954 (2010).
Bartzokis, G. Age-related myelin breakdown: a developmental model of cognitive decline and Alzheimer’s disease. Neurobiol. Aging 25, 5–18 (2004).
Behrendt, G. et al. Dynamic changes in myelin aberrations and oligodendrocyte generation in chronic amyloidosis in mice and men. Glia 61, 273–286 (2013).
Lake, B. B. et al. Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590 (2016).
John Lin, C.-C. et al. Identification of diverse astrocyte populations and their malignant analogs. Nat. Neurosci. 20, 396–405 (2017).
Mi, S. et al. LINGO-1 negatively regulates myelination by oligodendrocytes. Nat. Neurosci. 8, 745–751 (2005).
Santoro, M. et al. Expression profile of long non-coding RNAs in serum of patients with multiple sclerosis. J. Mol. Neurosci. 59, 18–23 (2016).
Fallin, M. D. et al. Bipolar I disorder and schizophrenia: a 440–single-nucleotide polymorphism screen of 64 candidate genes among Ashkenazi Jewish case-parent trios. Am. J. Hum. Genet. 77, 918–936 (2005).
Hall, C. N., Klein-Flügge, M. C., Howarth, C. & Attwell, D. Oxidative phosphorylation, not glycolysis, powers presynaptic and postsynaptic mechanisms underlying brain information processing. J. Neurosci. 32, 8940–8951 (2012).
Romito-DiGiacomo, R. R., Menegay, H., Cicero, S. A. & Herrup, K. Effects of Alzheimer’s disease on different cortical layers: the role of intrinsic differences in Abeta susceptibility. J. Neurosci. 27, 8496–8504 (2007).
Bis, J. C. et al. Whole exome sequencing study identifies novel rare and common Alzheimer’s-Associated variants involved in immune response and transcriptional regulation. Mol. Psychiatry https://doi.org/10.1038/s41380-018-0112-7 (2018).
Sims, R. et al. Rare coding variants in PLCG2, ABI3, and TREM2 implicate microglial-mediated innate immunity in Alzheimer’s disease. Nat. Genet. 49, 1373–1384 (2017).
Kajiho, H. et al. Characterization of RIN3 as a guanine nucleotide exchange factor for the Rab5 subfamily GTPase Rab31. J. Biol. Chem. 286, 24364–24373 (2011).
He, Z.-Y., Li, L., Wang, Y.-Z., Liu, X. & Yuan, L.-Y. Associations between thromboxane A synthase 1 gene polymorphisms and the risk of ischemic stroke in a Chinese Han population.Neural Regen. Res. 13, 463 (2018).
Diener, H. C. et al. European Stroke Prevention Study 2. Dipyridamole and acetylsalicylic acid in the secondary prevention of stroke. J. Neurol. Sci. 143, 1–13 (1996).
Paris, D. et al. Inhibition of Alzheimer’s beta-amyloid induced vasoactivity and proinflammatory response in microglia by a cGMP-dependent mechanism. Exp. Neurol. 157, 211–221 (1999).
Alzheimer’s Association. 2018 Alzheimer’s disease facts and figures. Alzheimers Dement. 14, 367–429 (2018).
Raj, T. et al. Integrative transcriptome analyses of the aging brain implicate altered splicing in Alzheimer’s disease susceptibility. Nat. Genet. 50, 1584–1592 (2018).
Tan, M. G. et al. Genome wide profiling of altered gene expression in the neocortex of Alzheimer’s disease. J. Neurosci. Res. 88, 1157–1169 (2010).
Blalock, E. M., Buechel, H. M., Popovic, J., Geddes, J. W. & Landfield, P. W. Microarray analyses of laser-captured hippocampus reveal distinct gray and white matter signatures associated with incipient Alzheimer’s disease. J. Chem. Neuroanat. 42, 118–126 (2011).
Lin, Y.-T. et al. APOE4 causes widespread molecular and cellular alterations associated with Alzheimer’s disease phenotypes in human iPSC-derived brain cell types. Neuron 98, 1294 (2018).
Bartzokis, G. et al. Apolipoprotein E genotype and age-related myelin breakdown in healthy individuals: implications for cognitive decline and dementia. Arch. Gen. Psychiatry 63, 63–72 (2006).
Chapuis, J. et al. Increased expression of BIN1 mediates Alzheimer genetic risk by modulating tau pathology. Mol. Psychiatry 18, 1225–1234 (2013).
