Abstract
Activity-driven transcription plays an important role in many brain processes, including those underlying memory and epilepsy. Here we combine genetic tagging of nuclei and ribosomes with RNA sequencing, chromatin immunoprecipitation with sequencing, assay for transposase-accessible chromatin using sequencing and Hi-C to investigate transcriptional and chromatin changes occurring in mouse hippocampal excitatory neurons at different time points after synchronous activation during seizure and sparse activation by novel context exploration. The transcriptional burst is associated with an increase in chromatin accessibility of activity-regulated genes and enhancers, de novo binding of activity-regulated transcription factors, augmented promoter–enhancer interactions and the formation of gene loops that bring together the transcription start site and transcription termination site of induced genes and may sustain the fast reloading of RNA polymerase complexes. Some chromatin occupancy changes and interactions, particularly those driven by AP1, remain long after neuronal activation and could underlie the changes in neuronal responsiveness and circuit connectivity observed in these neuroplasticity paradigms, perhaps thereby contributing to metaplasticity in the adult brain.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
References
Yap, E. L. & Greenberg, M. E. Activity-regulated transcription: bridging the gap between neural activity and behavior. Neuron 100, 330–348 (2018).
Benito, E. & Barco, A. The neuronal activity-driven transcriptome. Mol. Neurobiol. 51, 1071–1088 (2015).
West, A. E. & Greenberg, M. E. Neuronal activity-regulated gene transcription in synapse development and cognitive function. Cold Spring Harb. Perspect. Biol. 3, a005744 (2011).
Eagle, A. L., Gajewski, P. A. & Robison, A. J. Role of hippocampal activity-induced transcription in memory consolidation. Rev. Neurosci. 27, 559–573 (2016).
Sweatt, J. D. The emerging field of neuroepigenetics. Neuron 80, 624–632 (2013).
Lopez-Atalaya, J. P. & Barco, A. Can changes in histone acetylation contribute to memory formation? Trends Genet. 30, 529–539 (2014).
Kim, T. K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
Savell, K. E. et al. Extra-coding RNAs regulate neuronal DNA methylation dynamics. Nat. Commun. 7, 12091 (2016).
You, X. et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat. Neurosci. 18, 603–610 (2015).
Crepaldi, L. et al. Binding of TFIIIC to sine elements controls the relocation of activity-dependent neuronal genes to transcription factories. PLoS Genet. 9, e1003699 (2013).
Madabhushi, R. et al. Activity-induced DNA breaks govern the expression of neuronal early-response genes. Cell 161, 1592–1605 (2015).
Cho, J. et al. Multiple repressive mechanisms in the hippocampus during memory formation. Science 350, 82–87 (2015).
Mauger, O., Lemoine, F. & Scheiffele, P. Targeted intron retention and excision for rapid gene regulation in response to neuronal activity. Neuron 92, 1266–1278 (2016).
Mo, A. et al. Epigenomic signatures of neuronal diversity in the mammalian brain. Neuron 86, 1369–1384 (2015).
Ben-Ari, Y. & Cossart, R. Kainate, a double agent that generates seizures: two decades of progress. Trends Neurosci. 23, 580–587 (2000).
Elliott, R. C., Miles, M. F. & Lowenstein, D. H. Overlapping microarray profiles of dentate gyrus gene expression during development- and epilepsy-associated neurogenesis and axon outgrowth. J. Neurosci. 23, 2218–2227 (2003).
Huang, Y., Doherty, J. J. & Dingledine, R. Altered histone acetylation at glutamate receptor 2 and brain-derived neurotrophic factor genes is an early event triggered by status epilepticus. J. Neurosci. 22, 8422–8428 (2002).
Tsankova, N. M., Kumar, A. & Nestler, E. J. Histone modifications at gene promoter regions in rat hippocampus after acute and chronic electroconvulsive seizures. J. Neurosci. 24, 5603–5610 (2004).
Crosio, C., Heitz, E., Allis, C. D., Borrelli, E. & Sassone-Corsi, P. Chromatin remodeling and neuronal response: multiple signaling pathways induce specific histone H3 modifications and early gene expression in hippocampal neurons. J. Cell Sci. 116, 4905–4914 (2003).
Taniura, H., Sng, J. C. & Yoneda, Y. Histone modifications in status epilepticus induced by kainate. Histol. Histopathol. 21, 785–791 (2006).
Dougherty, J. D. The expanding toolkit of translating ribosome affinity purification. J. Neurosci. 37, 12079–12087 (2017).
