Abstract
Although genetics highlights the role of microglia in Alzheimer’s disease, one-third of putative Alzheimer’s disease risk genes lack adequate mouse orthologs. Here we successfully engraft human microglia derived from embryonic stem cells in the mouse brain. The cells recapitulate transcriptionally human primary microglia ex vivo and show expression of human-specific Alzheimer’s disease risk genes. Oligomeric amyloid-β induces a divergent response in human versus mouse microglia. This model can be used to study the role of microglia in neurological diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
References
Bohlen, C. J. et al. Diverse requirements for microglial survival, specification, and function revealed by defined-medium cultures. Neuron 94, 759–773.e8 (2017).
De Strooper, B. & Karran, E. The cellular phase of alzheimer’s disease. Cell 164, 603–615 (2016).
Zerbino, D. R. et al. Ensembl 2018. Nucl. Acids Res. 46, D754–D761 (2018).
Jansen, I. E. & et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019).
Claes, C. & et al. Human stem cell–derived monocytes and microglia-like cells reveal impaired amyloid plaque clearance upon heterozygous or homozygous loss of TREM2. Alzheimers Dement. 15, 453–464 (2019).
Rathinam, C. et al. Efficient differentiation and function of human macrophages in humanized CSF-1 mice. Blood 118, 3119–3128 (2011).
Hagemeyer, N. et al. Microglia contribute to normal myelinogenesis and to oligodendrocyte progenitor maintenance during adulthood. Acta Neuropathol. 134, 441–458 (2017).
Davis, B. M., Salinas-Navarro, M., Cordeiro, M. F., Moons, L. & Groef, L. De. Characterizing microglia activation: a spatial statistics approach to maximize information extraction. Sci. Rep. 7, 1–12 (2017).
Jordão, M. J. C. et al. Single-cell profiling identifies myeloid cell subsets with distinct fates during neuroinflammation. Science 363, eaat7554 (2019).
Sala Frigerio, C. et al. The major risk factors for Alzheimer’s disease: age, sex, and genes modulate the microglia response to Aβ plaques. Cell Rep. 27, 1293–1306.e6 (2019).
Friedman, B. A. et al. Diverse brain myeloid expression profiles reveal distinct microglial activation states and aspects of alzheimer’s disease not evident in mouse models. Cell Rep. 22, 832–847 (2018).
Brouillette, J. et al. Neurotoxicity and memory deficits induced by soluble low-molecular-weight amyloid-β1-42 oligomers are revealed in vivo by using a novel animal model. J. Neurosci. 32, 7852–7861 (2012).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of alzheimer’s disease. Cell 169, 1276–1290.e17 (2017).
Marsh, S. E. et al. The adaptive immune system restrains Alzheimer’s disease pathogenesis by modulating microglial function. Proc. Natl. Acad. Sci. USA 113, E1316–E1325 (2016).
Shay, T. et al. Conservation and divergence in the transcriptional programs of the human and mouse immune systems. Proc. Natl. Acad. Sci. USA 110, 2946–2951 (2013).
Butovsky, O. et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat. Neurosci. 17, 131–143 (2014).
Bhaskar, K. et al. Regulation of tau pathology by the microglial fractalkine receptor. Neuron 68, 19–31 (2010).
Deshmane, S. L., Kremlev, S., Amini, S. & Sawaya, B. E. Monocyte chemoattractant protein-1 (MCP-1): an overview. J. Interferon Cytokine Res. 29, 313–326 (2009).
Abud, E. M. et al. iPSC-derived human microglia-like cells to study neurological diseases. Neuron 94, 278–293.e9 (2017).
Espuny-Camacho, I. et al. Hallmarks of Alzheimer’s disease in stem-cell-derived human neurons transplanted into mouse brain. Neuron 93, 1066–1081.e8 (2017).
Kuperstein, I. et al. Neurotoxicity of Alzheimer’s disease Aβ peptides is induced by small changes in the Aβ42 to Aβ40 ratio. EMBO J. 29, 3408–3420 (2010).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinform. 10, 1–7 (2009).