Yu, L. et al. Association of brain DNA methylation in SORL1, ABCA7, HLA-DRB5, SLC24A4, and BIN1 with pathological diagnosis of Alzheimer disease. JAMA Neurol. 72, 15–24 (2015).
Lambert, J.-C. et al. Genome-wide haplotype association study identifies the FRMD4A gene as a risk locus for Alzheimer’s disease. Mol. Psychiatry 18, 461–470 (2013).
Hokama, M. et al. Altered expression of diabetes-related genes in Alzheimer’s disease brains: the Hisayama study. Cereb. Cortex 24, 2476–2488 (2014).
Shijo, M. et al. Association of adipocyte enhancer-binding protein 1 with Alzheimer’s disease pathology in human hippocampi. Brain Pathol. 28, 58–71 (2018).
Maynard, M. A. et al. Human HIF-3α4 is a dominant-negative regulator of HIF-1 and is down-regulated in renal cell carcinoma. FASEB J. 19, 1396–1406 (2005).
Martini-Stoica, H. et al. TFEB enhances astroglial uptake of extracellular tau species and reduces tau spreading. J. Exp. Med. 215, 2355–2377 (2018).
de Toledo-Morrell, L., Goncharova, I., Dickerson, B., Wilson, R. S. & Bennett, D. A. From healthy aging to early Alzheimer’s disease: in vivo detection of entorhinal cortex atrophy. Ann. N. Y. Acad. Sci. 911, 240–253 (2000).
McKenzie, A. T. et al. Brain cell type specific gene expression and co-expression network architectures. Sci. Rep. 8, 8868 (2018).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Zhu, Y., Wang, L., Yin, Y. & Yang, E. Systematic analysis of gene expression patterns associated with postmortem interval in human tissues. Sci. Rep. 7, 5435 (2017).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Lummertz da Rocha, E. et al. Reconstruction of complex single-cell trajectories using CellRouter. Nat. Commun. 9, 892 (2018).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Lambert, S. A. et al. The human transcription factors. Cell 175, 598–599 (2018).
Kolde, R. Pheatmap: pretty heatmaps. R package version 61, 926 (2012).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis. (Springer: 2016)..
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Buniello, A. et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47, D1005–D1012 (2019).
Skene, N. G. & Grant, S. G. N. Identification of vulnerable cell types in major brain disorders using single cell transcriptomes and expression weighted cell type enrichment. Front. Neurosci. 10, 16 (2016).
McKenzie, A. T., Katsyv, I., Song, W.-M., Wang, M. & Zhang, B. DGCA: a comprehensive R package for differential gene correlation analysis. BMC Syst. Biol. 10, 106 (2016).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Li, H. et al. Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nat. Genet. 49, 708–718 (2017).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Conway, J. R., Lex, A. & Gehlenborg, N. UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics 33, 2938–2940 (2017).
Acknowledgements
The authors acknowledge Flowcore and Micromon, Monash University, for the provision of instrumentation, training, and technical support. Tissues were received from the Victorian Brain Bank, supported by The Florey Institute of Neuroscience and Mental Health, The Alfred and the Victorian Institute of Forensic Medicine, and funded in part by Parkinson’s Victoria and MND Victoria. The Australian Regenerative Medicine Institute is supported by grants from the State Government of Victoria and the Australian Government. This work was supported by a Dementia Australia Research Foundation Grant to A.G., J.M.P., and E.P., a Monash Network of Excellence grant to J.M.P., E.P., and O.J.L.R., and a Yulgilbar Foundation grant to A.G. and G.S. A.G. was supported by a National Health and Medical Research Council (NHMRC) and Australian Research Council (ARC) Dementia Research Development Fellowship (GNT1097461). J.M.P. was supported by a Sylvia-Charles Viertel Fellowship. O.J.L.R. and J.F.O. were supported by a Singapore National Research Foundation Competitive Research Programme (NRF-CRP20-2017-0002). S.B. was supported by a National Health and Medical Research Council (NHMRC) and Australian Research Council (ARC) Dementia Research Development Fellowship (GNT1111206). This work was supported by a Sylvia and Charles Viertel Senior Medical Research Fellowship to R.L. and and a Howard Hughes Medical Institute International Research Scholarship to R.L. We acknowledge S. Freytag (University of Western Australia) for her advice regarding disentangling individuals from mixed-individual libraries.