Schott, J. et al. Translational regulation of specific mRNAs controls feedback inhibition and survival during macrophage activation. PLoS Genet. 10, e1004368 (2014).
Yamazaki, S., Muta, T., Matsuo, S. & Takeshige, K. Stimulus-specific induction of a novel nuclear factor-κB regulator, IκB-ζ, via Toll/interleukin-1 receptor is mediated by mRNA stabilization. J. Biol. Chem. 280, 1678–1687 (2005).
Behrens, G. et al. A translational silencing function of MCPIP1/regnase-1 specified by the target site context. Nucleic Acids Res. 46, 4256–4270 (2018).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Su, Y. et al. Neuronal activity modifies the chromatin accessibility landscape in the adult brain. Nat. Neurosci. 20, 476–483 (2017).
Wang, S. et al. Target analysis by integration of transcriptome and ChIP-seq data with BETA. Nat. Protoc. 8, 2502–2515 (2013).
Saha, R. N. et al. Rapid activity-induced transcription of Arc and other IEGs relies on poised RNA polymerase II. Nat. Neurosci. 14, 848–856 (2011).
Schep, A. N. et al. Structured nucleosome fingerprints enable high-resolution mapping of chromatin architecture within regulatory regions. Genome Res. 25, 1757–1770 (2015).
Wang, Y., Lu, J. J., He, L. & Yu, Q. Triptolide (TPL) inhibits global transcription by inducing proteasome-dependent degradation of RNA polymerase II (Pol II). PLoS One 6, e23993 (2011).
Jonkers, I., Kwak, H. & Lis, J. T. Genome-wide dynamics of Pol II elongation and its interplay with promoter proximal pausing, chromatin, and exons. eLife 3, e02407 (2014).
Creyghton, M. P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl Acad. Sci. USA 107, 21931–21936 (2010).
Malik, A. N. et al. Genome-wide identification and characterization of functional neuronal activity–dependent enhancers. Nat. Neurosci. 17, 1330–1339 (2014).
Hardingham, G. E., Pruunsild, P., Greenberg, M. E. & Bading, H. Lineage divergence of activity-driven transcription and evolution of cognitive ability. Nat. Rev. Neurosci. 19, 9–15 (2018).
Guzowski, J. F. et al. Mapping behaviorally relevant neural circuits with immediate-early gene expression. Curr. Opin. Neurobiol. 15, 599–606 (2005).
Lacar, B. et al. Nuclear RNA-seq of single neurons reveals molecular signatures of activation. Nat. Commun. 7, 11022 (2016).
Tai, X. Y. et al. Hyperphosphorylated tau in patients with refractory epilepsy correlates with cognitive decline: a study of temporal lobe resections. Brain 139, 2441–2455 (2016).
Halder, R. et al. DNA methylation changes in plasticity genes accompany the formation and maintenance of memory. Nat. Neurosci. 19, 102–110 (2016).
Rowley, M. J. & Corces, V. G. Organizational principles of 3D genome architecture. Nat. Rev. Genet. 19, 789–800 (2018).
Bonev, B. et al. Multiscale 3D genome rewiring during mouse neural development. Cell 171, 557–572.e524 (2017).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Xiao, M. F. et al. NPTX2 and cognitive dysfunction in Alzheimer’s disease. eLife 6, e23798 (2017).
Fernandes, D. & Carvalho, A. L. Mechanisms of homeostatic plasticity in the excitatory synapse. J. Neurochem. 139, 973–996 (2016).
Hampsey, M., Singh, B. N., Ansari, A., Laine, J. P. & Krishnamurthy, S. Control of eukaryotic gene expression: gene loops and transcriptional memory. Adv. Enzym. Regul. 51, 118–125 (2011).
Laine, J. P., Singh, B. N., Krishnamurthy, S. & Hampsey, M. A physiological role for gene loops in yeast. Genes Dev. 23, 2604–2609 (2009).
DeNardo, L. & Luo, L. Genetic strategies to access activated neurons. Curr. Opin. Neurobiol. 45, 121–129 (2017).
Maurano, M. T. et al. Large-scale identification of sequence variants influencing human transcription factor occupancy in vivo. Nat. Genet. 47, 1393–1401 (2015).
Vierbuchen, T. et al. AP-1 transcription factors and the BAF complex mediate signal-dependent enhancer selection. Mol. Cell 68, 1067–1082.e1012 (2017).
Del Blanco, B. et al. CBP and SRF co-regulate dendritic growth and synaptic maturation. Cell Death Differ. https://doi.org/10.1038/s41418-019-0285-x (2019).