Acknowledgements
Work in the laboratory of B.D.S. was supported by the European Union (grant no. ERC-834682 CELLPHASE_AD), the Fonds voor Wetenschappelijk Onderzoek, KU Leuven, Flanders Institute for Biotechnology, UK Dementia Research Institute (Medical Research Council, Alzheimer’s Research UK and Alzheimer’s Society), a Methusalem grant from KU Leuven and the Flemish Government, Vlaams Initiatief voor Netwerken voor Dementie Onderzoek (Strategic Basic Research Grant no. 135043), the ‘Geneeskundige Stichting Koningin Elisabeth’, Opening the Future campaign of the Leuven Universitair Fonds, the Belgian Alzheimer Research Foundation and the Alzheimer’s Association USA. B.D.S. is holder of the Bax-Vanluffelen Chair for Alzheimer’s Disease. B.D.S. receives funding from the Medical Research Council, the Alzheimer’s Society and Alzheimer’s Research UK via the Dementia Research Institute. Cell sorting was performed at the KU Leuven FACS core facility, and sequencing was carried out by the Flanders Institute for Biotechnology Nucleomics Core. R.M. recieves funding from Fonds voor Wetenschappelijk Onderzoek (grant no.G0C9219N) and is a recipient of a postdoctoral fellowship from the Alzheimer’s Association USA.
Author information
Authors and Affiliations
Contributions
R.M. conceived and designed the study, performed all of the experiments and wrote the manuscript. J.V.D.D. conceived and designed the study, performed all of the experiments and wrote the manuscript. N.F. conceived and designed the study, performed all of the experiments and wrote the manuscript. L.W. performed all of the experiments. S.B. contributed to the preparation of oligomeric amyloid-β and intracerebral injections. O.B. assisted with the flow cytometry experiments. A.L. contributed to the interpretation of the data. A.S. assisted on human genetics and human-to-mouse orthology. Y.F. assisted with the analysis of single-cell RNA sequencing data. S.P. assisted with the single-cell RNA sequencing experiments. A.A.-M. optimized the xenograft experiments. C.S.F. optimized the single-cell sequencing experiments and library preparations. C.C. assisted with the differentiation of microglia from ESCs. L.S. established and maintained the mouse colonies. T.T. recruited the human subjects, performed the neurosurgeries and provided the human tissue specimens. V.H.P. contributed to the design of the study and interpretation of the data. C.V. contributed to the design of the study and interpretation of the data. M.F. contributed to the design of the study, and analysis and interpretation of the data. B.D.S. conceived and designed the study and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests. B.D.S. receives grants from different companies that support his research and is a consultant for several companies, but nothing is directly related to the current publication.
Additional information
Peer review information Nature Neuroscience thanks Frederick Bennett and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Gating strategy for the isolation of H9-microglia from the mouse brain and graft efficiency.
(a) Human cells were sorted according to the expression of CD11b, hCD45, and GFP, whereas mouse cells only expressed CD11b but were negative for hCD45 and GFP. (b) H9-microglia graft efficiency. Percentage of CD11b cells in the total sample, and proportion of human cells amongst them. Graph shows mean ± s.e.m., n = 6 mice per group.
Extended Data Fig. 2 H9-microglia showed a widespread distribution across multiple areas of the brain.
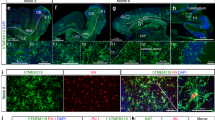
(a) Representative overview of the extent of H9-microglia graft in the mouse brain. Human microglia are stained for P2RY12 across consecutive sections separated by 500 μm to capture multiple anatomical areas. Scale bar, 1mm. (b) Higher magnification images of multiple anatomical areas including meninges, cortex, striatum, white matter, choroid plexus and hippocampus. Labelling shows DAPI (in blue), GFP (in green) and P2RY12 (in cyan). Images are representative of a staining performed in n=4 mice. Scale bar, 100 μm.
Extended Data Fig. 3 Extended clustering and distribution of in vitro, in vivo (engrafted) H9 and primary microglia.
(a) PCA shows clear separation between in vitro (MNC and MG) and in vivo (engrafted H9 and primary) microglia. The colours correspond to the clustering shown in Fig. 2a. (b) t-SNE plots as in Fig. 2a, coloured by the combined level of expression of groups of genes that characterize distinct microglial states. The original clusters from Fig. 2a are outlined. (c) Selected genes defining the different transcriptomic scores shown in b. The full list of genes is shown in Supplementary Table 3. (d) Distribution of the different samples across the tSNE plot, and (e) percentage of each sample across the different clusters. All the data shown represents 2,246 transplanted H9-microglia (in vivo) (n=3/1, 3 mice in 1 combined sequencing pool), 4496 H9-derived monocytes (n=2/1, 2 differentiations in 1 combined sequencing pool) and 3385 microglia in vitro (n=2/1), and 22,846 human primary microglia obtained from cortical surgical resections (n=7/7 Methods).
Extended Data Fig. 4 Direct comparison of in vitro, in vivo (engrafted) H9 and primary microglia.