Author information
Authors and Affiliations
Contributions
A.G. and J.M.P. conceived the study and designed experiments, and together with E.P. and O.J.L.R., designed the bioinformatics analyses. A.G., G.S., and X.Y.C. performed nuclei isolation and fluorescence-activated cell sorting (FACS). C.M. performed the pathological assessment of human control and Alzheimer’s disease cases. G.C. and E.P. performed GWAS integration, and CellRouter and network analyses. J.F.O. and O.J.L.R. performed cell-type and cell-subcluster identification and performed differential gene expression and GSEA analyses. J.F.O. and O.J.L.R. developed the shiny web interface. J.P., R.S., S.B., D.V.L., D.P., and R.L. worked up the protocol for single-nuclei sequencing from the human brain. A.G., G.C., J.F.O., O.J.L.R., E.P., and J.M.P. wrote the manuscript. All authors approved of, and contributed to, the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
O.J.L.R. and J.M.P. are co-inventors of the patent (WO/2017/106932) and are co-founders, shareholders, and directors of Mogrify Ltd., a cell therapy company. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended data Fig. 1 Gating strategy for FACS isolation of single nuclei.
Nuclei isolated using the EZ-Prep kit from entorhinal cortex of AD and control patients were FACS sorted on BD Influx using 70 μm nozzle, 21–22 psi. Single DAPI+ events were considered nuclei.
Extended data Fig. 2 Single nuclei metadata and analysis of hybrid cells.
a, cell proportion plots by library for eight libraries showing exclusion of two libraries due to high neuronal count and comparison of cell type proportions recovered in this study compared to cell proportions in other single nuclei studies24 and52. Cell proportions are shown by sequencing library, where this information is available. b, Frequency histogram of number of Unique Molecular Identifiers (UMIs) across all 8 cell type groups. c, Frequency histogram of number of detected genes across all 8 cell type groups. d, Barplot showing number of top two identified cell types in hybrid cell types in AD and control libraries as proportion. e, UpSetR69 plot showing the BRETIGEA number of distinct and overlapping cell type markers across the six major cell types - microglia, astrocyte, neuron, oligodendrocyte, OPC, and endothelial cells.
Extended data Fig. 3 Comparison of differentially expressed genes (DEGs) within microglia, OPCs and oligodendrocytes with Mathys et al. 2019.
a, d, g, Venn diagram showing overlap of DEGs detected in Mathys et al. (n = 24 AD-pathology vs n = 24 no-pathology individuals) and those identified in the present study (n = 6 AD vs n = 6 control individuals) in microglia (a), OPCs (d) and oligodendrocytes (g), and the table shows the number of and concordance of dysregulation in AD of genes within the overlap of DEGs in both studies. Hypergeometric test was used to test for significance of overlap. b, e, h, List of concordant and discordant overlapping DEGs in microglia (b), OPCs (e) and oligodendrocytes (h). c, f, i, Significant gene ontology (GO) terms for each gene set comparison in microglia (c), OPCs (f) and oligodendrocytes (i). Multiple testing correction was done using the Benjamini–Hochberg method. Enrichment analysis was performed using hypergeometric test.
Extended data Fig. 4 Comparison of differentially expressed genes (DEGs) within astrocytes, excitatory neurons and inhibitory neurons with Mathys et al. 2019.
a, d, g, Venn diagram showing overlap of DEGs detected in Mathys et al. (n = 24 AD-pathology vs n = 24 no-pathology individuals) and those identified in the present study (n = 6 AD vs n = 6 control individuals) in astrocytes (a), excitatory neurons (d), and inhibitory neurons (g), and the table shows the number of and concordance of dysregulation in AD of genes within the overlap of DEGs in both studies. Hypergeometric test was used to test for significance of overlap. b, e, h, List of concordant and discordant overlapping DEGs in astrocytes (b), excitatory neurons (e), and inhibitory neurons (h). c, f, i, Significant gene ontology (GO) terms for each gene set comparison in astrocytes (c), excitatory neurons (f), and inhibitory neurons (i). Multiple testing correction was done using the Benjamini–Hochberg method. Enrichment analysis was performed using hypergeometric test.