Scandaglia, M. et al. Loss of Kdm5c causes spurious transcription and prevents the fine-tuning of activity-regulated enhancers in neurons. Cell Rep. 21, 47–59 (2017).
Erdmann, G., Schutz, G. & Berger, S. Inducible gene inactivation in neurons of the adult mouse forebrain. BMC Neurosci. 8, 63 (2007).
Stanley, S. et al. Profiling of glucose-sensing neurons reveals that GHRH neurons are Activated by hypoglycemia. Cell Metab. 18, 596–607 (2013).
Fiorenza, A. et al. Blocking miRNA Biogenesis in adult forebrain neurons enhances seizure susceptibility, fear memory, and food intake by increasing neuronal responsiveness. Cereb. Cortex 26, 1619–1633 (2016).
Vogel-Ciernia, A. et al. The neuron-specific chromatin regulatory subunit BAF53b is necessary for synaptic plasticity and memory. Nat. Neurosci. 16, 552–561 (2013).
Heiman, M., Kulicke, R., Fenster, R. J., Greengard, P. & Heintz, N. Cell type-specific mRNA purification by translating ribosome affinity purification (TRAP). Nat. Protoc. 9, 1282–1291 (2014).
Lopez-Atalaya, J. P., Ito, S., Valor, L. M., Benito, E. & Barco, A. Genomic targets, and histone acetylation and gene expression profiling of neural HDAC inhibition. Nucleic Acids Res. 41, 8072–8084 (2013).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R., & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Bauer-Mehren, A., Rautschka, M., Sanz, F. & Furlong, L. I. DisGeNET: a Cytoscape plugin to visualize, integrate, search and analyze gene-disease networks. Bioinformatics 26, 2924–2926 (2010).
Queralt-Rosinach, N., Pinero, J., Bravo, A., Sanz, F. & Furlong, L. I. DisGeNET-RDF: harnessing the innovative power of the Semantic Web to explore the genetic basis of diseases. Bioinformatics 32, 2236–2238 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Ye, T. et al. seqMINER: an integrated ChIP-seq data interpretation platform. Nucleic Acids Res. 39, e35 (2011).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Pollard, K. S., Hubisz, M. J., Rosenbloom, K. R. & Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 20, 110–121 (2010).
Piper, J. et al. Wellington: a novel method for the accurate identification of digital genomic footprints from DNase-seq data. Nucleic Acids Res. 41, e201 (2013).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution hi-C experiments. Cell Syst. 3, 95–98 (2016).
Ay, F., Bailey, T. L. & Noble, W. S. Statistical confidence estimation for Hi-C data reveals regulatory chromatin contacts. Genome Res. 24, 999–1011 (2014).
Acknowledgements
The authors thank E. Herrera, T. Ferrar, J. P. Lopez-Atalaya and Y. Ruan for their critical reading of the manuscript, and R. Olivares, N. Cascales, A. Caler and the personnel of the sequencing facility at the CRG (Barcelona, Spain) and HudsonAlpha (Alabama, USA) for technical assistance. J.F.-A. and M.T.L.-C. are recipients of fellowships from the Spanish Ministry of Science and Innovation (MICINN, SVP-2014-068387 and BES-2017-081298, respectively). The research of A.B. is supported by grants SAF2017-87928-R and SEV-2017-0723 from the MICINN co-financed by the ERDF, PROMETEO/2016/026 from the Generalitat Valenciana and RGP0039/2017 from the Human Frontiers Science Program Organization (HFSPO). M.J.R. is supported by the National Institutes of Health (NIH) Pathway to Independence Award K99/R00 GM127671. The research of V.G.C. is supported by the US Public Health Service Award (R01) GM035463 from the NIH. The content of the article is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. The Instituto de Neurociencias is a “Centre of Excellence Severo Ochoa”.
Author information
Authors and Affiliations
Contributions
J.F.-A. performed most of the experiments and bioinformatic analyses. M.L. collaborated in the characterization of the models and performed ChIP-seq experiments. M.T.L.-C. collaborated in the bioinformatic analyses and preparation of the figures. M.J.R. conducted the Hi-C analyses. A.M.M.-G. collaborated in the preparation of Hi-C samples and performed some of the immunostaining analyses. B.d.B. provided important reagents. V.G.C. supervised the Hi-C experiments and analyses. A.B. supervised all other aspects of the project. A.B. and J.F.-A. conceived the study, designed the experiment and wrote the original draft. All the authors discussed the findings and revised the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Neuroscience thanks Schahram Akbarian, Michael Greenberg, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Integrated supplementary information
Supplementary Figure 1 KA-induced neuronal activation causes broad changes in the nuclear transcriptome.