(a) Volcano plots showing paired comparisons between average gene expression in vitro MNC, in vitro MG, in vivo (engrafted) MG and primary cells. (b) Individual comparisons of in vivo (engrafted) H9-microglia and each human subject (human cases 1–7). The dashed line corresponds to an arbitrary threshold logFC of 0.2. Blue labels correspond to homeostatic genes whereas red labels correspond to microglial activation genes (Supplementary Table 3). All the data shown represents 2,246 transplanted H9-microglia (in vivo) (n=3/1, 3 mice in 1 combined sequencing pool), 4496 H9-derived monocytes (n=2/1, 2 differentiations in 1 combined sequencing pool) and 3385 microglia in vitro (n=2/1), and 22,846 human primary microglia obtained from cortical surgical resections (n=7/7). In all cases, Wilcoxon Rank Sum test, P values adjusted with Bonferroni correction based on the total number of genes in the dataset.
Extended Data Fig. 5 Characterization of oligomeric amyloid-β preparation.
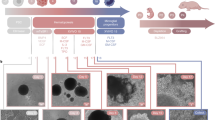
Freshly eluted recombinant amyloid β1–42 monomers follow a rapid aggregation course in Tris-EDTA buffer. (a) After 2 hours of incubation, amyloid β1–42 monomers oligomerize and run as dimers and trimers (indicated with *) on SDS-PAGE/Coomassie staining, and they are proteinase-K sensitive. (b) Early amyloid β1–42 oligomers form A11 and OC-positive aggregates. Two µl of either scrambled or amyloid-β1–42 from different time points (0 hours, 2 hours and 2 weeks) of incubation was spotted on blots. These dot blots were probed with A11 antibody (Invitrogen; #AHB0052), which recognizes amino acid sequence-independent oligomers of proteins or peptides. A11 epitope is transient and is present only in the early oligomers (2 h), in contrast to the mature fibers (2 w) formed after 2 weeks of incubation. No fibrillary material is detected after 2 hours of incubation. (c) OC antibody (Millipore; #AB2286) recognize epitopes common to monomers, amyloid-β oligomers, and fibrils. (d) 4G8 antibody (Eurogentec; #SIG-39220) detects N-terminal of the amyloid-β aggregates (epitope between amino acids 17–24).
Extended Data Fig. 6 Preparation of the datasets for the analysis of (a-c) host mouse and (d-f) H9-microglia response to oligomeric amyloid-β.
(a) t-SNE plot of the 13342 cells passing quality control, coloured by clusters. (b) t-SNE plots as in a, coloured by the level of ln normalized expression of selected genes for microglia (Cx3cr1, Tmem119), monocytes (Ccr2) and neutrophils (Ccrl2). (c) Violin plots of selected marker genes for homeostatic microglia (Cx3cr1, Tmem119), CRM (Il1b), ARM (Cd74, H2-Eb1, Ifit3), neutrophils (Ccrl2), monocytes (Ccr2, Ly6c1), astrocytes (Clu), oligodendrocytes (Mbp), neurons (Npy), and cycling cells (Top2a). Analysis shown in Fig. 2 was performed after removal of clusters 4 (neutrophils), 7 (monocytes), 10 (astrocytes), 12 (oligodendrocytes) and 9 (neurons). (d) t-SNE plot of the 6444 H9-microglia cells passing quality control, coloured by clusters. (e) t-SNE plot as in a, coloured by treatment (naïve; scrambled peptide, Scr; and oligomeric amyloid-β, oAβ). (f) t-SNE plot as in a, coloured by the level of ln normalized expression of selected genes for microglia (CX3CR1), cycling cells (MKI67) and brain resident macrophages (MRC1, CD163). Analysis shown in Fig. 2 was performed after removal of clusters 1 and 4 (brain resident macrophages), 6 (cycling cells) and 8 (doublets).
Extended Data Fig. 7 Expanded analysis, clustering and trajectory inference of the analysis of the response of H9-microglia upon oligomeric amyloid-β.
(a) PCA of 4880 H9-microglia isolated from the mouse brain (n=3 mice in 1 combined sequencing pool) shows clear separation of the different clusters identified in our analysis in PC1 and PC2. (b) t-SNE plots as in Fig. 3a, coloured by the combined level of expression of groups of genes that characterise distinct microglial states. (c) Selected genes defining the different transcriptomic scores shown in Fig. 3b: homeostatic score (1), cytokine score (2) activated score (3). The full list of genes is shown in Supplementary Table 3. (d) Volcano plots showing paired comparisons between H9.HM, H9.CRM and H9.PM clusters (Wilcoxon rank-sum test, P-values adjusted with Bonferroni correction based on the total number of genes in the dataset). (e) Proportion of the different experimental groups across clusters in Fig. 3a (Chi2 test, *** P < 10−250). (f, g) Phenotypic trajectory inferred by Monocle 2 as shown in Fig. 3a, coloured by (f) treatment and (g) clusters from Fig. 3a.