Extended data Fig. 5 Analysis of sources of variation.
a, Box plot (center line: median, box limits: upper and lower quartiles, whiskers: 1.5x interquartile range, points outside whiskers: outliers) showing percentage variance explained by each covariate for genes that are differentially expressed between AD and Control groups in any of the identified cell types. b, Proportion of AD-vs-Control differentially expressed genes (DEGs) that are also individual-associated genes and genes that are only DE between AD and Control. c, Proportion of AD-vs-Control DEGs that are also sex-associated genes and genes that are only DE between AD and Control. The number of DEGs are given in parenthesis.
Extended data Fig. 6 Cell-type-specific and shared AD related changes.
a–i, Bubble plot showing the top 15 differentially expressed genes (DEGs) within the DEG1-DEG9 groups (see Fig. 1f), where the colour represents the relative gene expression and the bubble size is the proportion of cells in the group expressing the gene.
Extended data Fig. 7 AD related changes in excitatory and inhibitory neurons.
Scatter plot showing the relationship between cell-type-specific and common differentially expressed genes (DEGs) in control and AD, excitatory and inhibitory neurons. Cutoff for significance is abs(log fold change) > 0.5 and false discovery rate (FDR) < 0.01.
Extended data Fig. 8 Single nuclei sequencing of human AD and control entorhinal cortex reveals homeostatic, AD-specific and shared ontological cell subclusters.
a,e,i, Uniform Manifold Approximation and Projection (UMAP) visualization of subclusters of microglia (a), OPCs (e) and endothelial cells (i) showing b, f, j, the composition of cells in subclusters by disease state, c, g, k, hierarchical clustering and heatmap colored by single-cell gene expression of subcluster-specific genes (top eight genes were shown per cluster). d, h, l, Gene Set Enrichment Analysis (GSEA) results of subcluster specific genes coloured by normalised gene set enrichment scores for the gene ontologies shown in each cell subcluster.
Extended data Fig. 9 Comparison of signatures across subclusters.
Comparison of astrocyte signatures to ai;17 aii25. b, Comparison of neuronal subcluster signatures to24(Ex1–8, In1–8). c, Functional annotation of gene regulatory networks (GRNs) controlled by TFs in Fig. 5a. GRNs are shown in Extended data Fig. 10, ETS2 (n = 9 genes), GTF2IRD1 (n = 339 genes), HIF3A-neuron (n = 112 genes), HIF3A-OPC (n = 96 genes), KDM5B (n = 44 genes), MAX (n = 273 genes), NKX6–2 (n = 332 genes), RELA (n = 224 genes), SOX10 (n = 114 genes), and ZKSCAN1 (n = 37 genes). Multiple correction testing was done using the Benjamini–Hochberg method. Enrichment analysis was performed using the hypergeometric test in R package GOstats.
Extended data Fig. 10 Gene regulatory network analysis predicts transcription factors regulating GWAS genes for conversion of control to AD subcluster signatures.
Trajectories of the top transcription factors (TFs): NKX6–2, ETS2, ZKSCAN1, HIF3A, GTF2IRD1, RELA, SOX10, KDM5B and MAX, with downstream GWAS gene targets. For each cell type, the reported TFs-driven gene regulatory networks include GWAS gene hits, which are direct target genes of the TF and have the highest average log fold change in gene expression between the source subcluster/s and target subcluster/s. Each gene is colored according to its average log fold change between the source subcluster/s and target subcluster/s. Only significantly differentially expressed GWAS gene hits are reported (false discovery rate < 0.05).
Supplementary information
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 7
Statistical Source Data
Source Data Extended Data Fig. 8
Statistical Source Data
Source Data Extended Data Fig. 9
Statistical Source Data
Rights and permissions
About this article
Cite this article
Grubman, A., Chew, G., Ouyang, J.F. et al. A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat Neurosci 22, 2087–2097 (2019). https://doi.org/10.1038/s41593-019-0539-4
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0539-4
This article is cited by
-
Navigating the metabolic maze: anomalies in fatty acid and cholesterol processes in Alzheimer’s astrocytes
Alzheimer's Research & Therapy (2024)
-
Single cell transcriptome analysis of the THY-Tau22 mouse model of Alzheimer’s disease reveals sex-dependent dysregulations
Cell Death Discovery (2024)
-
A phenotypic screening platform for identifying chemical modulators of astrocyte reactivity
Nature Neuroscience (2024)
-
Xenografted human microglia display diverse transcriptomic states in response to Alzheimer’s disease-related amyloid-β pathology
Nature Neuroscience (2024)
-
Microglia regulation of central nervous system myelin health and regeneration
Nature Reviews Immunology (2024)