a. Step by step analysis and channel filtering (percentage of the filtered population indicated within the channel) of flow cytometry signal for the specific isolation of fluorescent singlet nuclei. Shown the comparison of intrinsic fluorescence in Sun1-GFP- (above) vs Sun1-GFP+ (below) mice. Similar results were obtained in 25 independent experiments. b. Images of isolated nuclei immunostained with antibodies against GFP and the neuronal marker NeuN. Similar results were obtained in 3 independent experiments. c. Comparison of nuclear size of Sun1- GFP+ (green) and wildtype nuclei stained with NeuN (pink). Similar results were obtained in 3 independent experiments. d. RT-qPCR in nuclear RNA showing Fos transcript levels in GFP positive nuclei after KA-induced SE (levels referred to GAPDH expression). e. ChIP-qPCR assay analysis for H3K4me3 in GFP positive and negative nuclei showing FANS specificity (values are normalized to input). Hchrom: heterochromatin; pr: promoter. Sun1-GFP(-) n = 3; Sun1-GFP(+) n = 3, biologically independent samples. Bars indicate mean ± s.e.m. Two-sided t-test, **: p-value < 0.005. f. ATAC-qPCR assay showing specificity and promoter changes 1 h after KA-induced neuronal activation in GFP positive nuclei (levels referred to the Gapdh promoter). Hchrom: heterochromatin; pr: promoter. g. Pearson correlation matrix between normalized samples clustered by Euclidian dendrogram. Sal n = 2; KA-1h n = 2, biologically independent samples. h. Percentage of nuRNA-seq reads aligned into exons and introns. For comparison, we also present the values for riboRNA-seq samples. i. GO enrichment analysis for protein-coding DETs that are activity-induced (AI, red bars) or activity-depleted (AD, blue bars) by SE after PANTHER analysis. The number of genes associated with each GO term is indicated. Sal n = 2; KA-1h n = 2, biologically independent samples.
Supplementary Figure 2 SE-induced RNA translation activates nucleus-related functions.
a. Confocal images of principal hippocampal neurons in the DG and CA1 subfield of CaMKII-creERT2::GFP-L10a mice stained against GFP (green) and the interneuron marker GAD67 (red). Cells positive for GAD67 do not express GFP. Similar results were obtained in 3 independent experiments. b. RT-qPCR assay comparing transcript levels before IP (Bulk) and after immunoprecipitation (IP). Note that fold changes for IEG induction were larger after TRAPping. Sal n = 3; KA-1h n = 3, biologically independent samples. Bars indicate mean ± s.e.m. 2-way ANOVA p-values: * = treatment effect (KA 1h vs Sal); # = procedure effect (IP vs Bulk). */#: p < 0.05; **/##: p < 0.01; ***/###: p < 0.001; ns: non significant. c. Pearson correlation matrix between normalized riboRNA-seq samples clustered by Euclidian dendrogram. Sal n = 3; KA-1h n = 3, biologically independent samples. d. Principal component analysis of riboRNA-seq samples. Sal n = 3; KA-1h n = 3, biologically independent samples. e. Genomic snapshots for each replicate riboRNA-seq track at representative examples of non-neuronal (Plp1), housekeeping (ActB) and activity-induced (Fos) genes. Values indicate the number of counts in RPM. f. Gene biotype classification for DTGs 1h after KA. g. GO enrichment analysis for protein-coding genes displaying enhanced (red bars) or reduced (blue bars) translation upon SE after PANTHER analysis. The number of genes associated with each GO term is indicated. Sal n = 3; KA-1h n = 3, biologically independent samples. h. Comparison of nuRNAseq and riboRNAseq tracks for representative transcripts that are either detected as upregulated or downregulated in both screens (but showing largest changes in nuRNA-seq), or that show discordant changes in the two screens. Values indicate the levels of counts in RPM. i. Metagene mapability profiles for nuRNA-seq at a group of constitutive highly expressed genes. Compare the region downstream of the TTS for this group and the IEGs presented in Fig. 2k; the two groups contain the same number of genes.