Extended Data Fig. 8 Expanded analysis, clustering and trajectory inference of the analysis of the response of mouse host microglia upon oligomeric amyloid-β.
(a) PCA of 9942 endogenous mouse cells (n=3 mice in 1 combined sequencing pool) shows clear separation of the different clusters identified in our analysis in PC1 and PC2. (b) t-SNE plots as in Fig. 3d, coloured by the combined level of expression of groups of genes that characterise distinct microglial states. (c) Selected genes defining the different transcriptomic scores shown in b: homeostatic score (1), cytokine score (2) activated score (3). The full list of genes is shown in Supplementary Table 3. (d) Volcano plots showing paired comparisons between ms.HM, ms.CRM and ms.ARM clusters (Wilcoxon Rank Sum test, P-values adjusted with Bonferroni correction based on the total number of genes in the dataset). (e) Proportion of the different experimental groups across clusters in Fig. 3d (Chi2 test, *** P < 10−250). (f, g) Phenotypic trajectory inferred by Monocle 2 as shown in Fig. 3c, coloured by (f) treatment and (g) clusters from Fig. 3d.
Extended Data Fig. 9 Cytokine response microglia (CRM) are also present in APPNL-G-F mice.
(a) Original clustering analysis from Sala Frigerio et al. (2019)11 consisting of 10,801 microglial cells from 3 to 21 months old APPNL-G-F mice and aged matched wild type controls. (b) Clusters shown in a, coloured with CRM, HM, ARM and IRM transcriptomic signatures. Note the small population of cells displaying CRM features embeded into the ARM response in APPNL-G-F microglia. (c) Significant enrichment of either homeostatic (HM) or activated (ARM) microglia gene sets from Sala Frigerio et al. (2019)11 in our ms.HM and ms.ARM clusters, respectively (ANOVA with Turkey HSD multiple comparisons correction, *** P ≈ 0; box plots represent median, with 25th and 75th percentiles and 1.5 times the inter-quartile range as minima and maxima). (d) Subselection of CRM cells from the main clusters shown in a. (e) Microglia cells enriched with a CRM transcriptomic profile are largely located at early stages of the response to amyloid-β in APPNL-G-F mice11. The top panel shows the trajectory analysis coloured by clusters as represented in panel a, whereas the bottom panel highlights the cells displaying a CRM profile.
Extended Data Fig. 10 Differential responses of human and host mouse microglia to oligomeric amyloid-β.
(a) Pathway enrichment analysis (GOrilla) shows that the differentially expressed genes in CRM vs. HM clusters are involved in immune and inflammatory processes. (b) Top differentially expressed genes in H9-microglia upon oligomeric amyloid-β challenge relative to scrambled peptide, and expression of their mouse orthologs by endogenous mouse cells. Coloured marks indicate the functional category as shown in b. (c) Differentially expressed genes that show opposite behaviour in H9- and mouse host (Rag2−/− Il2rγ−/−) microglia. Coloured marks indicate the functional category as shown in b. (d) Volcano plots showing paired comparisons between H9.HM, H9.CRM, but including all genes (even those with no clear orthology to mouse, Wilcoxon Rank Sum test, P-values adjusted with Bonferroni correction based on the total number of genes in the dataset). (e) Further pathway enrichment analysis (GOrilla) performed on the human-specific (with no clear orthology) differentially expressed genes in H9.CRM vs. H9.HM clusters are involved in cytokine/chemokine responses.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2.
Supplementary tables
Supplementary Tables 1–4.
Rights and permissions
About this article
Cite this article
Mancuso, R., Van Den Daele, J., Fattorelli, N. et al. Stem-cell-derived human microglia transplanted in mouse brain to study human disease. Nat Neurosci 22, 2111–2116 (2019). https://doi.org/10.1038/s41593-019-0525-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0525-x
This article is cited by
-
Beyond genetics: including the environmental dimension in amyloidosis mouse models for Alzheimer’s disease
Acta Neuropathologica Communications (2024)
-
Regulation of human microglial gene expression and function via RNAase-H active antisense oligonucleotides in vivo in Alzheimer’s disease
Molecular Neurodegeneration (2024)
-
Updates on mouse models of Alzheimer’s disease
Molecular Neurodegeneration (2024)
-
Switch of innate to adaptative immune responses in the brain of patients with Alzheimer’s disease correlates with tauopathy progression
npj Aging (2024)
-
Transcriptional characterization of iPSC-derived microglia as a model for therapeutic development in neurodegeneration
Scientific Reports (2024)