Supplementary Figure 3 Transcriptional bursting increases accessibility at IEGs.
a. Fragment size distribution in ATAC-seq libraries. b. ATAC-seq signal at the TSS of highly expressed genes in NeuN+ vs NeuN- CA1 cells (Halder et al. 2016, Nat Neurosci 19, 102-10). c. PCA of accessibility profiles shows similarity between cortical (Mo et al. 2015, Neuron 86, 1369-84) and hippocampal excitatory neurons (this study). CTX_exc: cortical excitatory neurons, n = 2 ; CTX_VIP: vasoactive intestinal peptide-expressing interneurons, n = 2; CTX_PV: parvalbumin-expressing interneurons, n = 2; HIPP_exc: hippocampal excitatory neurons, n = 2, biologically independent samples. d. Percentage of reads mapped into nuclear and mitochondrial chromosomes in this study and (Su et al. 2017, Nat Neurosci 20, 476-83). e. Genome coverage in this study and (Su et al. 2017, Nat Neurosci 20, 476-83). E0 and E1: chromatin profiling coverage before (E0) and 1 h after synchronous electrical neuronal activation (E1). f. Comparison of genomic profiles at control and activity-regulated genes, including an extended view of the Npas4 locus. Discrepancies may result from the superior genomic coverage and intrinsic differences between stimulation paradigms (chemical vs. electrical) and composition of the samples (hippocampal glutamatergic neurons nuclei vs. microdissected DG tissue). g. Distribution of genomic features along differentially accessible regions.
Supplementary Figure 4 Correlation of changes in chromatin accessibility and transcription.
a. Predictive analysis of activating/repressive function for accessibility changes at regions displaying increased (IA) or reduced accessibility (RA) within the gene body and correlation with changes in nuclear transcript levels. b. Left: Overlap between activity-induced genes detected in the nuRNA-seq and riboRNA-seq screens that show increased accessibility. Right: Heatmap representation of changes in accessibility and transcript levels in the 119 overlapping genes. c. Scatter plots showing the correlation between chromatin accessibility changes at gene bodies (Y-axis) and transcript levels determined by DESeq2 analysis (X-axis: left graph, riboRNA; right graph, nucRNA). Blue dots label genes significantly regulated by KA (FDR < 0.1). riboRNA: Sal n = 3; KA-1h n = 3; nuRNA: Sal n = 2; KA-1h n = 2, biologically independent samples. The Pearson’s correlation indexes are shown d. Metagene plot of RNAPII occupancy in AI genes (nuRNA-seq) split according to increased gene body accessibility (FDR > 0.1) after DESeq2 analysis. RNAPII: Sal n = 1, KA-1h n = 1. e. RT-qPCR assays show the inhibition of IEG induction by TPL after KA-induced neuronal activation. Sal n = 4; KA-1h n = 4; TPL+KA-1h n = 3, biologically independent samples; p < 0.001 for the three genes. Bars indicate mean ± s.e.m. f. Metagene plot of ATAC-seq signal in Sal, KA-1h and KA+TPL samples; genes were split as in panel d. g. TPL effect on the accessibility at gene bodies and TSSs of activity-regulated genes.
Supplementary Figure 5 Activity-dependent TF-biding.
a. BETA analysis of accessibility changes at extragenic regions and associated changes in expression. IA: increased accessibility; RA: reduced accessibility. b. Bar plot comparing the percentages of DARs at accessible promoters and enhancers. Upper sector plots indicate percentage of strong/weak signal at IA and RA regions. c. Signal for RNAPII and CBP binding 1 h after KA. We compare IA and RA regions; IA regions are split between weak and strong promoters/enhancers. The solid lines indicate the mean and the shaded lines the s.e.m. RNAPII: Sal n = 1, KA-1h n = 1; CBP: Sal n = 1, KA-1h n = 2, biologically independent samples. d. Signal of activity regulated profiles for H3K27ac 1 h after KCl (Malik et al. 2014, Nat Neurosci 17, 1330-39) using the same classification than in panel c. e. Signal for RNAPII and CBP 1h after KA in transcriptional-dependent and independent regions. The solid lines indicate the mean and the shaded lines the s.e.m. RNAPII: Sal n = 1, KA-1h n=1; CBP: Sal n = 1, KA-1h n = 2, biologically independent samples. f. Signal of H3K27ac 1 h after KCl (Malik et al. 2014, Nat Neurosci 17, 1330-39) using the same classification than in panel e. The solid lines indicate the mean and the shaded lines the s.e.m. Untreated n = 2; KCl n = 2, biologically independent samples. g. Violin plots show the log2FC distribution of activity-regulated genes in KA-1h annotated with reduced (RA) or increased accessibility (IA) sites at promoters and enhancers. Note the detection of opposing changes in chromatin accessibility and transcript levels suggests the activity-dependent release of transcriptional repressors. Sector plot shows the percentage of strong and weak enhancers annotated at those genes. The boxplot indicates the median, interquartile range and min./max. Dots are colored in according to their weak or strong enhancer annotation. nuRNA: Sal n = 2, KA-1h n = 2; ATAC-seq: Sal n = 2, KA-1h n = 2, biologically independent samples. h. Bar plot comparing the percentage of promoter/enhancer DARs annotated to nuRNA-seq regulated genes (FDR < 0.1) after DESeq2 analysis. Upper sector plots show percentage of activity-induced (AI) and depleted (AD) genes associated with IA and RA regions. nuRNA: Sal n = 2, KA-1h n = 2; ATAC-seq: Sal n = 2, KA-1h n = 2, biologically independent samples.
Supplementary Figure 6 Activity-dependent changes of TF footprints.
a. Genomic snapshot of nuRNAs, ATAC-seq, RNAPII and CBP binding in saline, KA-1h and TPL+KA-1h samples at the Npas4 and Nr4a1 loci (values in RPM). Annotations label the detected footprints (in red, less stringent footprints) and classification for the regions (blue: promoters; gray: enhancers). Zoom-in inset shows upstream eRNA activity and H3K27 acetylation profile. b. Motif enrichment at transcription-dependent and -independent enhancers for the indicated activity-regulated TFs. c. Plots compare the digital footprint at AP1 and CTCF motifs in saline, KA-1h and TPL+KA-1h datasets (value correspond to normalized tn5 insertions). The bottom numbers correspond to the motifs detected in KA-1h. d. Signal for CREB, SRF and Fos binding 1 h after KCl stimulation of neuronal cultures (Malik et al. 2014, Nat Neurosci 17, 1330-39) at the detected footprints 1 h after KA in IA promoters/enhancers. The solid lines indicate the mean and the shaded lines the s.e.m. CREB: unt n = 2, KCl n = 2; SRF: unt n = 2, KCl n = 2; Fos: unt n = 2, KCl n = 2. e. Conservation score for the detected footprints at IA promoter/enhancer regions. The shaded lines indicate the s.e.m.
Supplementary Figure 7 Chromatin changes upon physiological neuronal activation.
a. Number of Fos+ nuclei in the dentate gyrus of mice in their home cage (HC) and after 1 h of novelty exploration (NE). HC n = 2, NE n = 2, biologically independent samples. Bars indicate mean ± s.e.m. Two-sided t-test, **: p-value < 0.005. b. Flow cytometry analysis of populations by its fluorescent intensities with the presence or absence of Fos (percentage of the filtered population indicated within the channel) after channel filtering following our protocol for the specific isolation of fluorescent singlet nuclei in Supplementary Fig. 2a. Right panel it is the co-localization of the size distribution of Fos+ (purple) and GFP+ cells (gray). Similar results were obtained in 3 independent experiments. c. Heat-map comparing DARs from SE and NE datasets, indicating common and exclusive regions between conditions. Last column is indicating the ratio of genic/intergenic regions on each subset. d. Principal component analysis for ATAC-seq datasets form physiologically (Fos+ vs Fos- neurons) and chemically activated (Sal vs KA-1h) neurons. Sal n = 2, KA-1h n = 2, Fos+ n = 1, Fos- n = 1, biologically independent samples. e. Volcano plot showing the significance value distribution after differential accessible regions after DESeq2 analysis in Fos- neurons vs neurons from saline-treated mice. Sal n = 2, Fos- n = 1. f. Predictive significance of accessibility changes at the regions annotated as promoter or enhancer and the change in expression of the gene after basic Activating/Repressive Function Prediction in BETA analysis. AI: activity-induced genes; AD: activity-depleted genes; IA: increased accessibility regions; RA: reduced accessibility regions. Single nucleus RNA-seq data from Lacar et al. 2016 (Nat Comm 7, 11022): NE n = 96, HC n = 23; ATAC-seq: Fos+ n = 1, Fos- n = 1. g. Bar plot indicating the percentage of DARs in NE dataset at promoter and enhancer regions. h. Signal of activity regulated profiles 1 h after KA for RNAPII and CBP, comparing exclusive regions at detected enhancers’ footprints in NE and SE. i. Signal for H3K4me1, H3K27ac and DNA methylation after fear conditioning (Halder et al. 2016, Nat Neurosci 19, 102-10) at detected NE-responding enhancers footprints.
Supplementary Figure 8 Chromatin accessibility dynamics shows changes that stay long after the transcriptional burst.
a. Pearson correlation matrix between normalized samples clustered by Euclidian dendrogram for nuRNA-seq. Sal n = 2, KA-1h n = 2, KA-6h n = 2, KA-48h n = 2, biologically independent samples. b. Volcano plot showing the significance of differential gene expression by DESeq2 analysis in nuRNA-seq samples 1 h, 6 h and 48 h after KA administration. AI: activity-induced; AD: activity-depleted. Sal n = 2, KA-1h n = 2, KA-6h n = 2, KA-48h n = 2. c. Venn diagram presenting the overlaps between the sets of DETs at each time point in the longitudinal nuRNA-seq analysis. Almost 20% of the changes observed 1 h after KA-treatment are still detected 5 h later, whereas only 2% remain 2 days later. d. Heatmap for FC in nuRNA-seq analysis. e. Heatmap of fold enrichment for biological process GO terms detected in AI and AD genes by PANTHER analysis in the nuRNA-seq longitudinal analysis using DESeq2. The numbers indicate the number of genes associated with each term. Sal n = 2, KA-1h n = 2, KA-6h n = 2, KA-48h n = 2, biologically independent samples. f. Heatmap comparing DARs at the different time points. The last column (black/ yellow) shows the ratio of genic/intergenic regions on each subset. IA: increased accessibility; RA: reduced accessibility g. Pearson correlation matrix between normalized samples clustered by Euclidian dendrogram for ATAC-seq. Sal n = 2, KA-1h n = 2, KA-6h n = 2, KA-48h n = 2. h. Volcano plot showing the significance value of differential accessible regions after DESeq2 analysis in ATAC-seq samples 1 h, 6 h and 48 h after KA administration. IA: increased accessibility; RA: reduced accessibility. Sal n = 2, KA-1h n = 2, KA-6h n = 2, KA-48h n = 2. i. Venn diagram presenting the overlaps between the sets of DARs at each time point in the longitudinal ATAC-seq analysis.
Supplementary Figure 9 Long-lasting chromatin accessibility changes are associated with AP1 binding and disease.
a. Signal for Fos binding 1 h after KCl stimulation of neuronal cultures (Malik et al. 2014, Nat Neurosci 17, 1330-39) at the detected footprints in enhancer regions showing long-lasting changes in accessibility. b. Plot shows the footprinted sites at the maintained AP1 motifs detected in our longitudinal analysis (profile indicate the normalized tn5 insertions). c. Immunostaining for Fos in granular neurons at the DG and pyramidal neurons at the CA1 subfield at different time points after KA. Similar results were obtained in 3 independent experiments. d. Top: Scheme of object location memory (NOL) test. The time intervals from KA-induced SE to NOL training and testing are indicated. Bottom: Preference index for the displaced object in naïve mice and in mice that suffered SE one week earlier in the NOL test. One-sided t-test (compared to 0) *: p-value < 0.05. Furthermore, saline-treated mice show a larger discrimination index during testing than training (p = 0.03), while the mice that received KA do not show a significant difference (p = 0.56). Sal n = 8, KA-1h n = 9, biologically independent samples. Bars indicate mean ± s.e.m. e. RT-qPCR assay for Fos and Npas4 in RNA samples extracted 1 h after NE in animals that suffered SE two weeks earlier. FC values are referred to the home cage situation. These two IEGs are strongly induced during SE but only Fos is in the proximity of DARs showing long-lasting changes in accessibility. HC: Sal n = 4, KA-1h n = 3; NE: Sal n = 5, KA-1h n = 5, biologically independent samples. Bars indicate mean ± s.e.m. Two-sided t-test, *: p-value < 0.05, **: p-value < 0.01, ***: p-value < 0.001. f. Genomic screenshots for the Fos/Jdp2 locus (an example of activity-regulated gene that returns to basal expression but still maintains some proximal chromatin accessibility changes after 48 h). Footprint motifs for SRF, AP1, CRE, MEF2C are labeled. DARs are colored depending on their location in a promoter or enhancer. The maintained DARs after 48 h are labeled in green. Values show counts in RPM.
Supplementary Figure 10 Hi-C analysis of activity-driven interactions.
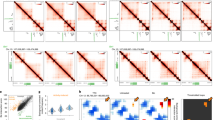
a. Pearson correlation between Hi-C replicate samples at 0 h (saline), 1 h and 48 h after KA administration. Sal n = 2, KA-1h n = 2, KA-48h n = 2, biologically independent samples. Comparison between all possible (over 600 millions) intrachromosomal bins. b. Comparison of compartmental eigenvectors in mature excitatory principal neurons (PN) and cortical neurons differentiated in vitro (iv-CN) (Bonev et al. 2017, Cell 171, 557-72). R = 0.75 Pearson correlation (n = 101,812 loci). c. Difference in gene expression between PN and iv-CN compared to embryonic stem cells (ES) (Bonev et al. 2017, Cell 171, 557-72). R = 0.69 Pearson correlation (n = 23,418 loci). d. Difference in the compartmental eigenvectors in cortical neurons (pink) and principle neurons (blue) vs embryonic stem cells for genes that are higher in ES vs iv-CN (ES vs CN) or vice versa (CN vs ES) (Bonev et al. 2017, Cell 171, 557-72). The boxplot indicates the median, interquartile range and min./max. Two-sided Wilcoxon rank sum test. Left side: n = 5,241. Right side: n = 14,735. All p-values are < 2.2e-16 (lower than the lowest calculated by R). e. Metaplot of CTCF loops where at least one anchor overlaps a differential ATAC-seq peak after DESeq2 analysis indicating that CTCF loops do not change during neuronal activation despite changes to other transcription factors. IA: increased accessibility, RA: reduced accessibility. Sample sizes are indicated at the right of the heatmaps. f. Venn Diagram of CTCF loops in KA-1h and saline samples. The low number of changes (< 4%, below the FDR level) indicates no real change. g. Snapshot of a locus presenting strong de novo Hi-C interactions in response to SE. Significant differences were tested for enrichment in KA-1h samples using a one-sided confidence interval using all interaction bins in the immediate vicinity (n = 81). Significance was assigned if fold change was above a 99.9% confidence interval giving rise to p < 0.001. Hi-C changes correlate with the reported changes in nuRNA production and chromatin accessibility. h. Heatmap of Hi-C interactions at genes with low or high nuRNA-seq signal (Hi-C signal normalized by distance). i. Snapshot for Bdnf (an example of activity-regulated gene that return to basal expression). The central area is shown with greater detail in Fig. 7g. Values show counts in RPM. Significant differences were tested and assigned as indicated in panel g (n = 16). j. GO enrichment after PANTHER analysis for the set of genes associated with long-lasting Hi-C interactions after Fit-Hi-C analysis. k. Total Fit-Hi-C interactions (gray) and their changes after 1 h (x-axis) or 48 h (y-axis) compared to 0 h. Dots correspond to the interactions that show more than a 4-fold increase (red) or decrease (blue) at 1 h. Most blue points are slightly below the horizontal, while most red points are above the horizontal, indicating that some of the SE-originated interactions are still maintained after 48 h. Pearson’s correlation index is shown (n = 148,732 loci).
Supplementary information
Supplementary Table 1
Differentially expressed transcripts in nuRNA-seq.
Supplementary Table 2
Differentially translated genes in riboRNA-seq.
Supplementary Table 3
Differentially accessed regions in ATAC-seq and comparison with DETs in nuRNA-seq (a) and DGTs in riboRNA-seq (b) and impact of TPL treatment (c): 3 sheets.
Supplementary Table 4
ATAC-seq analysis upon novelty exploration.
Supplementary Table 5
Longitudinal analysis of differentially expressed transcripts in nuRNA-seq.
Supplementary Table 6
Longitudinal analysis of chromatin accessibility changes.
Supplementary Table 7
Fit Hi-C values at different time points after SE.
Supplementary Table 8
NGS samples and datasets generated in this study (a) and generated in other studies and analyzed here (b): 2 sheets.
Rights and permissions
About this article
Cite this article
Fernandez-Albert, J., Lipinski, M., Lopez-Cascales, M.T. et al. Immediate and deferred epigenomic signatures of in vivo neuronal activation in mouse hippocampus. Nat Neurosci 22, 1718–1730 (2019). https://doi.org/10.1038/s41593-019-0476-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0476-2
This article is cited by
-
Kdm1a safeguards the topological boundaries of PRC2-repressed genes and prevents aging-related euchromatinization in neurons
Nature Communications (2024)
-
Characterization of transcriptional profiles associated with stress-induced neuronal activation in Arc-GFP mice
Molecular Psychiatry (2024)
-
Using deep learning to quantify neuronal activation from single-cell and spatial transcriptomic data
Nature Communications (2024)
-
Synaptically-targeted long non-coding RNA SLAMR promotes structural plasticity by increasing translation and CaMKII activity
Nature Communications (2024)
-
Cell-type specific transcriptional adaptations of nucleus accumbens interneurons to amphetamine
Molecular Psychiatry (2023)